Comparing machine learning methods and selecting a final model is a common operation in applied machine learning.
Models are commonly evaluated using resampling methods like k-fold cross-validation from which mean skill scores are calculated and compared directly. Although simple, this approach can be misleading as it is hard to know whether the difference between mean skill scores is real or the result of a statistical fluke.
Statistical significance tests are designed to address this problem and quantify the likelihood of the samples of skill scores being observed given the assumption that they were drawn from the same distribution. If this assumption, or null hypothesis, is rejected, it suggests that the difference in skill scores is statistically significant.
Although not foolproof, statistical hypothesis testing can improve both your confidence in the interpretation and the presentation of results during model selection.
In this tutorial, you will discover the importance and the challenge of selecting a statistical hypothesis test for comparing machine learning models.
After completing this tutorial, you will know:
- Statistical hypothesis tests can aid in comparing machine learning models and choosing a final model.
- The naive application of statistical hypothesis tests can lead to misleading results.
- Correct use of statistical tests is challenging, and there is some consensus for using the McNemar’s test or 5×2 cross-validation with a modified paired Student t-test.
Kick-start your project with my new book Statistics for Machine Learning, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.
- Update Oct/2018: Added link to an example of using McNemar’s test.

Statistical Significance Tests for Comparing Machine Learning Algorithms
Photo by Fotografías de Javier, some rights reserved.
Tutorial Overview
This tutorial is divided into 5 parts; they are:
- The Problem of Model Selection
- Statistical Hypothesis Tests
- Problem of Choosing a Hypothesis Test
- Summary of Some Findings
- Recommendations
Need help with Statistics for Machine Learning?
Take my free 7-day email crash course now (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
The Problem of Model Selection
A big part of applied machine learning is model selection.
We can describe this in its simplest form:
Given the evaluation of two machine learning methods on a dataset, which model do you choose?
You choose the model with the best skill.
That is, the model whose estimated skill when making predictions on unseen data is best. This might be maximum accuracy or minimum error in the case of classification and regression problems respectively.
The challenge with selecting the model with the best skill is determining how much can you trust the estimated skill of each model. More generally:
Is the difference in skill between two machine learning models real, or due to a statistical chance?
We can use statistical hypothesis testing to address this question.
Statistical Hypothesis Tests
Generally, a statistical hypothesis test for comparing samples quantifies how likely it is to observe two data samples given the assumption that the samples have the same distribution.
The assumption of a statistical test is called the null hypothesis and we can calculate statistical measures and interpret them in order to decide whether or not to accept or reject the null hypothesis.
In the case of selecting models based on their estimated skill, we are interested to know whether there is a real or statistically significant difference between the two models.
- If the result of the test suggests that there is insufficient evidence to reject the null hypothesis, then any observed difference in model skill is likely due to statistical chance.
- If the result of the test suggests that there is sufficient evidence to reject the null hypothesis, then any observed difference in model skill is likely due to a difference in the models.
The results of the test are probabilistic, meaning, it is possible to correctly interpret the result and for the result to be wrong with a type I or type II error. Briefly, a false positive or false negative finding.
Comparing machine learning models via statistical significance tests imposes some expectations that in turn will impact the types of statistical tests that can be used; for example:
- Skill Estimate. A specific measure of model skill must be chosen. This could be classification accuracy (a proportion) or mean absolute error (summary statistic) which will limit the type of tests that can be used.
- Repeated Estimates. A sample of skill scores is required in order to calculate statistics. The repeated training and testing of a given model on the same or different data will impact the type of test that can be used.
- Distribution of Estimates. The sample of skill score estimates will have a distribution, perhaps Gaussian or perhaps not. This will determine whether parametric or nonparametric tests can be used.
- Central Tendency. Model skill will often be described and compared using a summary statistic such as a mean or median, depending on the distribution of skill scores. The test may or may not take this directly into account.
The results of a statistical test are often a test statistic and a p-value, both of which can be interpreted and used in the presentation of the results in order to quantify the level of confidence or significance in the difference between models. This allows stronger claims to be made as part of model selection than not using statistical hypothesis tests.
Given that using statistical hypothesis tests seems desirable as part of model selection, how do you choose a test that is suitable for your specific use case?
