Human activity recognition is the problem of classifying sequences of accelerometer data recorded by specialized harnesses or smart phones into known well-defined movements.
Classical approaches to the problem involve hand crafting features from the time series data based on fixed-sized windows and training machine learning models, such as ensembles of decision trees. The difficulty is that this feature engineering requires strong expertise in the field.
Recently, deep learning methods such as recurrent neural networks like as LSTMs and variations that make use of one-dimensional convolutional neural networks or CNNs have been shown to provide state-of-the-art results on challenging activity recognition tasks with little or no data feature engineering, instead using feature learning on raw data.
In this tutorial, you will discover three recurrent neural network architectures for modeling an activity recognition time series classification problem.
After completing this tutorial, you will know:
- How to develop a Long Short-Term Memory Recurrent Neural Network for human activity recognition.
- How to develop a one-dimensional Convolutional Neural Network LSTM, or CNN-LSTM, model.
- How to develop a one-dimensional Convolutional LSTM, or ConvLSTM, model for the same problem.
Kick-start your project with my new book Deep Learning for Time Series Forecasting, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.

How to Develop RNN Models for Human Activity Recognition Time Series Classification
Photo by Bonnie Moreland, some rights reserved.
Tutorial Overview
This tutorial is divided into four parts; they are:
- Activity Recognition Using Smartphones Dataset
- Develop an LSTM Network Model
- Develop a CNN-LSTM Network Model
- Develop a ConvLSTM Network Model
Activity Recognition Using Smartphones Dataset
Human Activity Recognition, or HAR for short, is the problem of predicting what a person is doing based on a trace of their movement using sensors.
A standard human activity recognition dataset is the ‘Activity Recognition Using Smart Phones Dataset’ made available in 2012.
It was prepared and made available by Davide Anguita, et al. from the University of Genova, Italy and is described in full in their 2013 paper “A Public Domain Dataset for Human Activity Recognition Using Smartphones.” The dataset was modeled with machine learning algorithms in their 2012 paper titled “Human Activity Recognition on Smartphones using a Multiclass Hardware-Friendly Support Vector Machine.”
The dataset was made available and can be downloaded for free from the UCI Machine Learning Repository:
The data was collected from 30 subjects aged between 19 and 48 years old performing one of six standard activities while wearing a waist-mounted smartphone that recorded the movement data. Video was recorded of each subject performing the activities and the movement data was labeled manually from these videos.
Below is an example video of a subject performing the activities while their movement data is being recorded.
The six activities performed were as follows:
- Walking
- Walking Upstairs
- Walking Downstairs
- Sitting
- Standing
- Laying
The movement data recorded was the x, y, and z accelerometer data (linear acceleration) and gyroscopic data (angular velocity) from the smart phone, specifically a Samsung Galaxy S II. Observations were recorded at 50 Hz (i.e. 50 data points per second). Each subject performed the sequence of activities twice; once with the device on their left-hand-side and once with the device on their right-hand side.
The raw data is not available. Instead, a pre-processed version of the dataset was made available. The pre-processing steps included:
- Pre-processing accelerometer and gyroscope using noise filters.
- Splitting data into fixed windows of 2.56 seconds (128 data points) with 50% overlap.Splitting of accelerometer data into gravitational (total) and body motion components.
Feature engineering was applied to the window data, and a copy of the data with these engineered features was made available.
A number of time and frequency features commonly used in the field of human activity recognition were extracted from each window. The result was a 561 element vector of features.
The dataset was split into train (70%) and test (30%) sets based on data for subjects, e.g. 21 subjects for train and nine for test.
Experiment results with a support vector machine intended for use on a smartphone (e.g. fixed-point arithmetic) resulted in a predictive accuracy of 89% on the test dataset, achieving similar results as an unmodified SVM implementation.
The dataset is freely available and can be downloaded from the UCI Machine Learning repository.
The data is provided as a single zip file that is about 58 megabytes in size. The direct link for this download is below:
Download the dataset and unzip all files into a new directory in your current working directory named “HARDataset”.
Need help with Deep Learning for Time Series?
Take my free 7-day email crash course now (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
Develop an LSTM Network Model
In this section, we will develop a Long Short-Term Memory network model (LSTM) for the human activity recognition dataset.
LSTM network models are a type of recurrent neural network that are able to learn and remember over long sequences of input data. They are intended for use with data that is comprised of long sequences of data, up to 200 to 400 time steps. They may be a good fit for this problem.
The model can support multiple parallel sequences of input data, such as each axis of the accelerometer and gyroscope data. The model learns to extract features from sequences of observations and how to map the internal features to different activity types.
The benefit of using LSTMs for sequence classification is that they can learn from the raw time series data directly, and in turn do not require domain expertise to manually engineer input features. The model can learn an internal representation of the time series data and ideally achieve comparable performance to models fit on a version of the dataset with engineered features.
This section is divided into four parts; they are:
- Load Data
- Fit and Evaluate Model
- Summarize Results
- Complete Example
Load Data
The first step is to load the raw dataset into memory.
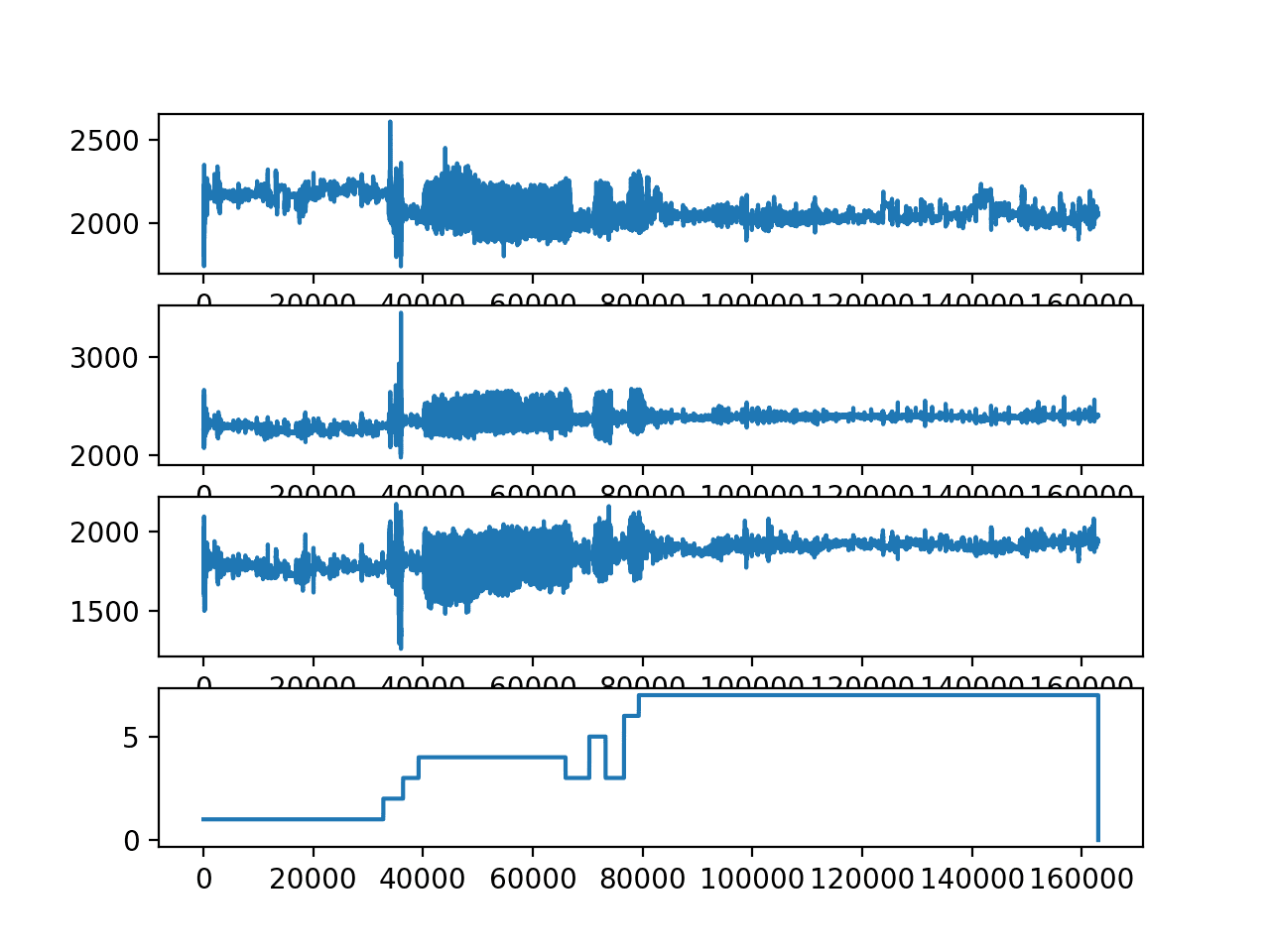
There are three main signal types in the raw data: total acceleration, body acceleration, and body gyroscope. Each has 3 axises of data. This means that there are a total of nine variables for each time step.
Further, each series of data has been partitioned into overlapping windows of 2.56 seconds of data, or 128 time steps. These windows of data correspond to the windows of engineered features (rows) in the previous section.
This means that one row of data has (128 * 9), or 1,152 elements. This is a little less than double the size of the 561 element vectors in the previous section and it is likely that there is some redundant data.
The signals are stored in the /Inertial Signals/ directory under the train and test subdirectories. Each axis of each signal is stored in a separate file, meaning that each of the train and test datasets have nine input files to load and one output file to load. We can batch the loading of these files into groups given the consistent directory structures and file naming conventions.
The input data is in CSV format where columns are separated by whitespace. Each of these files can be loaded as a NumPy array. The load_file() function below loads a dataset given the fill path to the file and returns the loaded data as a NumPy array.
|
1 2 3 4 |
# load a single file as a numpy array def load_file(filepath): dataframe = read_csv(filepath, header=None, delim_whitespace=True) return dataframe.values |
We can then load all data for a given group (train or test) into a single three-dimensional NumPy array, where the dimensions of the array are [samples, time steps, features].
To make this clearer, there are 128 time steps and nine features, where the number of samples is the number of rows in any given raw signal data file.
The load_group() function below implements this behavior. The dstack() NumPy function allows us to stack each of the loaded 3D arrays into a single 3D array where the variables are separated on the third dimension (features).
|
1 2 3 4 5 6 7 8 9 |
# load a list of files into a 3D array of [samples, timesteps, features] def load_group(filenames, prefix=''): loaded = list() for name in filenames: data = load_file(prefix + name) loaded.append(data) # stack group so that features are the 3rd dimension loaded = dstack(loaded) return loaded |
We can use this function to load all input signal data for a given group, such as train or test.
The load_dataset_group() function below loads all input signal data and the output data for a single group using the consistent naming conventions between the directories.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 |
# load a dataset group, such as train or test def load_dataset_group(group, prefix=''): filepath = prefix + group + '/Inertial Signals/' # load all 9 files as a single array filenames = list() # total acceleration filenames += ['total_acc_x_'+group+'.txt', 'total_acc_y_'+group+'.txt', 'total_acc_z_'+group+'.txt'] # body acceleration filenames += ['body_acc_x_'+group+'.txt', 'body_acc_y_'+group+'.txt', 'body_acc_z_'+group+'.txt'] # body gyroscope filenames += ['body_gyro_x_'+group+'.txt', 'body_gyro_y_'+group+'.txt', 'body_gyro_z_'+group+'.txt'] # load input data X = load_group(filenames, filepath) # load class output y = load_file(prefix + group + '/y_'+group+'.txt') return X, y |
Finally, we can load each of the train and test datasets.
The output data is defined as an integer for the class number. We must one hot encode these class integers so that the data is suitable for fitting a neural network multi-class classification model. We can do this by calling the to_categorical() Keras function.
The load_dataset() function below implements this behavior and returns the train and test X and y elements ready for fitting and evaluating the defined models.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 |
# load the dataset, returns train and test X and y elements def load_dataset(prefix=''): # load all train trainX, trainy = load_dataset_group('train', prefix + 'HARDataset/') print(trainX.shape, trainy.shape) # load all test testX, testy = load_dataset_group('test', prefix + 'HARDataset/') print(testX.shape, testy.shape) # zero-offset class values trainy = trainy - 1 testy = testy - 1 # one hot encode y trainy = to_categorical(trainy) testy = to_categorical(testy) print(trainX.shape, trainy.shape, testX.shape, testy.shape) return trainX, trainy, testX, testy |
Fit and Evaluate Model
Now that we have the data loaded into memory ready for modeling, we can define, fit, and evaluate an LSTM model.
We can define a function named evaluate_model() that takes the train and test dataset, fits a model on the training dataset, evaluates it on the test dataset, and returns an estimate of the model’s performance.
First, we must define the LSTM model using the Keras deep learning library. The model requires a three-dimensional input with [samples, time steps, features].
This is exactly how we have loaded the data, where one sample is one window of the time series data, each window has 128 time steps, and a time step has nine variables or features.
The output for the model will be a six-element vector containing the probability of a given window belonging to each of the six activity types.
Thees input and output dimensions are required when fitting the model, and we can extract them from the provided training dataset.
|
1 |
n_timesteps, n_features, n_outputs = trainX.shape[1], trainX.shape[2], trainy.shape[1] |
The model is defined as a Sequential Keras model, for simplicity.
We will define the model as having a single LSTM hidden layer. This is followed by a dropout layer intended to reduce overfitting of the model to the training data. Finally, a dense fully connected layer is used to interpret the features extracted by the LSTM hidden layer, before a final output layer is used to make predictions.
The efficient Adam version of stochastic gradient descent will be used to optimize the network, and the categorical cross entropy loss function will be used given that we are learning a multi-class classification problem.
The definition of the model is listed below.
|
1 2 3 4 5 6 |
model = Sequential() model.add(LSTM(100, input_shape=(n_timesteps,n_features))) model.add(Dropout(0.5)) model.add(Dense(100, activation='relu')) model.add(Dense(n_outputs, activation='softmax')) model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy']) |
The model is fit for a fixed number of epochs, in this case 15, and a batch size of 64 samples will be used, where 64 windows of data will be exposed to the model before the weights of the model are updated.
Once the model is fit, it is evaluated on the test dataset and the accuracy of the fit model on the test dataset is returned.
Note, it is common to not shuffle sequence data when fitting an LSTM. Here we do shuffle the windows of input data during training (the default). In this problem, we are interested in harnessing the LSTMs ability to learn and extract features across the time steps in a window, not across windows.
The complete evaluate_model() function is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 |
# fit and evaluate a model def evaluate_model(trainX, trainy, testX, testy): verbose, epochs, batch_size = 0, 15, 64 n_timesteps, n_features, n_outputs = trainX.shape[1], trainX.shape[2], trainy.shape[1] model = Sequential() model.add(LSTM(100, input_shape=(n_timesteps,n_features))) model.add(Dropout(0.5)) model.add(Dense(100, activation='relu')) model.add(Dense(n_outputs, activation='softmax')) model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy']) # fit network model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose) # evaluate model _, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0) return accuracy |
There is nothing special about the network structure or chosen hyperparameters, they are just a starting point for this problem.
Summarize Results
We cannot judge the skill of the model from a single evaluation.
The reason for this is that neural networks are stochastic, meaning that a different specific model will result when training the same model configuration on the same data.
This is a feature of the network in that it gives the model its adaptive ability, but requires a slightly more complicated evaluation of the model.
We will repeat the evaluation of the model multiple times, then summarize the performance of the model across each of those runs. For example, we can call evaluate_model() a total of 10 times. This will result in a population of model evaluation scores that must be summarized.
|
1 2 3 4 5 6 7 |
# repeat experiment scores = list() for r in range(repeats): score = evaluate_model(trainX, trainy, testX, testy) score = score * 100.0 print('>#%d: %.3f' % (r+1, score)) scores.append(score) |
We can summarize the sample of scores by calculating and reporting the mean and standard deviation of the performance. The mean gives the average accuracy of the model on the dataset, whereas the standard deviation gives the average variance of the accuracy from the mean.
The function summarize_results() below summarizes the results of a run.
|
1 2 3 4 5 |
# summarize scores def summarize_results(scores): print(scores) m, s = mean(scores), std(scores) print('Accuracy: %.3f%% (+/-%.3f)' % (m, s)) |
We can bundle up the repeated evaluation, gathering of results, and summarization of results into a main function for the experiment, called run_experiment(), listed below.
By default, the model is evaluated 10 times before the performance of the model is reported.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 |
# run an experiment def run_experiment(repeats=10): # load data trainX, trainy, testX, testy = load_dataset() # repeat experiment scores = list() for r in range(repeats): score = evaluate_model(trainX, trainy, testX, testy) score = score * 100.0 print('>#%d: %.3f' % (r+1, score)) scores.append(score) # summarize results summarize_results(scores) |
Complete Example
Now that we have all of the pieces, we can tie them together into a worked example.
The complete code listing is provided below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 |
# lstm model from numpy import mean from numpy import std from numpy import dstack from pandas import read_csv from keras.models import Sequential from keras.layers import Dense from keras.layers import Flatten from keras.layers import Dropout from keras.layers import LSTM from keras.utils import to_categorical from matplotlib import pyplot # load a single file as a numpy array def load_file(filepath): dataframe = read_csv(filepath, header=None, delim_whitespace=True) return dataframe.values # load a list of files and return as a 3d numpy array def load_group(filenames, prefix=''): loaded = list() for name in filenames: data = load_file(prefix + name) loaded.append(data) # stack group so that features are the 3rd dimension loaded = dstack(loaded) return loaded # load a dataset group, such as train or test def load_dataset_group(group, prefix=''): filepath = prefix + group + '/Inertial Signals/' # load all 9 files as a single array filenames = list() # total acceleration filenames += ['total_acc_x_'+group+'.txt', 'total_acc_y_'+group+'.txt', 'total_acc_z_'+group+'.txt'] # body acceleration filenames += ['body_acc_x_'+group+'.txt', 'body_acc_y_'+group+'.txt', 'body_acc_z_'+group+'.txt'] # body gyroscope filenames += ['body_gyro_x_'+group+'.txt', 'body_gyro_y_'+group+'.txt', 'body_gyro_z_'+group+'.txt'] # load input data X = load_group(filenames, filepath) # load class output y = load_file(prefix + group + '/y_'+group+'.txt') return X, y # load the dataset, returns train and test X and y elements def load_dataset(prefix=''): # load all train trainX, trainy = load_dataset_group('train', prefix + 'HARDataset/') print(trainX.shape, trainy.shape) # load all test testX, testy = load_dataset_group('test', prefix + 'HARDataset/') print(testX.shape, testy.shape) # zero-offset class values trainy = trainy - 1 testy = testy - 1 # one hot encode y trainy = to_categorical(trainy) testy = to_categorical(testy) print(trainX.shape, trainy.shape, testX.shape, testy.shape) return trainX, trainy, testX, testy # fit and evaluate a model def evaluate_model(trainX, trainy, testX, testy): verbose, epochs, batch_size = 0, 15, 64 n_timesteps, n_features, n_outputs = trainX.shape[1], trainX.shape[2], trainy.shape[1] model = Sequential() model.add(LSTM(100, input_shape=(n_timesteps,n_features))) model.add(Dropout(0.5)) model.add(Dense(100, activation='relu')) model.add(Dense(n_outputs, activation='softmax')) model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy']) # fit network model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose) # evaluate model _, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0) return accuracy # summarize scores def summarize_results(scores): print(scores) m, s = mean(scores), std(scores) print('Accuracy: %.3f%% (+/-%.3f)' % (m, s)) # run an experiment def run_experiment(repeats=10): # load data trainX, trainy, testX, testy = load_dataset() # repeat experiment scores = list() for r in range(repeats): score = evaluate_model(trainX, trainy, testX, testy) score = score * 100.0 print('>#%d: %.3f' % (r+1, score)) scores.append(score) # summarize results summarize_results(scores) # run the experiment run_experiment() |
Running the example first prints the shape of the loaded dataset, then the shape of the train and test sets and the input and output elements. This confirms the number of samples, time steps, and variables, as well as the number of classes.
Next, models are created and evaluated and a debug message is printed for each.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Finally, the sample of scores is printed, followed by the mean and standard deviation. We can see that the model performed well, achieving a classification accuracy of about 89.7% trained on the raw dataset, with a standard deviation of about 1.3.
This is a good result, considering that the original paper published a result of 89%, trained on the dataset with heavy domain-specific feature engineering, not the raw dataset.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 |
(7352, 128, 9) (7352, 1) (2947, 128, 9) (2947, 1) (7352, 128, 9) (7352, 6) (2947, 128, 9) (2947, 6) >#1: 90.058 >#2: 85.918 >#3: 90.974 >#4: 89.515 >#5: 90.159 >#6: 91.110 >#7: 89.718 >#8: 90.295 >#9: 89.447 >#10: 90.024 [90.05768578215134, 85.91788259246692, 90.97387173396675, 89.51476077366813, 90.15948422124194, 91.10960298608755, 89.71835765184933, 90.29521547336275, 89.44689514760775, 90.02375296912113] Accuracy: 89.722% (+/-1.371) |
Now that we have seen how to develop an LSTM model for time series classification, let’s look at how we can develop a more sophisticated CNN LSTM model.
Develop a CNN-LSTM Network Model
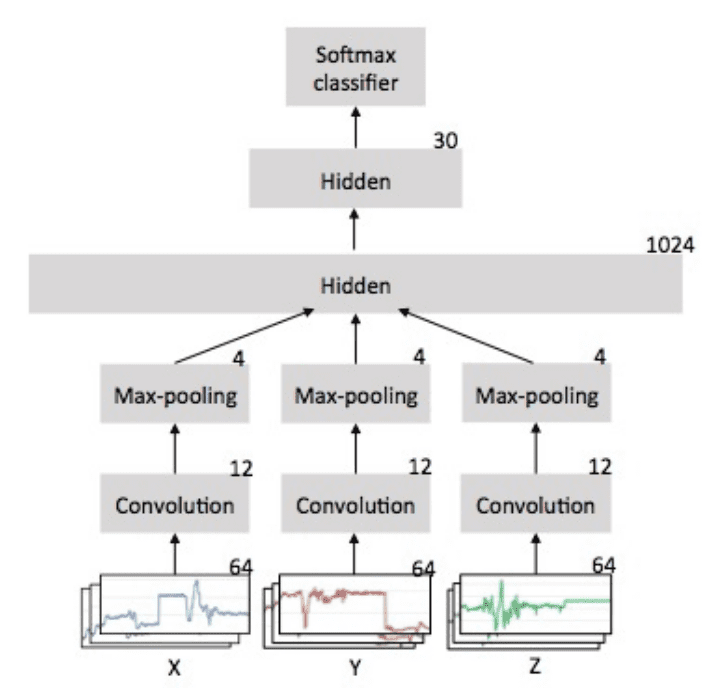
The CNN LSTM architecture involves using Convolutional Neural Network (CNN) layers for feature extraction on input data combined with LSTMs to support sequence prediction.
CNN LSTMs were developed for visual time series prediction problems and the application of generating textual descriptions from sequences of images (e.g. videos). Specifically, the problems of:
- Activity Recognition: Generating a textual description of an activity demonstrated in a sequence of images.
- Image Description: Generating a textual description of a single image.
- Video Description: Generating a textual description of a sequence of images.
You can learn more about the CNN LSTM architecture in the post:
To learn more about the consequences of combining these models, see the paper:
The CNN LSTM model will read subsequences of the main sequence in as blocks, extract features from each block, then allow the LSTM to interpret the features extracted from each block.
One approach to implementing this model is to split each window of 128 time steps into subsequences for the CNN model to process. For example, the 128 time steps in each window can be split into four subsequences of 32 time steps.
|
1 2 3 4 |
# reshape data into time steps of sub-sequences n_steps, n_length = 4, 32 trainX = trainX.reshape((trainX.shape[0], n_steps, n_length, n_features)) testX = testX.reshape((testX.shape[0], n_steps, n_length, n_features)) |
We can then define a CNN model that expects to read in sequences with a length of 32 time steps and nine features.
The entire CNN model can be wrapped in a TimeDistributed layer to allow the same CNN model to read in each of the four subsequences in the window. The extracted features are then flattened and provided to the LSTM model to read, extracting its own features before a final mapping to an activity is made.
|
1 2 3 4 5 6 7 8 9 10 11 |
# define model model = Sequential() model.add(TimeDistributed(Conv1D(filters=64, kernel_size=3, activation='relu'), input_shape=(None,n_length,n_features))) model.add(TimeDistributed(Conv1D(filters=64, kernel_size=3, activation='relu'))) model.add(TimeDistributed(Dropout(0.5))) model.add(TimeDistributed(MaxPooling1D(pool_size=2))) model.add(TimeDistributed(Flatten())) model.add(LSTM(100)) model.add(Dropout(0.5)) model.add(Dense(100, activation='relu')) model.add(Dense(n_outputs, activation='softmax')) |
It is common to use two consecutive CNN layers followed by dropout and a max pooling layer, and that is the simple structure used in the CNN LSTM model here.
The updated evaluate_model() is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 |
# fit and evaluate a model def evaluate_model(trainX, trainy, testX, testy): # define model verbose, epochs, batch_size = 0, 25, 64 n_timesteps, n_features, n_outputs = trainX.shape[1], trainX.shape[2], trainy.shape[1] # reshape data into time steps of sub-sequences n_steps, n_length = 4, 32 trainX = trainX.reshape((trainX.shape[0], n_steps, n_length, n_features)) testX = testX.reshape((testX.shape[0], n_steps, n_length, n_features)) # define model model = Sequential() model.add(TimeDistributed(Conv1D(filters=64, kernel_size=3, activation='relu'), input_shape=(None,n_length,n_features))) model.add(TimeDistributed(Conv1D(filters=64, kernel_size=3, activation='relu'))) model.add(TimeDistributed(Dropout(0.5))) model.add(TimeDistributed(MaxPooling1D(pool_size=2))) model.add(TimeDistributed(Flatten())) model.add(LSTM(100)) model.add(Dropout(0.5)) model.add(Dense(100, activation='relu')) model.add(Dense(n_outputs, activation='softmax')) model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy']) # fit network model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose) # evaluate model _, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0) return accuracy |
We can evaluate this model as we did the straight LSTM model in the previous section.
The complete code listing is provided below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 |
# cnn lstm model from numpy import mean from numpy import std from numpy import dstack from pandas import read_csv from keras.models import Sequential from keras.layers import Dense from keras.layers import Flatten from keras.layers import Dropout from keras.layers import LSTM from keras.layers import TimeDistributed from keras.layers.convolutional import Conv1D from keras.layers.convolutional import MaxPooling1D from keras.utils import to_categorical from matplotlib import pyplot # load a single file as a numpy array def load_file(filepath): dataframe = read_csv(filepath, header=None, delim_whitespace=True) return dataframe.values # load a list of files and return as a 3d numpy array def load_group(filenames, prefix=''): loaded = list() for name in filenames: data = load_file(prefix + name) loaded.append(data) # stack group so that features are the 3rd dimension loaded = dstack(loaded) return loaded # load a dataset group, such as train or test def load_dataset_group(group, prefix=''): filepath = prefix + group + '/Inertial Signals/' # load all 9 files as a single array filenames = list() # total acceleration filenames += ['total_acc_x_'+group+'.txt', 'total_acc_y_'+group+'.txt', 'total_acc_z_'+group+'.txt'] # body acceleration filenames += ['body_acc_x_'+group+'.txt', 'body_acc_y_'+group+'.txt', 'body_acc_z_'+group+'.txt'] # body gyroscope filenames += ['body_gyro_x_'+group+'.txt', 'body_gyro_y_'+group+'.txt', 'body_gyro_z_'+group+'.txt'] # load input data X = load_group(filenames, filepath) # load class output y = load_file(prefix + group + '/y_'+group+'.txt') return X, y # load the dataset, returns train and test X and y elements def load_dataset(prefix=''): # load all train trainX, trainy = load_dataset_group('train', prefix + 'HARDataset/') print(trainX.shape, trainy.shape) # load all test testX, testy = load_dataset_group('test', prefix + 'HARDataset/') print(testX.shape, testy.shape) # zero-offset class values trainy = trainy - 1 testy = testy - 1 # one hot encode y trainy = to_categorical(trainy) testy = to_categorical(testy) print(trainX.shape, trainy.shape, testX.shape, testy.shape) return trainX, trainy, testX, testy # fit and evaluate a model def evaluate_model(trainX, trainy, testX, testy): # define model verbose, epochs, batch_size = 0, 25, 64 n_timesteps, n_features, n_outputs = trainX.shape[1], trainX.shape[2], trainy.shape[1] # reshape data into time steps of sub-sequences n_steps, n_length = 4, 32 trainX = trainX.reshape((trainX.shape[0], n_steps, n_length, n_features)) testX = testX.reshape((testX.shape[0], n_steps, n_length, n_features)) # define model model = Sequential() model.add(TimeDistributed(Conv1D(filters=64, kernel_size=3, activation='relu'), input_shape=(None,n_length,n_features))) model.add(TimeDistributed(Conv1D(filters=64, kernel_size=3, activation='relu'))) model.add(TimeDistributed(Dropout(0.5))) model.add(TimeDistributed(MaxPooling1D(pool_size=2))) model.add(TimeDistributed(Flatten())) model.add(LSTM(100)) model.add(Dropout(0.5)) model.add(Dense(100, activation='relu')) model.add(Dense(n_outputs, activation='softmax')) model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy']) # fit network model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose) # evaluate model _, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0) return accuracy # summarize scores def summarize_results(scores): print(scores) m, s = mean(scores), std(scores) print('Accuracy: %.3f%% (+/-%.3f)' % (m, s)) # run an experiment def run_experiment(repeats=10): # load data trainX, trainy, testX, testy = load_dataset() # repeat experiment scores = list() for r in range(repeats): score = evaluate_model(trainX, trainy, testX, testy) score = score * 100.0 print('>#%d: %.3f' % (r+1, score)) scores.append(score) # summarize results summarize_results(scores) # run the experiment run_experiment() |
Running the example summarizes the model performance for each of the 10 runs before a final summary of the models performance on the test set is reported.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
We can see that the model achieved a performance of about 90.6% with a standard deviation of about 1%.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 |
>#1: 91.517 >#2: 91.042 >#3: 90.804 >#4: 92.263 >#5: 89.684 >#6: 88.666 >#7: 91.381 >#8: 90.804 >#9: 89.379 >#10: 91.347 [91.51679674244994, 91.04173736002714, 90.80420766881574, 92.26331862911435, 89.68442483881914, 88.66644044791313, 91.38106549032915, 90.80420766881574, 89.37902952154734, 91.34713267729894] Accuracy: 90.689% (+/-1.051) |
Develop a ConvLSTM Network Model
A further extension of the CNN LSTM idea is to perform the convolutions of the CNN (e.g. how the CNN reads the input sequence data) as part of the LSTM.
This combination is called a Convolutional LSTM, or ConvLSTM for short, and like the CNN LSTM is also used for spatio-temporal data.
Unlike an LSTM that reads the data in directly in order to calculate internal state and state transitions, and unlike the CNN LSTM that is interpreting the output from CNN models, the ConvLSTM is using convolutions directly as part of reading input into the LSTM units themselves.
For more information for how the equations for the ConvLSTM are calculated within the LSTM unit, see the paper:
The Keras library provides the ConvLSTM2D class that supports the ConvLSTM model for 2D data. It can be configured for 1D multivariate time series classification.
The ConvLSTM2D class, by default, expects input data to have the shape:
|
1 |
(samples, time, rows, cols, channels) |
Where each time step of data is defined as an image of (rows * columns) data points.
In the previous section, we divided a given window of data (128 time steps) into four subsequences of 32 time steps. We can use this same subsequence approach in defining the ConvLSTM2D input where the number of time steps is the number of subsequences in the window, the number of rows is 1 as we are working with one-dimensional data, and the number of columns represents the number of time steps in the subsequence, in this case 32.
For this chosen framing of the problem, the input for the ConvLSTM2D would therefore be:
- Samples: n, for the number of windows in the dataset.
- Time: 4, for the four subsequences that we split a window of 128 time steps into.
- Rows: 1, for the one-dimensional shape of each subsequence.
- Columns: 32, for the 32 time steps in an input subsequence.
- Channels: 9, for the nine input variables.
We can now prepare the data for the ConvLSTM2D model.
|
1 2 3 4 5 |
n_timesteps, n_features, n_outputs = trainX.shape[1], trainX.shape[2], trainy.shape[1] # reshape into subsequences (samples, time steps, rows, cols, channels) n_steps, n_length = 4, 32 trainX = trainX.reshape((trainX.shape[0], n_steps, 1, n_length, n_features)) testX = testX.reshape((testX.shape[0], n_steps, 1, n_length, n_features)) |
The ConvLSTM2D class requires configuration both in terms of the CNN and the LSTM. This includes specifying the number of filters (e.g. 64), the two-dimensional kernel size, in this case (1 row and 3 columns of the subsequence time steps), and the activation function, in this case rectified linear (ReLU).
As with a CNN or LSTM model, the output must be flattened into one long vector before it can be interpreted by a dense layer.
|
1 2 3 4 5 6 7 |
# define model model = Sequential() model.add(ConvLSTM2D(filters=64, kernel_size=(1,3), activation='relu', input_shape=(n_steps, 1, n_length, n_features))) model.add(Dropout(0.5)) model.add(Flatten()) model.add(Dense(100, activation='relu')) model.add(Dense(n_outputs, activation='softmax')) |
We can then evaluate the model as we did the LSTM and CNN LSTM models before it.