Problem of Choosing a Hypothesis Test
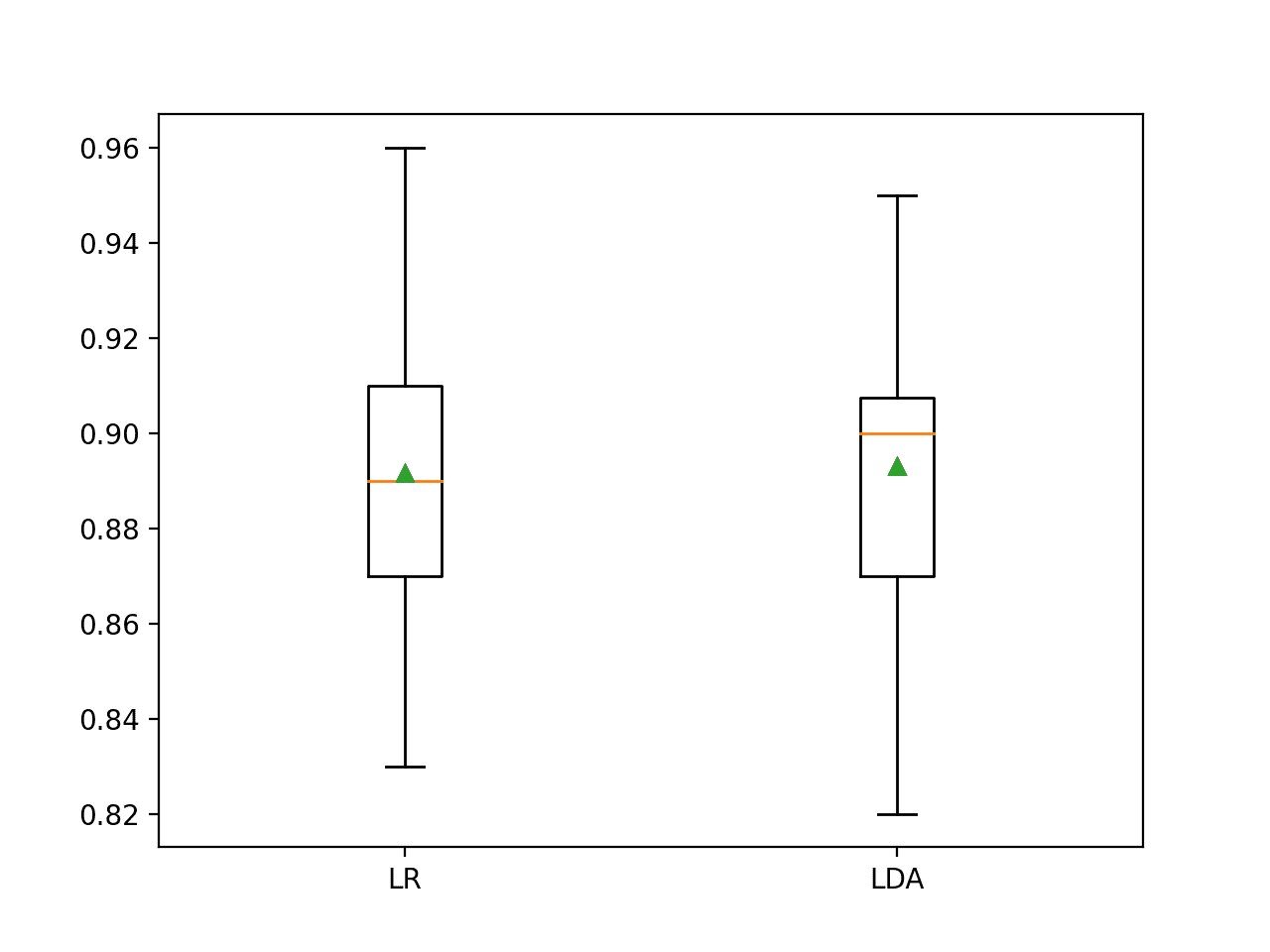
Let’s look at a common example for evaluating and comparing classifiers for a balanced binary classification problem.
It is common practice to evaluate classification methods using classification accuracy, to evaluate each model using 10-fold cross-validation, to assume a Gaussian distribution for the sample of 10 model skill estimates, and to use the mean of the sample as a summary of the model’s skill.
We could require that each classifier evaluated using this procedure be evaluated on exactly the same splits of the dataset via 10-fold cross-validation. This would give samples of matched paired measures between two classifiers, matched because each classifier was evaluated on the same 10 test sets.
We could then select and use the paired Student’s t-test to check if the difference in the mean accuracy between the two models is statistically significant, e.g. reject the null hypothesis that assumes that the two samples have the same distribution.