The complete example is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 |
# convlstm model from numpy import mean from numpy import std from numpy import dstack from pandas import read_csv from keras.models import Sequential from keras.layers import Dense from keras.layers import Flatten from keras.layers import Dropout from keras.layers import LSTM from keras.layers import TimeDistributed from keras.layers import ConvLSTM2D from keras.utils import to_categorical from matplotlib import pyplot # load a single file as a numpy array def load_file(filepath): dataframe = read_csv(filepath, header=None, delim_whitespace=True) return dataframe.values # load a list of files and return as a 3d numpy array def load_group(filenames, prefix=''): loaded = list() for name in filenames: data = load_file(prefix + name) loaded.append(data) # stack group so that features are the 3rd dimension loaded = dstack(loaded) return loaded # load a dataset group, such as train or test def load_dataset_group(group, prefix=''): filepath = prefix + group + '/Inertial Signals/' # load all 9 files as a single array filenames = list() # total acceleration filenames += ['total_acc_x_'+group+'.txt', 'total_acc_y_'+group+'.txt', 'total_acc_z_'+group+'.txt'] # body acceleration filenames += ['body_acc_x_'+group+'.txt', 'body_acc_y_'+group+'.txt', 'body_acc_z_'+group+'.txt'] # body gyroscope filenames += ['body_gyro_x_'+group+'.txt', 'body_gyro_y_'+group+'.txt', 'body_gyro_z_'+group+'.txt'] # load input data X = load_group(filenames, filepath) # load class output y = load_file(prefix + group + '/y_'+group+'.txt') return X, y # load the dataset, returns train and test X and y elements def load_dataset(prefix=''): # load all train trainX, trainy = load_dataset_group('train', prefix + 'HARDataset/') print(trainX.shape, trainy.shape) # load all test testX, testy = load_dataset_group('test', prefix + 'HARDataset/') print(testX.shape, testy.shape) # zero-offset class values trainy = trainy - 1 testy = testy - 1 # one hot encode y trainy = to_categorical(trainy) testy = to_categorical(testy) print(trainX.shape, trainy.shape, testX.shape, testy.shape) return trainX, trainy, testX, testy # fit and evaluate a model def evaluate_model(trainX, trainy, testX, testy): # define model verbose, epochs, batch_size = 0, 25, 64 n_timesteps, n_features, n_outputs = trainX.shape[1], trainX.shape[2], trainy.shape[1] # reshape into subsequences (samples, time steps, rows, cols, channels) n_steps, n_length = 4, 32 trainX = trainX.reshape((trainX.shape[0], n_steps, 1, n_length, n_features)) testX = testX.reshape((testX.shape[0], n_steps, 1, n_length, n_features)) # define model model = Sequential() model.add(ConvLSTM2D(filters=64, kernel_size=(1,3), activation='relu', input_shape=(n_steps, 1, n_length, n_features))) model.add(Dropout(0.5)) model.add(Flatten()) model.add(Dense(100, activation='relu')) model.add(Dense(n_outputs, activation='softmax')) model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy']) # fit network model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose) # evaluate model _, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0) return accuracy # summarize scores def summarize_results(scores): print(scores) m, s = mean(scores), std(scores) print('Accuracy: %.3f%% (+/-%.3f)' % (m, s)) # run an experiment def run_experiment(repeats=10): # load data trainX, trainy, testX, testy = load_dataset() # repeat experiment scores = list() for r in range(repeats): score = evaluate_model(trainX, trainy, testX, testy) score = score * 100.0 print('>#%d: %.3f' % (r+1, score)) scores.append(score) # summarize results summarize_results(scores) # run the experiment run_experiment() |
As with the prior experiments, running the model prints the performance of the model each time it is fit and evaluated. A summary of the final model performance is presented at the end of the run.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
We can see that the model does consistently perform well on the problem achieving an accuracy of about 90%, perhaps with fewer resources than the larger CNN LSTM model.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 |
>#1: 90.092 >#2: 91.619 >#3: 92.128 >#4: 90.533 >#5: 89.243 >#6: 90.940 >#7: 92.026 >#8: 91.008 >#9: 90.499 >#10: 89.922 [90.09161859518154, 91.61859518154056, 92.12758737699356, 90.53274516457415, 89.24329826942655, 90.93993892093654, 92.02578893790296, 91.00780454699695, 90.49881235154395, 89.92195453003053] Accuracy: 90.801% (+/-0.886) |
Extensions
This section lists some ideas for extending the tutorial that you may wish to explore.
- Data Preparation. Consider exploring whether simple data scaling schemes can further lift model performance, such as normalization, standardization, and power transforms.
- LSTM Variations. There are variations of the LSTM architecture that may achieve better performance on this problem, such as stacked LSTMs and Bidirectional LSTMs.
- Hyperparameter Tuning. Consider exploring tuning of model hyperparameters such as the number of units, training epochs, batch size, and more.
If you explore any of these extensions, I’d love to know.
Further Reading
This section provides more resources on the topic if you are looking to go deeper.
Papers
- A Public Domain Dataset for Human Activity Recognition Using Smartphones, 2013.
- Human Activity Recognition on Smartphones using a Multiclass Hardware-Friendly Support Vector Machine, 2012.
- Convolutional, Long Short-Term Memory, fully connected Deep Neural Networks, 2015.
- Convolutional LSTM Network: A Machine Learning Approach for Precipitation Nowcasting, 2015.
Articles
- Human Activity Recognition Using Smartphones Data Set, UCI Machine Learning Repository
- Activity recognition, Wikipedia
- Activity Recognition Experiment Using Smartphone Sensors, Video.
- LSTMs for Human Activity Recognition, GitHub Project.
Summary
In this tutorial, you discovered three recurrent neural network architectures for modeling an activity recognition time series classification problem.
Specifically, you learned:
- How to develop a Long Short-Term Memory Recurrent Neural Network for human activity recognition.
- How to develop a one-dimensional Convolutional Neural Network LSTM, or CNN LSTM, model.
- How to develop a one-dimensional Convolutional LSTM, or ConvLSTM, model for the same problem.
Do you have any questions?
Ask your questions in the comments below and I will do my best to answer.





Hi Jason,
So enjoy reading with your stuff, very helpful. As we can use CNN+LSTM to predict the spatial-temporal data, can we reverse the architecture as LSTM+CNN to do the same job? Any examples for LSTM + CNN?
Not that I have seen. What application did you have in mind exactly? Sequence to image?
Hi Jason,
I am working with a time series data which contains 1200 time series of different lengths and they all belong to one of the 4 classes. My goal is to classify them using LSTM, I can really understand how should i input these into the LSTM model. Can you help me out here please?
Perhaps use padding and a masking layer:
https://machinelearningmastery.com/data-preparation-variable-length-input-sequences-sequence-prediction/
Hi Jason,
Can you please explain the choice of parameters for the LSTM network?
Especially the LSTM layer and the dense layer?
What does the value 100 signify?
The model was configured via trial and error.
There is no analytical way to calculate how to configure a neural network model, more details here:
https://machinelearningmastery.com/faq/single-faq/how-many-layers-and-nodes-do-i-need-in-my-neural-network
Hi Jason, I’m just trying to understand correct me if I am wrong, does this value 100 in LSTM layer equal to 100 LSTM units in input layer? and each LSTM layer is fed with a sequence of length 128 (Time steps), right?
Yes, 100 is refers to the number of parallel units or nodes. It is unrelated to the number of timesteps.
Each node gets the full input sequence, not each layer.
Thank you for clarifying and I have one more question regarding dense layer. what is the input that dense layer is receiving from LSTM layer?( is it the time series itself or the last time step) and what happens if # of nodes in dense layer is not equal to # of nodes in LSTM(I mean is it possible to have more nodes in dense layer )?
The LSTM creates an internal representation / extracted features from the entire input sequence.
The term: „LSTM units” is very misleading. When it comes to the number of units we actually speak about the size of an internal state vector (either hidden, input or forget), so in the end it is just a mathematical thing. In my opinion it should not be called parallel because everything is done in one place at the same time (simple matrix-vector multiplication, where both are dimensionally-extended due to the number of “LSTM units”). BTW: Very good tutorial
Thanks!
Perhaps “nodes” would be more appropriate?
Definitely!
Note that I took the “unit” nomenclature from the original LSTM paper, and use it all the time across this site and my books:
https://www.bioinf.jku.at/publications/older/2604.pdf
Hi Jason. Could you please explain me the purpose of the first dense layer in the LSTM approach. Is it because of the dropout layer in front of it? Thanks.
Perhaps to interpet the output of the LSTM layers.
Thanks for such a comprehensive tutorial
I try it and it worked as expressed in the tutorial.
Now, I’m going to try it with my data that comes from a single axis accelerometer. It means I have only one feature so I don’t need a 3D array but a 2D.
You mentioned “The model (RNN LSTM) requires a three-dimensional input with [samples, time steps, features]. This is exactly how we have loaded the data.”
Then, it means that it won’t work with a 2D array? or I can consider a 3D array but the 3rd dimension has only one member?
and my second question is:
I need the model for a real-time classification, so I need to train once and then save the model and use it in my web application.
how can I save the model after training and use it?
Yes, even if the 3rd dimension has 1 parameter, it is still a 3D array.
You can call model.save() to save the model.
Hello Jason.
In my case I have 2 time series from EGG and I have to diseign a model that classify in two types de signal. I dont understand exactly how should i reshape the data.
The freq is 256 values per second so i could divide in windows like you did before. The problem is that i dont know how to put the 3rd dimension of caracteristics. From each window I have 7 caracteristics not from each moment (max, min, std, fft bandwidths, fft centroids, Arima 1, Arima 2)
Please, how could I do what you mean [samples, time steps, features] in my case??
Perhaps those 7 characteristics are 7 features.
If you have 256 samples per section, you can choose how many samples/seconds to use as time steps and perhaps change the resolution via averaging or removing samples.
Let’s see im going to make an easy example:
channel1 (values in one second)=2,5,6,8,54,2,8,4,7,8,…,5,7,8 (in total 256 values per second)
channel2 (values in one second)=2,5,6,8,54,2,8,4,7,8,…,5,7,8 (in total 256 values per second)
7 diferent features
[samples,timesteps,features]=[2, 256, 7]?
and another questions, for example the mean feature:
chanel 1:
feat_mean[0]=2
feat_mean[1]=(2+5)/2=3.5
feat_mean[2]=(2+5+6)/3=4.33
etc…
is it correct? what I understood is that I have to substract features for each each moment?
Yes, for one second of obs, that looks right.
You can use Pandas to up/downsample the series:
https://machinelearningmastery.com/resample-interpolate-time-series-data-python/
where does the “561 element vector of features” apply to?
That vector was the pre-processed data, prepared by the authors of the study that we do not use in this tutorial.
hello jason!
should we normalize each feature??
Perhaps try it and evaluate how it impacts model performance.
Does Batch Normalization layer serves the same purpose?
Not really, it is used to normalize a batch of activations between layers.
Hi Jason, thanks for the great article.
I notice in your load data section, you probably mean 2.56 seconds instead of 2.65, since 128(time step) * 1/50(record sampling rate) = 2.56.
Thanks, yes that is a typo. Fixed.
Hello Jason,
Why in Conv LSTM kernel_size=(1,3)? I don’t understand
For 1 row and 3 columns.
what what represents this in this example?
We split each sequence into sub-sequences. Perhaps re-read the section titled “Develop a ConvLSTM Network Model” and note the configuration that chooses these parameters.
Why we used 1 row? and why we used convLSTM2D? Can’t we model this problem like Conv1D?
A convlstm2d is not required, the tutorial demonstrates how to create different types of models for time series classification.
In dataset file, there aren’t label file (about y) Can i earn that files??
They are in a separate file with “y” in the filename.
i found it thank you!!
Glad to hear that.
thank you for answering me. I have one more question! you have 30 subjects in this experiment. so when you handle data, for example, in ‘body_acc_x_train’ all of the 30 subjects data is just merged?
Yes.
Also note, I did not run the experiment, I just analyzed the freely available data.
Oh thanks. Nowadays, i’m doing task about rare activity detection using sensor data, RNN&CRNN.
For example, i wanna detect scratch activity and non-scratch activity. But in my experiment, ratio of scratch and non-scratch window is not balanced (scratch is so rare..) then how to put my input ? Can you give me some advices?
Perhaps you can oversample the rare events? E.g. train more on the rare cases?
Hi Jason, i have question regarding to feeding feature data to LSTM. I have not used CNN, but if i use regular Auto encoder (sandwich like structure) instead of CNN for feature extraction, and if i define the time step of LSTM to be lets say 128,
1) should i extract feature from each time step and concatenate them to form a window before feeding to LSTM or
2) should i extract feature from window itself and feed the vector to LSTM
Thanks
The CNN must extract features from a sequence of obs (multiple time steps), not from a single observation (time step).
But LSTM will interpret features of each time step, not of a whole window right?
Hi Jason,
Thank you for the great material. My question is on data preprocessing. I have a squence of pressure data every 5 seconds with timestamp. How can I convert the 2D dataframe(Sample, feature) into 3D (Sample, Timestep, feature).
Good question, you can get started here:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Hi Jason,
In CNN LSTM model part, why we need to split the 128 time steps into 4 subsequences of 32 time steps?
Can’t we do this model with 128 time step directly?
Thank you
No, as this specific model expects sequences of sequences as input.
Hi there,
I have a question about your first model.
You set the batch_size for 64.
When I run your model (with verbose = 1), I got this :
Epoch 1/15
7352/7352 [==============================] – 12s 2ms/step – loss: 1.2669 – acc: 0.4528
It is mean 7352 * 64 ?
I ask this, because I want to overwrite your example which woulduse fit_generator, and I didn’t get the same results.
Here is my code:
………………
I don’t recall sorry, perhaps check the data file to confirm the number of instances?
any one cannot load the data? help, I think I follow the guide, unzip and rename in the working directory
solved, my mistake
Glad to hear it!
Sorry hear that, what is the problem exactly?
Hi Jason,
thank you for this tutorial. I am little be confused about the load the data set. Why we read_csv since no CVS files in the dataset. Sorry for this question because I am new to this subject. Also, I applied the code for (load_group), (load_dataset_group) and (load_dataset) ? Can you tell me if there is something need to add ?
In this tutorial we are loading datasets from CSV files, multiple files.
Sir, LSTM(100) means 100 LSTM cells with each cell having forget,input and output gate and each LSTM cell sends the output to other LSTM cells also and dinally every cell will give a 100 D vector as output..m i right?
100 cells or units means that each unit gets the output and creates an activation for the next layer.
The units in one layer do not communicate with each other.
Sorry Sir I didn’t get this.
1: Does these LSTM 100 cells communicate with each other?
2:If say we have 7352 samples with 128 timesteps and 9 features and my batch size is 64 then can i say that at time= 1 we will input first timestep of 64 samples to all the 100 LSTM cells then at time=2 we will input second time step of 64 samples and so on till we input 128th time step of 64 samples at time= 128 and then do BPTT and each lstm preserves its state from time =1 to time=128 ?
No, cells in one layer do not communicate with each other.
No, each sample is processed one at a time, at time step 1, all 100 cells would get the first time step of data with all 9 features.
BPTT refers to the end of the batch when model weights are updated. One way to think about it is to unroll each cell back through time into a deep network, more here:
https://machinelearningmastery.com/rnn-unrolling/
“No, each sample is processed one at a time, at time step 1, all 100 cells would get the first time step of data with all 9 features”
sir batch size means how many samples to show to network before weight updates.
If i have 64 as batch size….then is it that at time step 1, all 100 cells would get the first time step of each 64 data points with all 9 features?then at next time step =2, all 10 cells would get the second time step of each 64 data points with all 9 features and so on.
or is it like at time step 1, all cells get the first time step of one data point with all 9 features then at time step =2 all cells get the second time step of that data point and when all the 128 time steps of that point is fed to the network we compute the loss and do this same for remaining 63 points and then updates weights?
I am getting confused how batch size is working here..what i am visualizing wrong here ?
If the batch size is 64, then 64 samples are shown to the network before weights are updated and state is reset.
In a batch, samples are processed one at a time, e.g. all time steps of sample 1, then all time steps of samples 2, etc.
I strongly recommend reading this:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Also here we are using dropout after lstm 100 means only 50 values will be passed to a dense 100 layer right sir?
On average, yes 100 activations.
hello
I am beginner in area.. thanks for wonderful tutorial..
I wanted to workout your LSTM and CNN RNN example, I have downloaded HARDataset but
I have simple question here how to give input CSV file at the beginning ?
Do I have to generate that file? if so how to do that?
# load a single file as a numpy array
def load_file(filepath):
dataframe = read_csv(filepath, header=None, delim_whitespace=True)
return dataframe.values
Please help me
thanks in advance
You can follow the tutorial to learn how to load the CSV file.
Did you get your answer?
Sir i printed the model summary for CNN_LSTM output dimensions for timedistributed came as 4dimensions like (None,None,30,64)…it is 4D because we have partioned the windows into 4 subwindows of 32 size?
what is None,None representing here….30,64 I got that it is the output after 1st convolution
The “None” sized dimension means that the provided dataset will define the other dimensions.
Sir https://machinelearningmastery.com/cnn-long-short-term-memory-networks/ u have mentioned that…
“In both of these cases, conceptually there is a single CNN model and a sequence of LSTM models, one for each time step. We want to apply the CNN model to each input image and pass on the output of each input image to the LSTM as a single time step.”
So here we are dividing the 128 time steps into 4 subblocks..we will give each block to the CNN at once and our CNN model will give the output features.
means we have converted 128 time sequences to 4 time sequences here whih we will feed to our LSTM model now.
so earlier we were feeding 128 time steps to the LSTM(in simple LSTM) and now we will feed 4…am i right?
Yes, but the “4 time steps” are distilled from many more real time steps via the CNN model.
and sir CNN will process each of the 32 windows parallelly or will it first process first 32 sized window and feed it to lstm then another 32 and so on
Each window of data is processed by the n filters in the CNN layer in parallel – if that is what you mean.
yes sir go it ..thanks a lot
Sir means all four 32 sized windows from one 128 time step window are processed by th CNN parallelly then at time step 1 what we will input into our lstm?
like in normal lstm with 128 timesteps we input 1st time step with 9 features then 2nd timestep and so on..
here since we have processed 4 time steps parallely what we will input to lstm?
The LSTM takes the CNN activations (feature maps) as input. You can see the output shapes of each layer via the model.summary() output.
Sir I did that but getting a little confused….this is te summary
None,None,30,64….1st convoution
None,None,28,64….2nd convolution
None,None,28,64…dropout
None,None,896…maxpool
None,100…lstm 100
None,100…dropout
None,100…dense 100
None,6…softmax
Sir,My doubt is we input 32 sized window from 128 sized original window to lstm…so does lstm here predicting the activity for each 32 sized window treating it as one time sequence?
Dear Mr. Brownlee,
I was also trying to get behind the idea of how the data flows in the CNN-LSTM model, sadly I didn’t fully understand it yet. I was reading through the comment section and found a few helpful questions and answers! Still though, I’m stuck at this very exact point between the Flatten layer and the LSTM layer. How exactly are the flattened (invariant) features from the max pooling layer getting fed into the LSTM layer?
I tried to dig a little deeper and made some extra research on the internet. There I found these three posts
https://stackoverflow.com/questions/43237124/what-is-the-role-of-flatten-in-keras
https://medium.com/smileinnovation/how-to-work-with-time-distributed-data-in-a-neural-network-b8b39aa4ce00
https://datascience.stackexchange.com/questions/24909/the-model-of-lstm-with-more-than-one-unit
These three posts helped me to visualize the data flow in the network. Have I understood everything correctly?… So Keras will distribute the input in layers step by step to the LSTM cells. This means that it’ll first present the invariant flattened layers of the first subsequence to the LSTMs, then the features from the second subsequence…
To clearly show what I mean I made this visualization that also includes a short numerical example of the forwardpass from a smaller subsequence:
https://filebin.net/0kywmitsq5u0qgy5
In the numerical example I take the 2 first subsequences and show how they change from input, then through first and second convolution (here I just picked some random numbers), next the max-pooling operation and the flatten operation (note that here I was trying to correctly max-pool the random numbers from the previous convolution layers. Also I was flattening the max-pooled features, how they should be flattened according to the second post-link).
Is this understandable and correct what I visualized here?
And another questions that came up. If I feed the flattened features to the 100 LSTM cells (that are all in one layer, meaning that they are not stacked or not connected with each other or something like that). What is the exact benefit of using more then just one LSTM cell in one layer?
In terms of understanding the input to the model, this will help:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
In terms of how an LSTM layer processes input, this will help:
https://machinelearningmastery.com/faq/single-faq/how-is-data-processed-by-an-lstm
In terms of the flatten layer in the CNN-LSTM, we need to flatten the feature maps from the CNNs. Perhaps review the input of summary() function on the model or plot the model with output shapes to see the transition:
https://machinelearningmastery.com/visualize-deep-learning-neural-network-model-keras/
For more on the time distributed layer:
https://machinelearningmastery.com/timedistributed-layer-for-long-short-term-memory-networks-in-python/
I hope that helps as a first step.
Hey there!
Thankyou for your quick response. I have to say that I’ve already read through those articles, but none of them really explains exactly that detail, how the flattened features from the CNN layer are getting fed into the LSTM layer… so as I was trying to explain earlier, I tried to do a little bit more research (this included your articles) and now I think I came up with the correct solution, which I tried to visualize here
https://filebin.net/0kywmitsq5u0qgy5
I just wanted to ask you, whether you might want to take a look on that and tell me, whether it is correct.
Also what again is the benefit of using multiple LSTM cells in parallel, over using just a single LSTM cell?
Thankyou in advance and best wishes,
Alex
Sorry, I don’t have the capacity to review code/documents/data. I get 100s of similar requests daily.
Parallel LSTMs units in a layer (like parallel nodes in an MLP) give more “capacity” to the model:
https://machinelearningmastery.com/how-to-control-neural-network-model-capacity-with-nodes-and-layers/
Ok I understand that ! Thanks a lot anyways for all your effort and your answers so far!
Do you have an idea about the multi-class classification with the algorithm ECOC algorithm.
Can we use it for unsupervised classification (clustering)
What is the ECOC algorithm?
Error-correcting output coding (ECOC) algorithm
Thanks for sharing.
Does it makes sense to use dropout and maxpooling after convlstm layer, like we did in cnn?
Hmmm, maybe.
Always test, and use a configuration that gives best performance.
Hi Jason,
thank you for this tutorial.
Why is it necessary to perform signal windowing before training the neural network?
Can we consider the full signal instead?
We must transform the series into a supervised learning problem:
https://machinelearningmastery.com/time-series-forecasting-supervised-learning/
Let’s assume that the original signal is an acceleration representing a person walking and that the aim is to establish wheter the person fell on the floor.
Let’s say the original signal is composed by 1000 samples belonging to class1. The signal is processed and divided into fixed windows of N data points: now I have sub-signals each one labelled with class1. Is that correct to consider different windows even if the peak of the falling is present only in one of the them?
Perhaps. There are many ways to frame a given prediction problem, perhaps experiment with a few approaches and see what works best for your specific dataset?
Is their any role of LSTM units in convlstm. parameters of convlstm layer are similar to CNN, but nothing related to LSTM can be seen. In LSTM model, 100 LSTM units are used. How can we see convlstm in this context?
Not sure I follow what you’re asking, can you elaborate?
Following are input layers used in three different models.
model.add(LSTM(100, input_shape=(n_timesteps,n_features)))
model.add(TimeDistributed(Conv1D(filters=64, kernel_size=3, activation=’relu’), input_shape=(None,n_length,n_features)))
model.add(ConvLSTM2D(filters=64, kernel_size=(1,3), activation=’relu’, input_shape=(n_steps, 1, n_length, n_features)))
ConvLSTM2D layer I believe should be a mixture of CNN and LSTM. All parameters in convlstm layers are parameters of cnn like no of filters, filter size, activation function etc. LSTM as standalone uses 100 LSTM cells. My question was how many LSTM cells will be used by convlstm model? I believe convlstm operates with only one lstm cell?
I believe each filter is a cell – the idea behind these two concepts is merged.
I am generating sine and cosine curves for classification. However, I am not sure whether I need to pre-process data or just load it to the model. My understanding is further confounded by the following state in your post “Further, each series of data has been partitioned into overlapping windows of 2.56 seconds of data, or 128 time step”
Yes, perhaps this post will make things clearer:
https://machinelearningmastery.com/time-series-forecasting-supervised-learning/
Jason thanks a lot for his wonderful tutorial.
I want to use your approach for my problem. But my data set is a little different than what you used here.
I have 75 time series. Each of them shows two classes. From time 0 until time t ( which is different for each time series) is data for class 1 and from time t until the end of time series is class 2. Then, I want for test time series predict the time at which the class has changed from 1 to 2, or at each time the class is 1 or 2. can you help me how I can use your approach for my problem?
Perhaps you can pad each time series to the same length and use a masking layer to ignore the padded values.
This might help:
https://machinelearningmastery.com/data-preparation-variable-length-input-sequences-sequence-prediction/
Thanks Jason, but the difference is that each time series has two classes. But, the time series you used are for one class each.
Do you mean that each sequence is classified twice, e.g. {A,B} and {C,D}.
If so, perhaps use two models or perhaps two output layers, one for each classification?
I mean each time series shows two classes, healthy and unhealthy, for one system. From time 0 until time t, it shows healthy state and from time t until the failure of the system, it shows unhealthy state. We have 75 time series like that with different lengths for both classess. Now We want to determine for a test system, the time that it switches from healthy to unhealthy state.
Thanks
Perhaps you can predict the class value for each input time step, if you have the data?
May you please explain more? Do you think I can use your approach ?
I was suggesting perhaps try modeling the problem as a one-to-one mapping so each input time step has a classification.
More on sequence prediction here:
https://machinelearningmastery.com/models-sequence-prediction-recurrent-neural-networks/
I applied your tutorial to data similar to Amy’s (I guessed) where I tried to predict disease events, and for training and validation I used the n days before a disease event, and as comparison n days from an individual without disease events, and each window classified as either 1 (sick) or 0 (healthy).
The model was performing okay with an AUC of >0.85, but I’m not sure how I would apply this in practice because the time windows for the validation data were designed with a priori knowledge.
In practice one would have to construct a new input vector every time step, and I don’t think the classification of those vectors would be as good. But I did not try that out yet.
What I didn’t understand is how would I apply the on-to-one mapping from your article here? You state that the one-to-one approach isn’t appropriate for RNNs since it doesn’t capture the dependencies between time points.
@Amy you could investigate on heart rate classification with neural networks, I think that is a somewhat similar problem.
Not sure I follow the question completely.
Generally, you must frame the problem around the way the model is intended to be used, then evaluate under those constraints. If you build/evaluate a model based on a framing that you cannot use in practice, then the evaluation is next to useless.
Using training info or domain knowledge in the framing is fine, as long as you expect it to generalize. Again, this too can be challenged with experiments.
If I’ve missed the point, perhaps you can elaborate here, or email me?
https://machinelearningmastery.com/contact/
May you please explain more?
Hello sir,
Great tutorial, how to normalize or standardize this data.
I show how here:
https://machinelearningmastery.com/machine-learning-data-transforms-for-time-series-forecasting/
Dear Mr. Brownlee,
I am a student from Germany and first of all: thank you for your great Blog! It is so much better than all the lectures I have been visited so far!
I had a question regarding the 3D array and I hope you could help me. Let’s assume we have the following case, which is similar to the one from your example:
We measure the velocity in 3 dimensions (x,y,z direction) with a frequency of 50 Hz over one minute. We measure the velocity of 5 people in total.
– Would the 3D array have the following form: (5*60*50; 1 ; 3)?
– What do you mean by time steps? I am referring to “[samples, time steps, features]”.
– Is the form of the 3D array related to the batch size of our LSTM model?
Thank you in advance. I would really appreciate your help as I am currently stuck…
Best regards,
Simon
Great question, I believe this will make samples/timesteps/features clearer:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Dear Mr. Brownlee,
thank you for your replay and reference to the FAQ page.
Still, I am a little bit confused with the example at the FAQ page. Maybe there is a mistake?
I mean the following example about the 604.800 seconds. You said the input shape is [604800, 60, 2] for 60 time steps but doesn’t it supposed to be like [10080, 60, 2]?
Could give me a hint how the RNN handles different time steps in the input?
Thank you in advance!
Best regards,
Simon
I think you’re right re the FAQ, fixed.
We typically vectorize the data and pad it so each input sequence has the same number of timesteps and features, more on padding here:
https://machinelearningmastery.com/data-preparation-variable-length-input-sequences-sequence-prediction/
what is the use of those 561 features?
What do you mean exactly?
if in those 128*9 data, 64*9 represent standing and other 64*9 represent walking then how do i label that 128*9 data?
You can model the problem as a sequence classification problem, and try different length sequence inputs to see what works best for your specific dataset.
Amazing!
But how do I train the network with additional new data? I’m working on a project to detect suspicious activity from surveillance videos.
I have no idea on how to prepare such dataset. Appreciate your help!
You can save the model, load it later and update/train it on the new data or a mix of old and new. Or throw away the model and fit it anew.
Perhaps test diffrent approaches and compare the results.
Hi Jason,
Do you have the link for trained model? I would like to quickly check that how well it works on my data.
Also, what is size of your model? I am looking for the models which are less in size so as to be able to deploy on edge?
Sorry, I don’t share trained models.
Perhaps try fitting the model yourself, it only takes a few minutes.
what is the purpose of ‘prefix’ when loading the file?
In case you have the data located elsewhere and need to specify that location.
Hi Jason,
If I export this model to an android app should I do any preprocessing on input data from mobile sensors?
I don’t know about android apps sorry.
Hi Jason,
the first LSTM example, you mentioned that the multiple times of evaluation due to stochastic reason. BUT, how did you get the best performance parameter weights?
We don’t get the best weights, we evaluate the average performance of a model.
We can reduce the variance of the predictions of a given model using ensembles:
https://machinelearningmastery.com/start-here/#better
Hi Jason, thanks for the great article. I am currently working on human activity recognition (Kinetics-600) and I wanted to connect LSTM with 3D ResNet head for action prediction. Can you please tell how can I use LSTM on 1024 vectors obtained from the last layer and feed it to RNN-LSTM for action prediction?
Thank you.
That is challenging for me to answer off the cuff without writing custom code/digging into your data – which I cannot do.
Perhaps this will give you ideas on how to adapt your data for an LSTM:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
model = Sequential()
model.add(LSTM(100, input_shape=(n_timesteps,n_features)))
model.add(Dropout(0.5))
model.add(Dense(100, activation=’relu’))
model.add(Dense(n_outputs, activation=’softmax’))
model.compile(loss=’categorical_crossentropy’, optimizer=’adam’, metrics=[‘accuracy’])
In this code i am getting an this traceback
Traceback (most recent call last):
File “C:\Users\Tanveer\AppData\Local\Programs\Python\Python37-32\HARUP.py”, line 54, in
model = Sequential()
NameError: name ‘Sequential’ is not defined
How can resolve this issue kindly guide me
You may have skipped some lines from the full example.
Dear Sir i didn’t skipped any code which is given above but this problem still occuring kindly give me some guideline or hint to solve this issue
I recommend copying the complete code example, rather than the partial code examples.
hi Jason. I am running the code from here https://medium.com/@curiousily/human-activity-recognition-using-lstms-on-android-tensorflow-for-hackers-part-vi-492da5adef64 on anaconda and notebook and pycharm with TensorFlow at 1.4 version (i had to downgrade due to the fact that the author is using placeholders which are incompatible with TensorFlow 2) and python at 3.6. The problem here is that I always get train loss: nan
can you please suggest some ideas here because I cannot find anything when I search for the problem (there are some articles but are not helpful)
Sorry, i am not familiar with that tutorial, perhaps contact the authors?
Hello Jason,
Amazing tutorial. I am doing something very similar to this. My problem is: “If I am given a set of 128 time steps with 9 features data, i.e. an ndarray of the shape (128,9), how can I use the model.predict() method to make a prediction for the 128 time steps data?” Currently when I do model.predict( ndarray of shape (128, 9)), I get the error that “expected lstm_1_input to have 3 dimensions, but got array with shape (128, 9)”. From my understanding, I will be provided a time steps data with its feature values, and I have to predict the class for it. How can this data be 3d since I have to predict only one sample?
Thank you
Yes, the shape for one sample would be [1, 128, 9].
For more on this, see:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
hey Jason,
Thanks for the useful tutorial 🙂
I don’t really get how the data is shaped into windows of size 128. I know here the data has already been shaped but I am asking if you have a tutorial showing how this shaping is done for a classification problem.
Thanks,
This shaping cannot be performed for a classification problem on tabular data, it only makes sense for a sequence prediction problem.
so how can I shape my data for a classification problem? is there any tutorial showing similar problem?
All you need is a table, rows of examples with columns of features.
Stored in a CSV file.
Does that help?
my question is how to make the sliding window?
I have the csv file , I have a target with two classes 1 and 0. I have readings per day.
I want to make a sliding window of 5 days. Does this work for my problem or not?
and if it work, when I shape the window do i include the target variable or I exclude it?
the sliding window is done using the shift() in the to_supervised function or using the reshape only is enough?
Thanks,
This tutorial will show you how:
https://machinelearningmastery.com/time-series-forecasting-supervised-learning/
This tutorial will give you code:
https://machinelearningmastery.com/convert-time-series-supervised-learning-problem-python/
Thanx alot Jason 🙂
You’re welcome.
Hi,
do you think your CNN+LSTM example would achieve good results for pose estimation ? what necessary changes would have to be done to accomplish this ? thanks in advance
Perhaps try it and see?
Hello,
my dataset has dimensions (170,200,9) and i want to assign one class label to one sample/window of 200 timesteps, not individual labels to each of the 200 time steps in one window.
How can i do this? so my target class output has dimensions (170,1)
Yes, this is called time series classification.
The above tutorial does this.
Hello,
Can you please tell me which files do I need to load in the code below for stacking?
Thanks.
Sorry, I don’t understand your question. Perhaps you can elaborate?
I meant that which files in the dataset should I load and stack so that the features are in 3 dimensions?
I mean for applying the Numpy’s dstack function in the code above which files from the dataset should I load?
Thanks in advance.
The example in the tutorial shows exactly how to call this function to stack the dataset.
No need to invent anything – just copy the full code example.
Hello,
Can you please tell me why are we subtracting 1 in the code below?
# zero-offset class values
trainy = trainy – 1
testy = testy – 1
Thank you.
As the comment suggests, to change the integers for the class label to have a zero offset. To start at 0 and not 1.
Hi Jason, Have a good day.
Thank you for this tutorial. I have one question. I have data to online learning students, I want to prepare data to CNN LSTM, the data form ( students, weeks, videos for every week, video features ). now, I want to make inputs data(samples,n_steps,n_length, features) where n_length is Variable length such as ( first week 8 videos, second week of 13 videos, so on,). these data form as inputs data to CNN then LSTM. as like CNN-LSTM. Please, Can you guide me, how to prepare this data? my especially question, how can make n_length is variable length?
This might help:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Hi Jason,
I used this tutorial to do my time-series-classification / prediction.
Basically, I only use 1st part of this tutorial, the LSTM.
The predicted result should be either 0, 1, or 2
I had 3 classes in Y, the integer values are 0, 1, 2.
So after a calling to_categorical(), that will be:
0 –> [1, 0, 0]
1 –> [0, 1, 0]
2 –> [0, 0, 1]
However, after I make this kind of function call:
yhat = model.predict(test_X, verbose=verbose)
I got these kind of values:
…
[0.95416844 0.00828072 0.03755087]
[0.9620431 0.00921117 0.02874582]
[0.9660313 0.01018358 0.02378514]
[0.9682437 0.01118453 0.02057172]
[0.96942824 0.01226482 0.01830701]
[0.9697388 0.01347514 0.0167861 ]
…
Can you elaborate that a little bit about how to convert my predicted values back to one-hot form or integer form?