In fact, this is a common way to compare classifiers with perhaps hundreds of published papers using this methodology.
The problem is, a key assumption of the paired Student’s t-test has been violated.
Namely, the observations in each sample are not independent. As part of the k-fold cross-validation procedure, a given observation will be used in the training dataset (k-1) times. This means that the estimated skill scores are dependent, not independent, and in turn that the calculation of the t-statistic in the test will be misleadingly wrong along with any interpretations of the statistic and p-value.
This observation requires a careful understanding of both the resampling method used, in this case k-fold cross-validation, and the expectations of the chosen hypothesis test, in this case the paired Student’s t-test. Without this background, the test appears appropriate, a result will be calculated and interpreted, and everything will look fine.
Unfortunately, selecting an appropriate statistical hypothesis test for model selection in applied machine learning is more challenging than it first appears. Fortunately, there is a growing body of research helping to point out the flaws of the naive approaches, and suggesting corrections and alternate methods.
Summary of Some Findings
In this section, let’s take a look at some of the research into the selection of appropriate statistical significance tests for model selection in machine learning.
Use McNemar’s test or 5×2 Cross-Validation
Perhaps the seminal work on this topic is the 1998 paper titled “Approximate Statistical Tests for Comparing Supervised Classification Learning Algorithms” by Thomas Dietterich.
It’s an excellent paper on the topic and a recommended read. It covers first a great framework for thinking about the points during a machine learning project where a statistical hypothesis test may be required, discusses the expectation on common violations of statistical tests relevant to comparing classifier machine learning methods, and finishes with an empirical evaluation of methods to confirm the findings.
This article reviews five approximate statistical tests for determining whether one learning algorithm outperforms another on a particular learning task.
The focus of the selection and empirical evaluation of statistical hypothesis tests in the paper is that calibration of Type I error or false positives. That is, selecting a test that minimizes the case of suggesting a significant difference when no such difference exists.
There are a number of important findings in this paper.
The first finding is that using paired Student’s t-test on the results of skill estimated via random resamples of a training dataset should never be done.
… we can confidently conclude that the resampled t test should never be employed.
The assumptions of the paired t-test are violated in the case of random resampling and in the case of k-fold cross-validation (as noted above). Nevertheless, in the case of k-fold cross-validation, the t-test will be optimistic, resulting in a higher Type I error, but only a modest Type II error. This means that this combination could be used in cases where avoiding Type II errors is more important than succumbing to a Type I error.
The 10-fold cross-validated t test has high type I error. However, it also has high power, and hence, it can be recommended in those cases where type II error (the failure to detect a real difference between algorithms) is more important.
Dietterich recommends the McNemar’s statistical hypothesis test in cases where there is a limited amount of data and each algorithm can only be evaluated once.
McNemar’s test is like the Chi-Squared test, and in this case is used to determine whether the difference in observed proportions in the algorithm’s contingency table are significantly different from the expected proportions. This is a useful finding in the case of large deep learning neural networks that can take days or weeks to train.
Our experiments lead us to recommend […] McNemar’s test, for situations where the learning algorithms can be run only once.
Dietterich also recommends a resampling method of his own devising called 5×2 cross-validation that involves 5 repeats of 2-fold cross-validation.
Two folds are chosen to ensure that each observation appears only in the train or test dataset for a single estimate of model skill. A paired Student’s t-test is used on the results, updated to better reflect the limited degrees of freedom given the dependence between the estimated skill scores.
Our experiments lead us to recommend […] 5 x 2cv t test, for situations in which the learning algorithms are efficient enough to run ten times
Refinements on 5×2 Cross-Validation
The use of either McNemar’s test or 5×2 cross-validation has become a staple recommendation for much of the 20 years since the paper was published.
Nevertheless, further improvements have been made to better correct the paired Student’s t-test for the violation of the independence assumption from repeated k-fold cross-validation.
Two important papers among many include:
Claude Nadeau and Yoshua Bengio propose a further correction in their 2003 paper titled “Inference for the Generalization Error“. It’s a dense paper and not recommended for the faint of heart.
This analysis allowed us to construct two variance estimates that take into account both the variability due to the choice of the training sets and the choice of the test examples. One of the proposed estimators looks similar to the cv method (Dietterich, 1998) and is specifically designed to overestimate the variance to yield conservative inference.