Thanks a lot
Calculate the argmax() to get the index of the class label with the largest probability for each predicted sample.
Thank you so much. It solved my problem.
However, I got a second question.
For this kind of predicting results,
[0.95416844 0.00828072 0.03755087]
[0.9620431 0.00921117 0.02874582]
[0.9660313 0.01018358 0.02378514]
[0.9682437 0.01118453 0.02057172]
[0.96942824 0.01226482 0.01830701]
[0.9697388 0.01347514 0.0167861 ]
if I use the argmax(), I will have all of events classified to class 1 which is “0”
I was just wondering if there is anything wrong with my model?
model.add(LSTM(100, input_shape=(train_X.shape[1], train_X.shape[2])))
model.add(Dropout(0.5))
model.add(Dense(100, activation=’relu’))
model.add(Dense(n_outputs, activation=’softmax’))
model.compile(loss=’categorical_crossentropy’, optimizer=’adam’, metrics=[‘accuracy’])
Or anything I need to do in data pre-processing?
For example, this is my raw data piece for the feature I’m predicting on:
0
0
0
0
0
1
0
0
0
0
0
1
0
0
0
0
0
0
0
0
0
0
2
0
0
0
0
0
0
1
0
0
0
0
0
1
0
0
0
0
0
0
0
0
0
0
2
0
By the way, this seems like a more complicated problem of “Classification on imbalanced data” ?
Well done.
Yes, I recommend testing a suite of models, configs and pre-processing methods in order to discover what works best for your specific dataset.
Hey, can you share the code you used for Argmax () ? Thanks in advance 🙂
This will help:
https://machinelearningmastery.com/argmax-in-machine-learning/
HI Sir,
if we want to test the lstm model to new data, let’s say fro here(https://sensor.informatik.uni-mannheim.de/#dataset_realworld) which will be the procedure?
1. we will stop after the train and then we will go to evaluate to new data (model.evaluate(new_data), without some pre-processing?
2. or we must turn the data in a format that our lstm model understand
Thank you so much for giving me the opportunity to ask my question.
You must prepare all data the same way, including test data and new data when making a predictions.
thank you fo the answer,
when the model performs well on test data but not in new data what you think is the case?
it depends on the quality of the new data, or maybe some hyperparameters change will lead to better accuracy?
Perhaps the test dataset is too small or not representative of the problem.
thank you so much for your answers
You’re welcome.
as I do not find anywhere in the site I will ask here. what is the procedure for downsampling sensor signal?
if say we have a dataframe with same columns like
acticity, accx, accy, accz, gurx, gury, gurz
and the sampling array is 100hz. How can I downsample to 50 Hz?
I find pandas replace but I have not seen anywhere how is applied.
I believe pandas offers sampling methods that might help:
https://machinelearningmastery.com/resample-interpolate-time-series-data-python/
Hi,
I have a classification problem in which the transitions from one state to the next happen very abruptly. I can classify the sequences well with the above example but is there a way to accurately identify when the system changes from one state to the next?
Thanks
Good question.
I recommend looking into models for “regime changes” in time series.
Hello Sir,
Running your LSTM code yields me a shape of
(7352, 128, 1) (7352, 1)
(2947, 128, 1) (2947, 1)
(7352, 128, 1) (7352, 6) (2947, 128, 1) (2947, 6)
instead of
(7352, 128, 9) (7352, 1)
(2947, 128, 9) (2947, 1)
(7352, 128, 9) (7352, 6) (2947, 128, 9) (2947, 6)..
Can you please clarify the reason?
Perhaps confirm you followed the steps of the tutorial in order.
Also see this:
https://machinelearningmastery.com/faq/single-faq/why-does-the-code-in-the-tutorial-not-work-for-me
hi,
when the signal from the accelerometer or gyroscope data does not capture properly (i.e sharp peaks, edges, flat lines, etc), do you think that a auto-encoder lstm network will get read of this permutations?
It might smooth them out. Perhaps test?
hi Jason,
thank you for your answers in my previous posts.
can i use the suggestion from here https://machinelearningmastery.com/diagnose-overfitting-underfitting-lstm-models/
in order to check for overfitting or underfitting? and how i decide to the number of validation split during train?
Yes, but you may have to create the curve manually.
Depends on the problem and what you want to validate. E.g. perhaps try different size validation sets and see what is stable.
thank you Jason. What do you mean to create the curve manually. I usually use this code
def plot_learningCurve(history, epochs):
# Plot training & validation accuracy values
epoch_range = range(1, epochs+1)
plt.plot(epoch_range, history.history[‘accuracy’])
plt.plot(epoch_range, history.history[‘val_accuracy’])
plt.title(‘Model accuracy’)
plt.ylabel(‘Accuracy’)
plt.xlabel(‘Epoch’)
plt.legend([‘Train’, ‘Val’], loc=’upper left’)
plt.show()
# Plot training & validation loss values
plt.plot(epoch_range, history.history[‘loss’])
plt.plot(epoch_range, history.history[‘val_loss’])
plt.title(‘Model loss’)
plt.ylabel(‘Loss’)
plt.xlabel(‘Epoch’)
plt.legend([‘Train’, ‘Val’], loc=’upper left’)
plt.show()
But calculating validation accuracy via fit() might not be correct, instead we might want to estimate it using walk-forward validation.
ok thanks, i will try your suggestions
You’re welcome.
hi,
what will happen if someone omits this layer model.add(Dense(100, activation=’relu’))
i am performing different experiments with a dataset that i record (accelerometer, gyroscope signals at 100hz).
when i do not use the above layer the accuracy raises to 1.00 and all classes are property classified
when i use the layer the accuracy drops at 78% and some static activities like sitting can not be classified
what you think is the case here?
The dense layer interprets the output from the LSTM.
Try other configs and see if you can lift the skill of the model.
ok i see thanks. so it seems mandatory to have a dense layer before your final dense layer where you make predictions
No, some people use an global pooling layer.
oo I thought that global pooling layer is only for CNN.
Thanks i gonna search for this
hi ,
one more question, please
how do you decide the number of hidden units?
model.add(LSTM(100, input_shape=(n_timesteps,n_features)))
why 100 and not 200 or 50 etc? Are you aware of any paper that discusses this issue
Good question, I answer it here:
https://machinelearningmastery.com/faq/single-faq/how-many-layers-and-nodes-do-i-need-in-my-neural-network
I wish to form a model which could detect yawn of the driver. I wish to know how and by using what algorithm I can do that??
Good question, see this:
https://machinelearningmastery.com/faq/single-faq/what-algorithm-config-should-i-use
Great work Jason!
I am doing video classification for action detection using Keras (v.2.3.1). My model is CNN and LSTM. My dataset consists of videos of 3-7 seconds, each representing specific action. I use OpenCv to get frames from each video. But since video lengths are different, I get different number of frames for each video. However, I need to have the same number of frames for the LSTM layer. I searched a little bit and looks like padding along with a masking layer should do it, but I can’t figure out how to do this with Keras. Could you please provide some clues on how to achieve this?
Perhaps try zero padding the sequences of frames.
Is it possible to classify activities detected by bounding boxes ?
To elaborate on my above comment,
how to build a model to classify a multi-class video sequence? instead of the whole video as you show in the above tutor.
For example, let us imagine we have a video with 4 different clocks moving at varying speeds. I have folders as following:
Folder 1: 1000 multi-clocks in a Video
Folder 2: 1000 corresponding annotated JSON/XML files with the co-ordinates of the clocks with their respective labels (“Slow Speed”, “Normal Speed”, “Fast Speed”, “Varying Speed”).
How to build a model to classify such videos?
I don’t have examples of working with video, sorry.
Hi Jason, I found that you have
https://machinelearningmastery.com/how-to-develop-rnn-models-for-human-activity-recognition-time-series-classification/
Is there an expansion on that tutorial so that instead of recognition on the whole video, there are several recognitions on the same video multiple activities? i.e. Bounding with Slow Speed, Bounding box with Normal speed etc…
It will not be sufficient to do only “object recognition” since it needs the next frames in order to predict the clock speed.
Some methods I found online are:
1) 3D CNN
2) ConvNets + RNN
3) COnvent + MLP
4) Two steam convolutional networks
Do you cover any of these methods in one of your books perhaps with a video example?
Thank you for your prompt reply, and thank you for all your fantastic work.
I don’t have any examples of working with video.
In images? Yes, that is called “object recognition”:
https://machinelearningmastery.com/object-recognition-with-deep-learning/
Hello Dear,
I want to plot time series graph single activity..
For Example,
def plot_activity(activity, df):
data = df[df[‘activity’] == activity][[‘x-axis’, ‘y-axis’, ‘z-axis’]][:200]
axis = data.plot(subplots=True, figsize=(16, 12),
title=activity)
Can i plot this by using HARDataset?
Pleaze Guide me
This tutorial will show you how to plot a time series:
https://machinelearningmastery.com/time-series-data-visualization-with-python/
Thank you dear Jason for you kind response but i want to plot time series graph i.e., https://3qeqpr26caki16dnhd19sv6by6v-wpengine.netdna-ssl.com/wp-content/uploads/2016/12/Minimum-Daily-Temperature-Yearly-Line-Plots.png
For axis (X,Y and Z) only for Single Activity using HARDataset.
Kindly provide me code for it, I am not enough familiar with Python.
Sorry, I cannot write custom code for you.
Ok Dear, Can you give me some hint; HARDataset has many files, how can i relate to them so as it will be possible for me to solve above problem
Perhaps this tutorial will help:
https://machinelearningmastery.com/how-to-model-human-activity-from-smartphone-data/
Thank you Dear.
hi Jason,
do you have any examples in order to synchronize and interpolate accelerometer and gyroscope data
let’s say that we have a data frame
activity timestamp accx accy accz timestamp gurx gury gurz
0 walking 1585564269527 -0.625122 -0.343262 0.357971 1585564269513 10.785061 3.288872 -5.015244
the timestamps form the two sensors are in different scale
Perhaps you can adapt the examples here:
https://machinelearningmastery.com/resample-interpolate-time-series-data-python/
yeah, i have already seen this. First, i must convert the unix time into date and time and then use the examples
thanks Jason
Hi Jason,
Nice explanation, as always. I have a query that how to do the same (video classification) with multiple videos. I mean, any suggestion for one label for one video?
Perhaps explore a CNN-LSTM and ConvLSTM for this type of problem.
Any good resource/tutorial if you can please suggest?
Perhaps start here:
https://machinelearningmastery.com/start-here/#deep_learning_time_series
Hello Jason,
Thank you for this tutorial. In fact I have a question:
I can see that the size of the sliding window is 128. I can also see that the sliding window is moving by 64 values in the data (50% overlap). So in the new window, there is 128 / 2 = 64 redundant data. Do you have an explanation for the choice of 50%? What is the effect of choosing other options?
Thank you for your answer.
The choice of framing is arbitrary. Find what works well/best for your specific model and dataset.
sir how does this use for live data.
First fit your model on all available data, then make predictions on new data by calling model.predict()
Hey Jason,
I also enjoyed this article. I am currently working on a similar project and I wanted to know if the sliding window is always necessary in a time series classification problem, where the idea is to predict the classes of one multivariate time series data?
To be concrete, lets say I have 9 subjects, each of them has 22 features extracted and from the measurements i get for every feature 376 data points, which makes 22*376=8272 datapoints for each person. Each subject then belongs to either class 1,2,3 or 4. How do I decide on the window size and why is this necessary? I actually also read you articles about how to convert a Time Series to a supervised learning problem, but I did not see how this is applicable for my problem.
Thanks a lot!
Yes it is.
Choose the window size based on the size that results in the best performing models on your data.
Great tuto,
Is there any transfert learning to be applied in the CNN part of the CNN-LSTM model (like VGG, RestNet, Inception…) in order to make time series classification ?
You could. I don’t recall writing an example of exactly this, sorry.
Hi Jason, I stuck at loading the data. Should I include the directory of the workspace for example C:\Users\… ? I’ve done everything but the dataset just could not load.
Thank you in advance!
Yes, have the dataset and code in the same directory.
Sir, i am adapting the your example using my data. I have 4 input features and 1 target.
The target value is in 0s and 1s , y= [0,1,0]
and the input shapes is (timestep, features) [7,4] and output[1] since i have got only one target i.e num_y-signals.
The proposed model is:
model.add(LSTM(100, activation=’relu’, return_sequences=True,
input_shape=(X_train.shape[1], X_train.shape[2])))
model.add(Dense(num_y_signals, activation=’softmax’))
I am getting the following error
ValueError: A target array with shape (14388, 1) was passed for an output of shape (None, 7, 1) while using as loss
binary_crossentropy.This loss expects targets to have the same shape as the output.
Can you please help me where i have got wrong please
Sorry to hear that, preparing LSTM input can be tricky. Perhaps this will help:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Sir,
Thank you for the response
I have y classes with 0s and 1s and how to get the predicted label classes for the corresponding feature.
I got this output but donot know how to interpret.
# define vector
probs = np.array(yhat)
print(probs.shape)
# get argmax
labels = np.argmax(probs, axis=-1)
print(labels)
Output
(3592, 7, 2)
[[1 1 1 … 1 1 1]
[1 1 1 … 1 1 1]
[1 1 1 … 1 1 1]
…
[1 1 1 … 1 1 1]
[1 1 1 … 1 1 1]
[1 1 1 … 1 1 1]]
Thank You
You can use predict() and argmax on the results:
https://machinelearningmastery.com/argmax-in-machine-learning/
Or can use predict_classes() and get the class labels directly:
https://machinelearningmastery.com/how-to-make-classification-and-regression-predictions-for-deep-learning-models-in-keras/
Hi Jason,
Thanks for the great tutorial.
I want to play with CNN based model to make a Rank list. I have student’s profile(20 students) in a class and I want to create a model, which is able to make a rang list in the class.
The student profile is only one row with 10 features (grades from different curriculum and achievements).
Dou you have any idea, or suggestion for me?
Yes, perhaps this framework will help:
https://machinelearningmastery.com/how-to-develop-a-skilful-time-series-forecasting-model/
Hi, Jason,
I have a question, please. The samples and the timestamps are the same? I mean that when we have 50hz frequency and we decide to take 2s we then take 100-timestep which is equal to 100 samples. Am I write?
No, see this:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Hello. It was a great post, thank you.
When I use the following code to predict the model:
pred = model.predict( testX )
I get the following error for convlstm:
——————————————ERROR——————————————————————————–
ValueError: Error when checking input: expected conv_lst_m2d_4_input to have 5 dimensions, but got array with shape (2947, 128, 9)
————————————————————————————————————————–
and for cnn-lstm is:
——————————————————ERROR——————————————————————–
ValueError: Error when checking input: expected time_distributed_21_input to have 4 dimensions, but got array with shape (2947, 128, 9)
————————————————————————————————————————–
How can I fix it? Thank You
You need to change the shape of your data to match the expectation of your model or change your model to meet the shape of your data.
Sir I was trying to apply CNN+LSTM on EEG data whose X shape is (11540,20). How should I reshape it to feed into neural network? Thank you.
This will help you understand how to reshape your data:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
From there you can add one more dimension for the CNN-LSTM.
Hi Jason, amazing article for novices. I had a question regarding the Zero-one offset. Why there is a need to do it? Why do you want the class labels to start from zero and not from one? Also it changes the dimension of the “trainy” and “testy” to (7352, 6) and (2947, 6) respectively. Why is that happening?
Thanks!
Good question, in one sense it is arbitrary, it is all just code, in another sense this is a property of how we can efficiently represent the multinomial distribution in code:
https://machinelearningmastery.com/discrete-probability-distributions-for-machine-learning/
Also array offset in python is zero based, which also makes it simpler to read/understand.
Hi,
Thank you for your good tutorial.
I read your previous article.
https://machinelearningmastery.com/cnn-models-for-human-activity-recognition-time-series-classification/
Could you please explain the differences between 1D Convolutional Neural Network Models and CNN-LSTM / ConvLSTM?
if I understand correctly, there are 4 types:
recurrent neural networks include:
LSTM
CNN-LSTM
ConvLSTM
and one-dimensional convolutional neural networks include:
CNN
Am I correct? Thank you very much.
Yes, they are different model architectures, see this:
https://machinelearningmastery.com/how-to-develop-lstm-models-for-time-series-forecasting/
I have one doubt, my work is similar to this. But instead of human activity, I have to tell whether the vehicle is moving straight or zigzag path. I have sensor readings, how can I implement it with neural network model? Thanks in advance
Perhaps you can use the tutorial as a starting point and adapt it for your specific dataset.
Hello, I want to apply RNN on data extract from 2D skeleton of openpose. So I have 36 features (body co ordinates) and a label for that pose. So my X_train has a shape of (4000,36) and Y_train has a shape of (4000,5). How can I use RNN to classify this data?
I am currently using a dense network for pose estimation in real time
Y = (4000,5) because I have use one-hot encoding for 5 classes
This may help you prepare your data for LSTMs:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Hello,
I trained my LSTM network given the all vector (time series vector of n samples).
But what I need is to feed my network sample by sample and get the results after each sample and not to give the whole vector,
How do I do that?
Use a loop over your samples and pass them in one at a time and make a prediction.
Thank you, this is what I am doing now.
what do you mean by ‘samples’? sample of a given series?
if so:
1. I will do prediction sample after sample, but when I get new series what should I do?
The default is stateful = false, so there is reset between batches.
2. should I used stateful = True?
How do I use it ? should I use it during training\predication?
2. if I set return_sequences = true, than I get the hidden state… is it the outputs of the prediction for each sample in my series?
I
Good question, this explains the terms:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Perhaps experiment and discover what works best for your model+data.
If just ” pass them in one at a time and make a prediction” than after every predication the states will be reset, no? because every predication is like a new batch, no?
Correct.
Test if this matters or not for your model/data.
If not, you may as well use a CNN or MLP as it will be faster.
Hi jason,
i have a question please,
you said above that:
1. The model learns to extract features from sequences of observations and how to map the internal features to different activity types.
2. we do feature learning on raw data.
3. The benefit of using LSTMs for sequence classification is that they can learn from the raw time series data directly, and in turn do not require domain expertise to manually engineer input features. The model can learn an internal representation of the time series data and ideally achieve comparable performance to models fit on a version of the dataset with engineered features.
How can this possible? How can LSTM learn that in this timestep the activity is let’s say walking? which features it recognize in order to come to that decision,
any help will be higly valuable.
Thank you
It learns the unknown underlying function that maps patterns in the input data to a target class.
This is called function approximation and it is what neural nets do well, perhaps this will help:
https://machinelearningmastery.com/neural-networks-are-function-approximators/
oh so LSTM does not calculate ant features like min-max etc? Is function approximation valid in the unsupervised scenario right?
Unsupervised algorithms like clustering methods do a type of function approximation, e.g. best separation of data by natural groupings.
Hello,
First of all thanks for this code !!
I have a question, where did you write the path of your dataset location ?
I’m trying to run it but it always says : ‘FileNotFoundError: [Errno 2] File HARDataset/train/Inertial Signals/total_acc_x_train.txt does not exist: ‘HARDataset/train/Inertial Signals/total_acc_x_train.txt”
Thanks for your help,
Robin
The code is expecting to run in the same directory as the unzipped dataset.
You can change the path to the dataset in the code if you like.
Thanks ! It’s working 🙂
I’m happy to hear that.
Greetings!
Would you like to explain to me the number of layers, cells, and type of LSTM that you have performed in the first model HAR LSTM-RNN?
The models were configured with a little trial and error.
This may help:
https://machinelearningmastery.com/faq/single-faq/how-many-layers-and-nodes-do-i-need-in-my-neural-network
Thanks a lot, dear Jason
Please let me explain the above LSTM model that how many layers, memory cells, and neuron(s) that you have used?
From LSTM types (e.g., one-to-one, one-to-many, etc.,) which one that have you used?
It is a time series classification model. One prediction/classification per input time step.
The number of nodes and layers were chosen after some trial and error.
Hi Jason,
Hopefully this is a quick question, but I haven’t been able to find an answer anywhere.
What is the best way to preprocess timestamp data for an LSTM?
Should timestamps be scaled or normalised like other continuous variables? Or is there a more appropriate approach?
Specific context: I’m using an LSTM sequential model (from Keras in Python) for time series anomaly detection (classification) on network packet captures from a simulated network
This can help prepare data for LSTMs:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
It is a good idea to normalize or standarlize your data prior to modeling, perhaps test and confirm in your case.
Hello,
In this example you set lstm unit =100.
Does it means that the memory of the lstm is only of 100 time stamps?
It will forget what happend 101 time stamps before?
I found the following defenistion for units:
STM with units=1 the key values (f, i, C, h) are scalar; and with units=n they’ll be vectors of length n
I am confuzed because diagram of unrolled lstm network looks like a chain of lstm units…. is this the same unit above?
Thx
Yasmin
No, it means it has 100 units and each takes the entire input sequence.
This will help:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
And this:
https://machinelearningmastery.com/faq/single-faq/how-is-data-processed-by-an-lstm
Hi jason,
i have two questions, please. In the setting of human activity recognition, we have the many to one sequence? Meaning that in each time step we input a window of data and we have a prediction y hat.
also, I see that there been used different models like encoders-decoders, bidirectional lstm, transformer, models with attentions etc. Are these models applicable to the human activity recognition area?
In this case, one to one, e.g. one input observation, one classification.
Yes, try a suite of models and see what works best for your dataset.
Dear Jason,
Really V.good material thank you for it,
i want to apply the same ideas of the tutorial to my problem. I have raw data for 385 patients, each 2 hours i take some measurements for this patients to predict his state want to enter ICU or not where each patient has 2 rows of measurements. the shape of data for LSTM was (385, 2, 85) where 385 is number of samples/ patients, 2 is number of sequences for each patient, and 85 is a group of measurements as features that calculated for each sequence of time.
my problem is the shape of input for CNNLSTM model, i don’t know how to get n_steps, n_length because my problem is simple than the problem in the tutorial .
please help me to know how to reshape trainx and what is n_steps, n_length in my problem?
Also, what is the input shape of CNNLSTM for CONV1D layer?
the same thing for ConvLSTM Network Model???
Thanks
Thanks!
Good question, these suggestions will help (preparing data for LSTMs is the same as preparing it for 1D CNNs):
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Hi Jason,
In the similar way i am trying to implement Time series classification using keras. It is 33000×251 size data for each instance.
Training data is about 18x33000x251
test data is about 12x33000x251
When trained with model, validation accuracy is not going beyond 60%
Can you please suggest what we can do in this case to improve validation accuracy. my model is as below:
from tensorflow.keras.layers import Dropout
from tensorflow.keras.callbacks import EarlyStopping
early_stop=EarlyStopping(monitor=”val_loss”, mode=”min”, verbose=1, patience=25)
from tensorflow.keras.regularizers import l2
model3=Sequential()
model3.add(LSTM(256, input_shape=(seq_len, 251), kernel_regularizer=l2(0.01), recurrent_regularizer=l2(0.01), bias_regularizer=l2(0.01)))
model3.add(Dense(150, activation=”relu”))
model3.add(Dense(50, activation=”relu”))
model3.add(Dense(1, activation=”sigmoid”))
model3.compile(loss=”binary_crossentropy”, optimizer=”adam”, metrics=[“accuracy”])
model3.fit(x=train,
y=train_target,
epochs=600,
callbacks=[early_stop],
validation_data=(test, test_target))
These tutorials will give you ideas on how to diagnose and improve the performance of your model:
https://machinelearningmastery.com/start-here/#better
Hi Jason,
I’m using the following model for eye tracking time series data but accuracy is only 64%.Any suggestions how to improve it?
model = Sequential()
model.add(TimeDistributed(Conv1D(filters=128, kernel_size=1, activation=’relu’), input_shape=(1,3, 3)))
model.add(TimeDistributed(Conv1D(filters=128, kernel_size=1, activation=’relu’)))
model.add(TimeDistributed(Dropout(0.5)))
model.add(TimeDistributed(MaxPooling1D(pool_size=1)))
model.add(TimeDistributed(Flatten()))
model.add(LSTM(250))
model.add(Dropout(0.5))
model.add(Dense(100, activation=’relu’))
model.add(Dense(3, activation=’softmax’))
model.compile(loss=’categorical_crossentropy’, optimizer=’adam’, metrics=[‘accuracy’])
model.fit(X_train, y_train, epochs=30, validation_split=0.2)
_, accuracy = model.evaluate(X_test, y_test, batch_size=64, verbose=0)
print(“accuracy”, accuracy)
Yes, many suggestions right here:
https://machinelearningmastery.com/start-here/#better
Hello Jason,
Thank you for this amazing tutorial, as always.
Out of curiosity, a simple logistic regression with l1 penalty trained on the features (training set) provides 95.9% accuracy on the test set, while the whole LSTM one seems to be at around 92% max.
What are your thoughts on this?
Thank you in advance !
The tutorial is teaching how to use a method, not solve a specific standard dataset in the most effective manner.
Use the model that performs the best on your problem.
Hi, Jason:
In “model.compile(loss=’categorical_crossentropy’, optimizer=’adam’, metrics=[‘accuracy’]” , is it possible to change ‘accuray’ by ‘f1_macro’ or something similar? (I have an imbalanced dataset). I tried it but got an error.
Thanks in advance
Yes, see this tutorial:
https://machinelearningmastery.com/custom-metrics-deep-learning-keras-python/
And this:
https://machinelearningmastery.com/how-to-calculate-precision-recall-f1-and-more-for-deep-learning-models/
Hi Mr. Jason,
I’m new to machine learning and I have some questions.
I’m trying to plot a confusion matrix, displaying classification report for the CNN-LSTM model.
I comment the line #_, accuracy = model.evaluate(testX, testy, batch_size=batch_size)
and add this line
history = model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, validation_split=0.2)
then I add a function
But I keep getting errors like name ‘history’ is not defined
Even I write the plotting codes out of function it also return the same error, do you know how to solve this error please?
Or perhaps do you know how can I plot confusion matrix and display classification report?
Your reply will be highly appreciated, thank you very much.
Perhaps this example will help:
https://machinelearningmastery.com/how-to-calculate-precision-recall-f1-and-more-for-deep-learning-models/
Thank you Mr. Jason, I will try to read the tutorial again .
I have one more question, I have read the FAQ part for “What is the difference between “CNN-LSTMs” and “ConvLSTMs”?”
But I saw something called DeepConvLSTM (https://www.mdpi.com/1424-8220/16/1/115), is this same as CNN-LSTM?
I’m not familiar with that paper, thanks for the link.
Great article, I used the LSTM approach you highlighted and successfully could distinguish among 3 different activities. I created my own data from essentially multiple “runs” of executing an activity (e.g. throwing a ball). In a recording I would do the activity multiple times. As I converted sensor data (accelerometer, gyroscope) into the right shape using your series_to_supervised with ~30 lags and 100 observations. My question to you is once I successfully characterize the activity, how can I count the number of times the activity occurred in my original data. To clear my original sequence may represent say 10 instances of an activity (e.g. throwing a ball 10 times), the conversion of this sequence may with lags and observations create roughly 500-1000 samples. So though I classify the activity successfully I don’t understand how to count the number of times I did the activity. Any suggestions?
Well done!
You may need to write code to interpret the predictions from your model. E.g. classical programming/software engineering where the model is just one piece.
I guess I don’t understand what you are suggestion. For example if an original sequence of sensor data is say 20 seconds and you do an activity every 5 seconds (say throw a ball in my example) I would like to count “4 times a ball was thrown). Are you suggesting I should somehow understand there are 4 cycles of data in this 20 seconds? How? Furthermore if mixed kicking a ball versus throwing a ball the cycles would look different (and the model would see that) but again how to equate it back to original data.
Can you expand on your suggestion? Maybe an equivalent example would be if the activity was walking and model accurately determined walking, how would one go about counting the number of steps in that walking sequence?
I don’t know the full extent of your project, but it sounds like you are using the model to classify sequences of data.
Once classified, you can write some if-statements to interpret the predictions, e.g. in this interval there were this many of this and that actions.
If that is not the case, perhaps I don’t follow.
Let me explain. Say you ask a person to wear a sensor and perform a series of different activities. Example throw a football 10 times, then shoot a basketball 10 times, then hit a tennis ball with a racquet ten times. Say each sequence takes ~ 25 -30 seconds. Using your blog concepts I can create sequences of times series data with known labels and converting these time series with series_to_supervised. That is I can get convert these into samples, timesteps, features. Building a model on this gives great accuracy with LSTM. And if I then have the person repeat one aspect say throw the football 5 times, convert the data and do a model.predict the model accurately determines the person’s activity is throwing a football. The problem is I know the person is throwing a football but I don’t know how many times the person is throwing the football (other than I told them to throw it n times). So once I know what the person’s activity is I’d like to count the discrete number of times the person does that activity? Does this better explain the challenge?
Thanks for elaborating, I think you can address your concern by redefining the problem.
Some ideas:
Perhaps you can re-define what you classify from the activity to the identification of a specific sub-action of an activity, like catching the ball. Then count catches
Perhaps you can train the model to identify the beginning and end of each activity so you can count iterations?
Perhaps you can train the model to identify the activity performed between repetitions (e.g. standing waiting to throw/catch football before actually doing it)?
I have a query , is this for a sequence to label or for sequence to sequence classification problem.
Beacuse i too have a similar windowed input i.e trainX(462,250,6) and its corresponding trainY(462,250.12) where in trainX there are 6 features(3 accelerometer + 3 gyrometer) and 12 output classes in trainY.
So i am having an error with input shapes of the Conv + Lstm architecture which you were using beacuse i think i need to change the shape of trainY(as i a doing sequence to sequence prediction).
Could you please throw some light on it?
The above example is sequence classification.
Instead of classifying individual data point, what would be a good approach towards classifying the entire time series?
I recommend evaluating a suite of approaches in order to discover what works well or best for your dataset.
How can I use LSTM for real-time data classification?
Call model.predict() on new data.
Thanks Jason for quick response.
I’m collecting data from EEG and model is already pre-trianed, so I need to get real-time output every 10 seconds. How can I do this real-time classification output every 10 or 20 seconds?
TIA
Perhaps you can fit a model on historical data then make predictions on new data as it comes it. Try it and compare it to a model that is updated frequently.
Hey Jason,
I have adapted this example for a research project where I take in keypoints from a body instead of acceleration data.
I am a new to Keras and I was wondering if you know the syntax to print out each individual result in the LSTM. For example, I want to see which results are wrong and which are correctly estimated. Thanks!
The LSTM will output a matrix with one row per sample. You can print each row to get the output foe each input sample.
Hi Jason, I have a total of 16 simulation runs (Time Series) for a healthy and 15 faulty cases. I want to build a classifier to differentiate the type of fault. There are a total of 8 features (sensor signals) for each simulation run and each data set consists of 20,000 data points. I would like to know if any of the above mentioned methods can help to build this classifier.
Perhaps try an LSTM and compare results to other models to see if they have skill.
i am looking for similar data to the dataset used in this toturiel please
Maybe this will help:
https://machinelearningmastery.com/faq/single-faq/where-can-i-get-a-dataset-on-___
Hello jason, first this article is very helpful for my studying keras lstm.
And i have a question
In dataset there are 6 label, but why did you make zero-offset class value?
(trainy = trainy – 1
testy = testy – 1)
because i want to classify all of label.
We use a one hot encoding for class labels, requiring the data to be first ordinal encoded with a zero-offset.
hi Jason, Thank you for your post. I have a question about preparing dataset for training. I have a wide range of time series files, let say filename1, filename2, …. The sample rate is the same, but these files may be of different length. For instance, filename1 may contain 20 min of the data, and filename2 can contain 10 min of the data. I mean my data is not continuous and also it is coming from different sensors. To prepare my files as input for lstm/cnn I should merge them together and assign the class that each data point belongs to? is that right?
This may give you some ideas:
https://machinelearningmastery.com/data-preparation-variable-length-input-sequences-sequence-prediction/
Thank you. Whether a window in one row of the train dataset should be necessarily related to the next row? I mean should windows have overlap or they can be from different data files?
It’s your choice whether windows overlap or not.
Perhaps test a few approaches and compare results.
Thank you for your reply. Based on what I understand from lstm windows can contain a discontinues time intervals as well. Is that right? For instance, first window can contain time 0,…,128 and the second one can contain time 200, 328.
There is nothing stopping you.
It may or may not impact the performance of your model on your dataset.
Hi Jason, Thanks a lot for the valuable tutorial. I have a question though. I have a data set collected from a cooling water system sensors and there are 64 features/variables and nearly 12000 records to train with 7 types of faulty data. I’m working on conducting a fault detection & diagnosis on this data set with LSTM approach for multi-class classification(7 fault types and normal).
But Iim finding a bit confused here how to decide the number of time steps that i should use when shaping the input to the LSTM layer since it is now a two dimensional data with the shape of (12000,64). If i do this with a single time step am i not gonna get the full use out of LSTM due to there will be not recurrence of the same cell?
and secondly do i need to conduct a optimal feature extraction for the LSTM before the input layer setup. Or is it something that LSTM will take care of by it self? ( i can use all 64 features without having to do any selection?)
Perhaps you can evaluate different numbers of time steps and discover what works well or best for your specific model and dataset.
LSTM will perform feature extraction automatically.
Hi Jason,
I have dataset of a uni-axial accelerometer. The raw dataset only have one feature and class and the total datapoints are 36000. I injected sensor faults via simulations and added 5 more classes. Now, I have a dataset with one feature, 6 classes and total datapoints 216000.
With the help of your tutorials I made samples of the dataset, each sample has 1000 datapoints. I did sampling in two ways; one with no overlap in this case data shape is [samples= 216, n_timesteps=1000, n_features=1] and the second with overlap skipping just one step and the data shape in this case is [samples= 12006, n_timesteps=1000, n_features=1]. And each sample of the dataset is labeled.
I want to build a classification model using LSTM. I build a single layers LSTM with input layer having the shapes mentioned above and the output layer with 6 nodes/cells. But it’s not working I am getting the following error:
ValueError: Shapes (None, 1) and (None, 6) are incompatible
First, I was thinking to use auto-encoder (for feature extraction) with LSTM (for classification). But, then I decided to train a LSTM first and then go for a hybrid kinda model so that I have clear picture whether the autoencoder is useful or not.
I want to know if there is a problem with my dataset or the choice of algorithm?
I really need your help. Any sort of help and guidance would be great.
Thanks.
This will help you prepare your data for working with LSTMs:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
can’t i predict only one label output if i make multiple time steps in LSTM (which makes return a whole sequence). I think i find it very vague and unclear what the time step is and how we can use it in LSTM.