Remco Bouckaert and Eibe Frank in their 2004 paper titled “Evaluating the Replicability of Significance Tests for Comparing Learning Algorithms” take a different perspective and considers the ability to replicate results as more important than Type I or Type II errors.
In this paper we argue that the replicability of a test is also of importance. We say that a test has low replicability if its outcome strongly depends on the particular random partitioning of the data that is used to perform it
Surprisingly, they recommend using either 100 runs of random resampling or 10×10-fold cross-validation with the Nadeau and Bengio correction to the paired Student-t test in order to achieve good replicability.
The latter approach is recommended in Ian Witten and Eibe Frank’s book and in their open-source data mining platform Weka, referring to the Nadeau and Bengio correction as the “corrected resampled t-test“.
Various modifications of the standard t-test have been proposed to circumvent this problem, all of them heuristic and lacking sound theoretical justification. One that appears to work well in practice is the corrected resampled t-test. […] The same modified statistic can be used with repeated cross-validation, which is just a special case of repeated holdout in which the individual test sets for one cross- validation do not overlap.
— Page 159, Chapter 5, Credibility: Evaluating What’s Been Learned, Data Mining: Practical Machine Learning Tools and Techniques, Third Edition, 2011.
Recommendations
There are no silver bullets when it comes to selecting a statistical significance test for model selection in applied machine learning.
Let’s look at five approaches that you may use on your machine learning project to compare classifiers.
1. Independent Data Samples
If you have near unlimited data, gather k separate train and test datasets to calculate 10 truly independent skill scores for each method.
You may then correctly apply the paired Student’s t-test. This is most unlikely as we are often working with small data samples.
… the assumption that there is essentially unlimited data so that several independent datasets of the right size can be used. In practice there is usually only a single dataset of limited size. What can be done?
— Page 158, Chapter 5, Credibility: Evaluating What’s Been Learned, Data Mining: Practical Machine Learning Tools and Techniques, Third Edition, 2011.
2. Accept the Problems of 10-fold CV
The naive 10-fold cross-validation can be used with an unmodified paired Student t-test can be used.
It has good repeatability relative to other methods and a modest type II error, but is known to have a high type I error.
The experiments also suggest caution in interpreting the results of the 10-fold cross-validated t test. This test has an elevated probability of type I error (as much as twice the target level), although it is not nearly as severe as the problem with the resampled t test.
— Approximate Statistical Tests for Comparing Supervised Classification Learning Algorithms, 1998.
It’s an option, but it’s very weakly recommended.
3. Use McNemar’s Test or 5×2 CV
The two-decade long recommendations of McNemar’s test for single-run classification accuracy results and 5×2-fold cross-validation with a modified paired Student’s t-test in general stand.
Further, the Nadeau and Bengio further correction to the test statistic may be used with the 5×2-fold cross validation or 10×10-fold cross-validation as recommended by the developers of Weka.
A challenge in using the modified t-statistic is that there is no off-the-shelf implementation (e.g. in SciPy), requiring the use of third-party code and the risks that this entails. You may have to implement it yourself.
The availability and complexity of a chosen statistical method is an important consideration, said well by Gitte Vanwinckelen and Hendrik Blockeel in their 2012 paper titled “On Estimating Model Accuracy with Repeated Cross-Validation“:
While these methods are carefully designed, and are shown to improve upon previous methods in a number of ways, they suffer from the same risk as previous methods, namely that the more complex a method is, the higher the risk that researchers will use it incorrectly, or interpret the result incorrectly.
I have an example of using McNemar’s test here:
4. Use a Nonparametric Paired Test
We can use a nonparametric test that makes fewer assumptions, such as not assuming that the distribution of the skill scores is Gaussian.
One example is the Wilcoxon signed-rank test, which is the nonparametric version of the paired Student’s t-test. This test has less statistical power than the paired t-test, although more power when the expectations of the t-test are violated, such as independence.
This statistical hypothesis test is recommended for comparing algorithms different datasets by Janez Demsar in his 2006 paper “Statistical Comparisons of Classifiers over Multiple Data Sets“.
We therefore recommend using the Wilcoxon test, unless the t-test assumptions are met, either because we have many data sets or because we have reasons to believe that the measure of performance across data sets is distributed normally.
Although the test is nonparametric, it still assumes that the observations within each sample are independent (e.g. iid), and using k-fold cross-validation would create dependent samples and violate this assumption.