If the model is designed to predict a sequence, then you will always get a sequence as output for each input sample.
Thanks alot
Hi Jason,
One more question. I’m a little worried about if i have structured my data set in a way it is suitable for a LSTM time series multi-class classification problem. I have different machinery conditions (7 faults and normal condition) logged in 8 different times. And i have setup my data frame by appending all these 8 data files. each data file has 3000 records with 65 columns(sensors) with a label to mention which condition that certain data point falls into.
So i split this data set into x and y and reshape if i want to go with more than one time step.
Is this sort of a scenario valid with such a dataset for LSTM? Really appreciate if you can give me a help on this
Thanks
Perhaps try an LSTM and compare results to other models like 1d cnn and mlp, and other ml models like random forest.
I tried both LSTM and 1D CNN and I get a testing data accuracy of 97.59% for 1D CNN and 98.11 % for LSTM. Training accuracies of both of them are more than 99.20 %. Guess i should go with LSTM.
In LSTM I have done the classification by reshaping both X and Y into 3D with 100 time steps. So I have to take predicted value for the last time step returned from the prediction function as the output right? becoz i get a array of predicted values (many to many). Those should be the states calculate at each time steps with the effect of previous time step state ?
Not sure I follow your question, sorry.
Your model will make one prediction per input sample.
Perhaps this will help if you’re confused with the predictions:
https://machinelearningmastery.com/how-to-connect-model-input-data-with-predictions-for-machine-learning/
Hi Jason,
this is my input data shape for X and Y data.
X_train.shape, y_train.shape
((816, 100, 65), (816, 100, 8))
following is the model
model.add(LSTM(100, dropout=0.1, input_shape=(X_train.shape[1],
X_train.shape[2]),return_sequences=True))
model.add(LSTM(100, return_sequences=True))
model.add(Dense(100, activation=’relu’))
model.add(Dense(8,activation=’softmax’))
model.compile(loss=’categorical_crossentropy’,
optimizer=’adam’, metrics=[‘accuracy’])
Here I have to return the sequence since i have reshaped the Y also into 3D. Then calling the predict function for this i get a result like following for a prediction input of a 10 samples in a sequence
[[2,2,2,2,2,2,3,3,2,3]]
So here i get states for the each time step i have send to predict function right. and what should i get as the output value from these values as the predicted class label for my input sequence? Hope it is clear now
Thanks
Not sure I follow.
The output of your model is a vector of class predictions for a given input.
Also, your model is odd, e.g. it has return_sequences=true for the last LSTM layer, I would not recommend this.
Hi Jason,
I had to return the sequences since I have reshaped not only X_train to ((816, 100, 65) but also y_train to (816, 100, 8)) since I want to train my model using multi time steps (multiple recurrence). Other wise I will get an error saying Invalid output shapes since y output is defined with time steps (816, 100, 8).
Anyway thanks a lot for your response 😉
For sequence classification, when would you use an autoencoder first, and feed the compressed representation (from the trained encoder) into a subsequent classification model?
When the training datasets are “noisy”?
I’m just trying to synthesize across the various LSTM-based sequence classification articles you have presented?
That approach can be effective in many cases, noisy data as you say, also sequences with variable lengths. Try it and see.
Dear Jason,
Thanks for the post!
My data set was not obtained in a complete constant rate. For example, it is 50Hz but only for 5 seconds in every minute. What should I do?
Thanks,
Don
This will give you ideas:
https://machinelearningmastery.com/faq/single-faq/how-do-i-handle-discontiguous-time-series-data
Helpful, thanks for the quick reply!
You’re welcome.
Hi Jason,
Quick question. Is it normal that using my own data and running experiment 20 times it’s getting the same accuracy in all cases? What could be a possible reason for this? Increasing the training data obviously increases the accuracy but it’s not changing from run to run.
Thank you in advance.
If you get the same accuracy, may be you reset your random seed every time? You should not. See this FAQ on random seed https://machinelearningmastery.com/faq/single-faq/what-value-should-i-set-for-the-random-number-seed
Hi Adrian,
It’s solved thank you for your help. I would like to ask something related with input data.
I’m not too sure how should I address the problem of doing the same classification as I have done but instead of using a csv file using more csv’s files with completely different variables.
I’m not sure if I should create 3 different models and combine later or if there’s a different way to solve this. Any hint with this would help me a lot.
Thank you in advance.
“Bagging” or “ensemble learning” is a technique in machine learning doing exactly like what you said. You can definitely create different models and train them independently, and later combine them, such as by taking their average.
Dear Jason,
I am trying to implement a CNN network + LSTM to be able to predict the class based on the sequence of images. My X-train shape is (10, 256, 150, 150, 3). Images’ size is 150*150*3, and each image sequence includes 256 images (so I have 10 image sequences). I defined time-steps to be 256, channels (features) to be 3. This is a part of my code:
X_data = np.reshape(X_data, (10, 256, 150, 150, 3))
Y_data = np.reshape(Y_data, (10, 256, 1))
model = tf.keras.models.Sequential()
model.add(TimeDistributed(Conv2D(filters=64, kernel_size=3, activation=’relu’), input_shape=(None, 256, 150, 150,3)))
model.add(TimeDistributed(Conv2D(filters=64, kernel_size=3, activation=’relu’)))
model.add(TimeDistributed(Dropout(0.5)))
model.add(TimeDistributed(MaxPool2D(pool_size=2)))
model.add(TimeDistributed(Flatten()))
model.add(LSTM(100))
model.add(Dropout(0.5))
model.add(Dense(100, activation=’relu’))
model.add(Dense(3, activation=’softmax’))
model.compile(loss=’categorical_crossentropy’, optimizer=’adam’, metrics=[‘accuracy’])
model.fit(X_data, Y_data, verbose=1, epochs=epochs, callbacks=callbacks)
And I get this error:
ValueError: Input 0 of layer max_pooling2d is incompatible with the layer: expected ndim=4, found ndim=5. Full shape received: (None, 256, 146, 146, 64)
Can you please tell me What is the problem with input_shape and how can I give the network the desired input()?
I tried several different input shapes but every time I get an error regarding the dimension of input.
Sorry I can’t debug your code but what it might help is that the max_pooling2d layer in the error message is not your input layer. Please see if you can trace back where is causing that. One way to do is to remove some layers in the model until you see the error goes away. That can tell you which layer is configured wrong.
Dear Jason,
Thanks again for the post!
I have another question. In my case, I have only 10 time steps (vs. 128 in this post) with no overlap between the time windows, and I have only 2-3 features (vs. 9 in this post). How many LSTM units I should take (in this post 100 units were taken)? Any rule I should follow maybe even some thumb rule? And a similar question regarding the units in the dense layer, which number should I take? Any rules here?
Many thanks,
Don
No rules here. The best bet is to read other people’s code, especially those with similar problem as yours, and try to tweak around their design. Hyperparameters like the number of units can also be underfit or overfit. If you get too few units, for example, your model learn nothing. If you get too much, your model may take too long time to converge and fail to generalize.
Thanks Adrian for the quick reply!
I guess that when Jason set the number of LSTM units to be 100 it was not a random number, and there are reasons he didn’t set it to 10 or 1000. For example, the amount of data puts constraints on the number of free parameters (which are connected to the number of units). Is it possible to get Jason’s view on my question?
Thanks a lot,
Don
Checked with Jason. That’s arbitrary as it is just a demonstration. In reality, as I mentioned before, you need to try out different values and evaluate. Often it is a trade off between model complexity and output accuracy.
Thanks for checking with Jason! Ok, then I’ll try different values.
Thanks again,
Don
Thank you for this wonderful explanation.
Please, I am working on an experiment and would love to know if using LSTM algorithm will be most suitable for my experiment.
I am trying to learn the behavioral pattern of a single individual living in a 4 room house. I can get data (movement from one door to another and the time spent in each room) from the simulation using PIR sensors. I want to be able to predict when the occupant generates an abnormal behavior like, spending more time in the kitchen than usual (maybe he has fallen down and requires the Ambulance).
Also, if the sequence of data collected over a period of time seems normal and all of a sudden a sequence of some seemingly inaccurate data is generated over a short time period and then correct data sequence starts coming in again; I need to be able to predict that this would be a possible sensor malfunction. For instance. If all previous readings are normal and I notice a move from room1 to room4 which is clearly abnormal but all other subsequent readings continues normally; the prediction algorithm should be able to infer this situation as caused by the sensors in doors 2 and 3 not capturing when the occupant passed through them.
Thank you.
I cannot really tell because you didn’t give any design on how you would use the LSTM. This kind of problem can also be applied using other techniques. But if you’re using LSTM, you are remembering some pattern in the network and producing output based on input. If you can quantify what is your input, what would be the output, and why remembering something is required, then you should easily see whether LSTM is a good fit.
Hi Adrian,
I have three models using this method that are exactly the same. In each of these models the input has the same 3 dimensions (changing the values because data is different, but following the same structure of: time steps, 1 ,and features).
The problem I’m facing is that following all of them the same structure, only 2 of those 3 models work. I’m not sure why but in one of them it’s printing the same accuracy all the time, I didn’t really touch anything, the models are the same and the structure of input data is as well.
The only difference I see from this input data compared with the rest is that there are way more variables (160, while on the other cases it’s 2 or just 40 variables), can this be the reason? if not what do you think it could be?
Thank you in advance.
160 vs 2 is very imbalanced. Of course this will hurt (usually, also depends on variation of the data, e.g., whether the 160 are binary variables or floating point values).
Hi Adrian Tam
first of all I really appreciate your precious effort sharing the materials with us. As a beginner I have two questions,
1. the way to use the model for prediction of a new input or a test case.
2. <for another case using ategorical_crossentropy and sofmax, the output of the prediction was like this [[0.08681168 0.11935446 0.09112465 0.09589134 0.10852194 0.08946144
0.10187661 0.11885039 0.0876272 0.10048036]] while I expected to see the integer. I was wondering if you could answer my questions
(1) I think you just borrow the code from this post should work; (2) that’s the nature of softmax. It gives you a list of probability-like value and usually we just pick the one with highest “probability” as the classification output.
Hi Adrian,
I’m not sure why but accuracy is always the same, no matter if I change the data set always sets to a 25% or 0%. I had to make a padding of 0’s to make the second dimension the same in train and test and I’m not sure of this can be the reason for the model to not work properly.
So the structure (and input adding to the model) I have is trainX = (10,1000,140) trainy = (10,8) testX = (3,1000,140) testy = (3,8)
The majority of rows from that 1000 are the padding of 0’s, I got this 1000 because it was the max number of labels of a specific class from the 8 I have to classify, so we include all the data.
I believe this has nothing to do with reseting the seed, if you could please give me some idea of what can be happening I would be very thankful.
Thank you in advance.
I forgot to say that with a different data set of structure trainX = (43, 369, 49) trainy = (43, 8)
testX = (8, 369, 49) testy = (8, 8), the accuracy is always the same but in this case 12.5%. So it shouldn’t be a problem of having a high value on the second dimension.
Here I did a padding of 0’s as well, since originally the input was trainX = (15867, 49)
trainy = (43,8) testX = (2952, 49) testy = (8, 8) after reshaping it.
The original labels have been subdivided in “groups” of 8 (testy) and 43 (trainy) respectively because each group has actions happening many times
I’m looking forward for your response.
Can you explain what are each of the dimension (10,1000,140) means?
Hi Adrian,
The first dimension is the samples: I get this number by counting how many groups of labels are (for example if I have as labels [1,1,1,2,2,3,3,4,4,4,5,6,6,7,8,8,8,1,1,2] I’m putting it in groups of the same numbers that are one after the other so: (1,2,3,4,5,6,7,8,1,2) and counting how many are inside, therefore, 10 in this example case would be the first dimension. Note that (1,2,3,4,5,6,7,8,1,2) would be the new labels, with a to_categorical it gets the shape (10,8).
As second dimension there’s time steps: First I’m counting how many numbers are in each subgroup of labels, for example as we saw, [1,1,1,2,2,3,3,4,4,4,5,6,6,7,8,8,8,1,1,2] would count (3,2,2,3,1,2,1,3,2,1), I’m picking the highest number therefore 3 in this example, and what I do is padding rows of 0’s to all the subgroups so all of them count the same max number so (3,3,3,3,3,3,3,3,3,3). I want all to have same count because later I want to reshape it, and for this they have to be the same size. The second dimension here is that max count between counts of trainy and testy. After this padding I have (huge number due to the padding of many rows, features) and with this I reshape it to have the (samples of grouped labels, max count between trainy and testy,140 features).
As third dimension is the number of features: this is basically that I have 140 variables/features.
I hope I explained it correctly enough so you understand what is the input and why it’s not working and not giving errors. Thank you in advance, I’m stuck with this part and can’t advance more.
If you set most of your time series steps as zero, of course you don’t get a good result. You’re basically providing nothing as input and expecting something as output.
I used a mask layer to ignore the 0’s rows too and it’s working in the same way. What do you think it can be a solution for this?
It works if the input structure is (1000, 1, 140) and labels (1000,8) but I don’t think it’s the best solution.
I think there is a mistake in data reshaping here. If the purpose of ConvLSTM is to convolved both temporal and spatial information, then the input data to ConvLSTM network should be
(samples, n_subsequence , len_subsequence, n_features , channels). To give you an example, if the input to your LSTM model is (samples, len_sequence = 128, n_features =9), then this is how you must reshape it for ConvLSTM: (samples, n_subsequence =4 , len_subsequence =32, n_features = 9, channels=1)
Hi Prof, thanks for the tutorial, I want to ask.
I’m trying to create a new raw dataset, let’s call it data A, from accelerometer and gyroscope sensors like Anguita but it only consists of 6 features (x,y,z for each sensor). Then I want to create a transfer learning model, in this case pre trained data, from the data A. Then I want to classify it using LSTM as you did.
My question is, is it possible to make pre trained data from data A? Because based on what I read and found, pre trained data is used for image and classification methods such as CNN.
Then is the data suitable for me to use for classification using LSTM or should I use other data for pre trained or classification?
Please help, Professor. Thank you very much.
Transfer learning is useful because the training time is excessively long. This is the case of image applications with 100 layers of network. In LSTM, usually you don’t have that many and therefore it may not be very beneficial. The other use of transfer learning is to prevent some layers to be trained when transferred to a different application. This is useful when you think the feature extraction in application A is useful to application B. Do you see your use case fits this?
Thank you for your explanation, Prof. Adrian Tam.
I want to ask again, I think the output of transfer learning is a new dataset that can be used for classification using LSTM, is that wrong?
I actually also want to use transfer learning data to several methods such as LSTM and bidirectional LSTM because I am curious, whether the learning model is only suitable for images or not, and that requires experimentation.
Besides that, I also haven’t found a tutorial for creating pre trained data from data A (non-images), so this makes me even more curious.
Please help, thank you Prof Adrian Tam
Transfer learning is to train a new model weight based on the model weight of another application. See https://machinelearningmastery.com/transfer-learning-for-deep-learning/
Hi Adrian,
I would like to know what would be good ways to experiment with the results obtained with LSTM. Any way to show and compare within a graph? I have the accuracy and a confusion matrix but I want to know more about results.
Thank you
What is the purpose of comparing? I think answering this question helps you to drive forward.
Dear Prof Adrian, thanks for the tutorial, I want to ask.
I want to combine the attention mechanism on the page
https://machinelearningmastery.com/attention-long-short-term-memory-recurrent-neural-networks/
with 2 layers LSTM for HAR classification as on this page, but I’m confused about how to do it. I’m trying to add the code I got for the attention mechanism
===============================================
from tensorflow.hard.layers import Layer
from tensorflow.hard import backend as K
class Attention(Layer):
def __init__(self, return_sequences=True):
self.return_sequences = return_sequences
super(Attention,self).__init__()
def build(self, input_shape):
self.W=self.add_weight(name=”att_weight”, shape=(input_shape[-1],1),
initializer=”normal”)
self.b=self.add_weight(name=”att_bias”, shape=(input_shape[1],1),
initializer=”zeros”)
super(Attention,self).build(input_shape)
def call(self, x):
e = K.tanh(K.dot(x,self.W)+self.b)
a = K.softmax(e, axis=1)
output = x*a
if self.return_sequences:
return output
return K.sum(output, axis=1)
===============================================
and change the model in your coding lstm to
===============================================
# fit and evaluate a model
def evaluate_model(trainX, trainy, testX, testy):
verbose, epochs, batch_size = 0, 15, 64
n_timesteps, n_features, n_outputs = trainX.shape[1], trainX.shape[2], trainy.shape[1]
model = Sequential()
model.add(attention(return_sequences=True)) # receive 3D and output 3D
model.add(LSTM(32, return_sequences=True))
model.add(LSTM(32, return_sequences=True))
model.add(attention(return_sequences=False)) # receive 3D and output 2D
model.add(Dropout(0.1))
model.add(Dense(n_outputs, activation=’softmax’))
model.summary()
model.compile(loss=’categorical_crossentropy’, optimizer=’adam’, metrics=[‘accuracy’])
# fit network
model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose)
# evaluate models
_, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0)
return accuracy
===============================================
but i get error:
This model has not yet been built. Build the model first by calling
build()or callingfit()with some data, or specify aninput_shapeargument in the first layer(s) for automatic build.Please help. Thanks.
Update:
I changed evaluate_model to
===============================================
# fit and evaluate a model
def evaluate_model(trainX, trainy, testX, testy):
verbose, epochs, batch_size = 0, 15, 64
n_timesteps, n_features, n_outputs = trainX.shape[1], trainX.shape[2], trainy.shape[1]
model = Sequential()
model.add(LSTM(32, input_shape=(n_timesteps,n_features), return_sequences=True))
model.add(attention(return_sequences=True)) # receive 3D and output 3D
model.add(LSTM(32, input_shape=(n_timesteps,n_features), return_sequences=False))
model.add(Dropout(0.1))
model.add(Dense(n_outputs, activation=’softmax’))
model.summary()
model.compile(loss=’categorical_crossentropy’, optimizer=’adam’, metrics=[‘accuracy’])
# fit network
model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose)
# evaluate models
_, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0)
return accuracy
===============================================
And the program is running, but is the placement of the attention and lstm layers correct?
Thanks Prof.
It looks OK to me. Or you can try to put the attention before the Dense layer and see how much difference.
Thank you Prof. Adrian for your answer.
May i ask again?
1. So we can put attention not between LSTM layers? I think attention should be flanked by LSTM because it acts as an encoder-decoder.
2. I’ve tried with LSTM-Attention-LSTM-Dropout-Dense like in my code, but only get 54.913% accuracy, is this weird? Or is there something wrong with my attention mechanism code?
3. I tried as you suggested in the following order:
=========================================
model = Sequential()
model.add(LSTM(32, input_shape=(n_timesteps,n_features), return_sequences=True))
model.add(LSTM(32, input_shape=(n_timesteps,n_features), return_sequences=True))
model.add(Dropout(0.1))
model.add(attention(return_sequences=False)) # receive 3D and output 3D
model.add(Dense(n_outputs, activation=’softmax’))
model.summary()
model.compile(loss=’categorical_crossentropy’, optimizer=’adam’, metrics=[‘accuracy’])
model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose)
_, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0)
return accuracy
=========================================
And i got 89.793% (+/-0.807). That’s an amazing improvement, if I may know, why is this happening?
And why do you suggest putting attention after dropout and before dense? doesn’t the dropout select the neuron to be input?
Thanks and sorry to bother you.
Attention should be after encoder and before decoder so the decoder can find what is the most important to look at. So if your encoder is not doing a good job to encode, the potential of attention layer is not in good use. That maybe the reason for your 54% accuracy.
Dropout layer is for regularization prevent overfitting. It usually put before dense to make sure the first half of the network did not learn something not generalized.
Thank you for your explanation Prof. very reasonable.
Then I tried to compare it by putting the attention before the dropout, and strangely the resulting accuracy is better.
LSTM-LSTM-DROPOUT-ATTENTION-DENSE
89.708% (+/-0.977)
89.725% (+/-0.747)
LSTM-LSTM-ATTENTION-DROPOUT-DENSE
90.624% (+/-0.595)
90.445% (+/-1.024)
This is quite surprising for me.
If you notice the error ranges are overlapping, I would not be so confident to say one is better than another.
Dear Prof Adrian, I want to ask again.
I’m studying and having trouble, i’m trying to use the code like the following (which comes from your page) and now i’m curious how to find out the weight of each input before and after using the attention mechanism?
Because I want to know how attention actually works in graphics.
Thank you very much.
In keras, layerx.get_weight() would give you the weight of “layerx”
Hi Prof. Thank you before, I’m still trying to find out the weight changes of each epoch for each layer.
May I ask again? What is the difference between batch size on:
verbose, epochs, batch_size = 0, 15, 64
which will be used in:
model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose)
_, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0)
with a value of 100 in the following LSTM:
model.add(LSTM(100, input_shape=(n_timesteps,n_features)))
model.add(Dense(100, activation=’relu’))
Because from what I know, the value of 100 is also a batch size.
Thank you very much.
100 is not a batch size, but a unit size. You have 100 hidden states in the LSTM layers in this case.
I can’t figure out where is the error arose. Can you post your full code? The error you mentioned said what it complained. You probably called evaluate() without first calling fit().
Dear Jason,
Thank you for sharing such a great post with us!
I have a question about the input format of LSTM.
As we know that the input format of LSTM is that [samples, time steps, features].
In this post, you input the time series based on a fixed window of 2.56 seconds.
As the frequency is 50Hz, so per window contains 2.56*50 (128) points, so you set the time steps as 128.
as for features, actually, we should call them the variables. The number of features is 9.
So the input format of LSTM is [samples, 128,9].
I noticed that you said that there were 561 features are extracted from each window.
Could we take these features as input for the LSTM?
If we can, how should we set the timestep and the number of features for LSTM?
Best wishes,
Dandan
Hi Jason – we used this tutorial to help us get started in generating an LSTM model that we are using to analyze Drosophila behavior. Is there a specific way you would like us to cite you in a publication?
Thank you, John
This is one of the FAQ:
For example:
Jason Brownlee, Deep Learning with Time Series Forecasting, Machine Learning Mastery, Available from https://machinelearningmastery.com/machine-learning-with-python/, accessed November 6th, 2018.
Hi Sir Adrian, thank you for this tutorial. I want to ask. What is the meaning of _, in this code?
Sorry because i’m new to python and this machine learning world.
# fit network
model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose)
# evaluate model
_, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0)
return accuracy
That’s a Python special variable. It means “don’t care”. Your model.evaluate() should return you the loss and accuracy, and you assign to “_, accuracy” to mean you don’t care the loss, but save the accuracy to the variable “accuracy”
So if I want to use the loss function, I just write
loss, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0)
return loss, accuracy ?
Yes
Thank you Sir. May I ask again?
I have data with 2833 rows, 6 features and 8 classes.
I tried to apply your LSTM code as above but I get error:
expected shape=(None, 128, 6), found shape=(None, 6)
how to determine the correct shape? Thank you very much
I think you have specified that your input to LSTM to be a sequence of 128. So you need to reshape your data in such as way too.
I tried adding the following code to convert it to an array:
trainX = np.array(trainX)
then reshape it to:
trainX = np.reshape(trainX, (trainX.shape[0], 64, trainX.shape[1]))
but i get error:
ValueError: cannot reshape array of size 11874 into shape (1979,64,6)
I don’t know where 11874 came from whereas my shape data for trainX is
print(trainX.shape) = (1979, 6)
Can you help me ?
if trainX.shape is (1979,6), then you’re asking to reshape trainX into (1979,64,6) and 1979x64x6=11874
How can you rearrange a 1979×6 array into 1979x64x6 array?
Hi Sir, thank you for this amazing tutorial. May i ask?
When I print the shape trainx, trainy, testx, and testy. produce as follows:
train x = (7352, 128, 9)
train y = (7352, 6)
test x = (2947, 128, 9)
test y = (2947, 6)
But when I print the lstm model, it appears like this:
Layer (type) Output Shape Param #
=================================================================
lstm_2 (LSTM) (None, 128, 32) 5376
lstm_3 (LSTM) (None, 128, 32) 8320
dropout_1 (Dropout) (None, 32) 0
dense_1 (Dense) (None, 6) 198
1. On train x = (7352, 128, 9) and test x = (2947, 128, 9)
Value 128 indicates what? because as far as i understand, 7352 and 2947 is data rows and 9 is features
2. Then on the lstm layer (None, 128, 32), the number 128 reappears and there is none and the number 32. what do none and the number 32 mean?
3. There are also new numbers 5376 and 8320. What do these numbers show?
These things confuse me. Please help. Thank you very much
(1) the 128 is the sequence length
(2) 32 is the number of units of LSTM, so you’re producing a vector of 32 as the states
(3) those are the number of parameters in the layer. When you do training, you’re updating these much of parameters
So, is this right, Sir?
train x = (data train rows, sequence length, features)
train y = (data train rows, label)
test x = (data test rows , sequence length, features)
test y = (data test rows, label)
Layer (type) Output Shape Param #
=================================================================
lstm_2 (LSTM) (None, sequence length, units of LSTM) parameters
lstm_3 (LSTM) (None, sequence length, units of LSTM) parameters
dropout_1 (Dropout) (None, units of LSTM) 0
dense_1 (Dense) (None, label class) parameters
Then what does none mean? Sorry, I have to learn it slowly.
None is a placeholder for the batch size, which can be varying
Hi Sir, i want to ask again.
Do you have the dataset in excel form? Because I don’t know how to save the result file of your load_dataset() method and I want to try to learn between the data in excel and the data output method you use.
Thank you again.
No. Indeed rarely we use Excel because CSV or TSV are simpler formats. If you need Excel file, you may make use of the to_excel() function in pandas.
update: I found the data format in the form of excel (xlsx). but when I use it, why does the error appear?
ValueError: Input 0 of layer "sequential" is incompatible with the layer: expected shape=(None, 7352, 561), found shape=(None, 561)
I only change the code when reading the file to the dataframe:
path_data_train= r'/content/drive/MyDrive/train_data.xlsx'
df_train = pd.read_excel(path_data_train)
path_data_test= r'/content/drive/MyDrive/test_data.xlsx'
df_test = pd.read_excel(path_data_test)
trainX = df_train.iloc[:, 0:-1]
trainy = df_train['label']
testX = df_test.iloc[:, 0:-1]
testy = df_test['label']
and parameter of n_timesteps, n_features, n_outputs in evaluate_model
def evaluate_model(trainX, trainy, testX, testy):
verbose, epochs, batch_size = 0, 15, 64
n_timesteps, n_features, n_outputs = trainX.shape[0], trainX.shape[1], 1
model = Sequential()
model.add(LSTM(100, input_shape=(n_timesteps,n_features)))
model.add(Dropout(0.5))
model.add(Dense(100, activation='relu'))
model.add(Dense(n_outputs, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(trainX, trainy, epochs=epochs, batch_size=batch_size, verbose=verbose)
_, accuracy = model.evaluate(testX, testy, batch_size=batch_size, verbose=0)
return accuracy
Obviously the input array has a different shape now. You should check and compare the shape of the numpy arrays and trace down to what caused it.
If so, how many timesteps do I need, Sir?
and how to calculate good timesteps?
I use 64 because usually the number for timesteps between 32,64, and 128.
Thank you.
Not necessarily a power of 2 like 32, 64, 128; but it really has no way to tell how many timesteps you would need. It depends on the problem.
Hi Jason, thanks for this amazing tutorial! I have a few questions.
I’ve downloaded the dataset and tried to follow your tutorial, what I wanted to ask:
1. each dataset file in the train\inertial signal\ folder has 128 columns and 7352 rows (which combined there are 128*9 = 1152 columns), whereas there is also an X_train file in the train folder which only has 561 columns and 7352 rows which are not found in your code. is it true that you are not using this X_train file?
2. when I try one of the files (body_acc_x_train.txt) and try to split the first row data into 128 data in 1 row, there is a lot of redundant data, like row 2 and 3, row 4 and 5, etc.). Why this happened?
3. I tried to retrieve data from the accelerometer sensor myself, but there are only 3 columns, namely x, y, z with many rows of data, while in the file in this tutorial there are 128 data columns, where does that data come from?
Thank you.
(1) the training data is from a split; you should not combine the columns
(2) redundant data is not a problem, indeed you can’t expect human keep moving all the time
(3) check the feature description file in the UCI repository link will explain the features
Hi Jason:
I am trying to replicate the full script of your first example (an LSTM Network Model)
But this line: model.add(LSTM(100, input_shape=(n_timesteps,n_features))), is generating this error:
NotImplementedError: Cannot convert a symbolic Tensor (lstm/strided_slice:0) to a numpy array. This error may indicate that you’re trying to pass a Tensor to a NumPy call, which is not supported
May be It is keras-numpy compability
Do you know a workaround?
Thank you.
Hi Fernando…The following discussion addresses this common issue and provides some ideas on how to remedy it.
https://stackoverflow.com/questions/58479556/notimplementederror-cannot-convert-a-symbolic-tensor-2nd-target0-to-a-numpy
Also, you may want to try Google Colab for the time being if you cannot quickly resolve the compatibility issue with your local installation of Python and Tensor Flow.
Hi Jason:
Thanks for tutorial
I want to run CNN-1D or RNN on my dataset, I saw this error: ” ValueError: Input 0 of layer “sequential_14″ is incompatible with the layer: expected shape=(None, 25, 10), found shape=(1, 250)”
Dataset includes 250 columns for a feature only (like voltage time series) and 20,000 rows (each row has one label and problem is multi-class) , I generate this 250 data in 5 sec and 50 sample/sec (50 Hz). I have tried but I can’t set parameters, Please guide me.
Hi Saeid…Thanks for asking.
I’m eager to help, but I just don’t have the capacity to debug code for you.
I am happy to make some suggestions:
Consider aggressively cutting the code back to the minimum required. This will help you isolate the problem and focus on it.
Consider cutting the problem back to just one or a few simple examples.
Consider finding other similar code examples that do work and slowly modify them to meet your needs. This might expose your misstep.
Consider posting your question and code to StackOverflow.
These are my setting in your code:
verbose, epochs, batch_size = 0, 15, 1
n_timesteps, n_features, n_outputs = 25, 10, 1
error:
File “/usr/local/lib/python3.7/dist-packages/keras/engine/training.py”, line 1021, in train_function *
return step_function(self, iterator)
File “/usr/local/lib/python3.7/dist-packages/keras/engine/training.py”, line 1010, in step_function **
outputs = model.distribute_strategy.run(run_step, args=(data,))
File “/usr/local/lib/python3.7/dist-packages/keras/engine/training.py”, line 1000, in run_step **
outputs = model.train_step(data)
File “/usr/local/lib/python3.7/dist-packages/keras/engine/training.py”, line 859, in train_step
y_pred = self(x, training=True)
File “/usr/local/lib/python3.7/dist-packages/keras/utils/traceback_utils.py”, line 67, in error_handler
raise e.with_traceback(filtered_tb) from None
File “/usr/local/lib/python3.7/dist-packages/keras/engine/input_spec.py”, line 264, in assert_input_compatibility
raise ValueError(f’Input {input_index} of layer “{layer_name}” is ‘
ValueError: Input 0 of layer “sequential_14” is incompatible with the layer: expected shape=(None, 25, 10), found shape=(1, 250)
Hi Saeid…Thanks for asking.
I’m eager to help, but I just don’t have the capacity to debug code for you.
I am happy to make some suggestions:
Consider aggressively cutting the code back to the minimum required. This will help you isolate the problem and focus on it.
Consider cutting the problem back to just one or a few simple examples.
Consider finding other similar code examples that do work and slowly modify them to meet your needs. This might expose your misstep.
Consider posting your question and code to StackOverflow.
I simply wanted to write down a quick word to say thanks to you for that wonderful information you are showing on this site.
Hi Jason,
I’ve been trying to run the “attention mechanism” with my data:
The data is of shape (605,16) [ ( time_steps , features ) ]
I followed your discussion with Zainal and run the code:
model = Sequential()
model.add(LSTM(64,input_shape=(X.shape[1],X.shape[2]), return_sequences=True))
model.add(attention(return_sequences=True))
model.add(Dropout(0.1))
model.add(Dense(10, activation=’sigmoid’))
model.add(Dense(1, activation=’sigmoid’))
model.summary()
model.compile(loss=’binary_crossentropy’, optimizer=’adam’, metrics=[‘accuracy’])
This same code run perfectly fine on one machine but I get the following error on different machine:
Dimensions must be equal, but are 605 and 64 for ‘dense_1/MatMul’ (op: ‘MatMul’) with input shapes: [?,605], [64,10]
Can you please tell what causes the error and how to resolve it.
Thank you
Best regards
Hi Ahmad…Thanks for asking.
I’m eager to help, but I just don’t have the capacity to debug code for you.
I am happy to make some suggestions:
Consider aggressively cutting the code back to the minimum required. This will help you isolate the problem and focus on it.
Consider cutting the problem back to just one or a few simple examples.
Consider finding other similar code examples that do work and slowly modify them to meet your needs. This might expose your misstep.
Consider posting your question and code to StackOverflow.
Hi Jason
Thank you for this amazing tutorial
I’m a novice in this field so I’m quite confused about the filepath( I have provided path of one txt file here). I have downloaded and saved the data as you have instructed but my code is showing error:
” categorical[np.arange(n), y] = 1
IndexError: index -2 is out of bounds for axis 1 with size 1″.
It would be of great help if you could guide me through this.
Thanks.
Hi Anam…The following may be of interest:
https://rollbar.com/blog/python-indexerror/
how to deploy the model on live data. i mean webcam or any recorded videos.
Hi akhil…The following may be of interest:
https://towardsdatascience.com/neural-networks-for-real-time-audio-stateless-lstm-97ecd1e590b8
Hello, I would like to thank you for this very interesting tutorial.
Actually, I want to track the position of the person using the same features. Let’s assume that we have used the same sensors but the output here is different. The model will predict a 2D position, so we are predicting (x,y). This will be a regression problem.
The question is: should I normalize the position variable or I train the model with the original variables’ values?
Hi Jenny…I would recommend that you try it both ways. There are instances for which normalization is not critical. Let us know what you find out.
Hi, I enjoyed the tutorial so much.
I need to classify the faulty transmission line connected to a 5-line bus of a power system according to the three-phase currents of all lines by LSTM. Then, the fault types including 3-phase, 2-phase, 2-phase-ground, and 1-phase-ground must be classified along with the faulty phases determined. The data labels show the faulty line and the fault types at each timestep. How can I develop the LSTM network?
Hi Hossein…The following resource may be of interest:
https://machinelearningmastery.com/how-to-develop-lstm-models-for-multi-step-time-series-forecasting-of-household-power-consumption/