5. Use Estimation Statistics Instead
Instead of statistical hypothesis tests, estimation statistics can be calculated, such as confidence intervals. These would suffer from similar problems where the assumption of independence is violated given the resampling methods by which the models are evaluated.
Tom Mitchell makes a similar recommendation in his 1997 book, suggesting to take the results of statistical hypothesis tests as heuristic estimates and seek confidence intervals around estimates of model skill:
To summarize, no single procedure for comparing learning methods based on limited data satisfies all the constraints we would like. It is wise to keep in mind that statistical models rarely fit perfectly the practical constraints in testing learning algorithms when available data is limited. Nevertheless, they do provide approximate confidence intervals that can be of great help in interpreting experimental comparisons of learning methods.
— Page 150, Chapter 5, Evaluating Hypotheses, Machine Learning, 1997.
Statistical methods such as the bootstrap can be used to calculate defensible nonparametric confidence intervals that can be used to both present results and compare classifiers. This is a simple and effective approach that you can always fall back upon and that I recommend in general.
In fact confidence intervals have received the most theoretical study of any topic in the bootstrap area.
— Page 321, An Introduction to the Bootstrap, 1994.
Extensions
This section lists some ideas for extending the tutorial that you may wish to explore.
- Find and list three research papers that incorrectly use the unmodified paired Student’s t-test to compare and choose a machine learning model.
- Summarize the framework for using statistical hypothesis tests in a machine learning project presented in Thomas Dietterich’s 1998 paper.
- Find and list three research papers that correctly use either the McNemar’s test or 5×2 Cross-Validation for comparison and choose a machine learning model.
If you explore any of these extensions, I’d love to know.
Further Reading
This section provides more resources on the topic if you are looking to go deeper.
Papers
- Approximate Statistical Tests for Comparing Supervised Classification Learning Algorithms, 1998.
- Inference for the Generalization Error, 2003.
- Evaluating the Replicability of Significance Tests for Comparing Learning Algorithms, 2004.
- On estimating model accuracy with repeated cross-validation, 2012.
- Statistical Comparisons of Classifiers over Multiple Data Sets, 2006.
Books
- Chapter 5, Evaluating Hypotheses, Machine Learning, 1997.
- Chapter 5, Credibility: Evaluating What’s Been Learned, Data Mining: Practical Machine Learning Tools and Techniques, Third Edition, 2011.
- An Introduction to the Bootstrap, 1994.
Articles
- Student’s t-test on Wikipedia
- Cross-validation (statistics) on Wikipedia
- McNemar’s test on Wikipedia
- Wilcoxon signed-rank test on Wikipedia
Discussions
- For model selection/comparison, what kind of test should I use?
- How to perform hypothesis testing for comparing different classifiers
- Wilcoxon rank sum test methodology
- How to choose between t-test or non-parametric test e.g. Wilcoxon in small samples
Summary
In this tutorial, you discovered the importance and the challenge of selecting a statistical hypothesis test for comparing machine learning models.
Specifically, you learned:
- Statistical hypothesis tests can aid in comparing machine learning models and choosing a final model.
- The naive application of statistical hypothesis tests can lead to misleading results.
- Correct use of statistical tests is challenging, and there is some consensus for using the McNemar’s test or 5×2 cross-validation with a modified paired Student t-test.
Do you have any questions?
Ask your questions in the comments below and I will do my best to answer.





Thanks again for awesome post and simplifying it for better understanding.
Thanks, I’m glad it helped.
This article seems to suggest we should be comparing models on the training set. But shouldn’t we focus on the differences in performances on the test set?
Yes, models are trained on the training set and their skill evaluated and compared on a hold out test set.
Is it possible to run a statistical test on repeated measurements of the same model? Let’s say we have 10 run of the same neural network on a particular dataset, but with different initializations. Does it make any sense? Which test should be used?
Yes, this is exactly the situation described in the above post for 5×2 CV.
Is it okay to simply use a Paired Sample T-test, which has the underlying assumption that the samples are dependent?
No, I recommend re-reading this post.
If you have a highly unbalanced dataset and you are predicting the minority class. Even if you do down-sampling and you don’t mind having more false positives as you are predicting a rare event. Then, a t-test may not be that bad, isn’t it?
(By the way, it was a good summary on the topic)
I’m not sure that these issues interact.
Can McNemar’s Test or 5×2 CV be used for compare different machine learning regressors?
5×2 CV can be used for regression, McNemar’s Test is for classification only.
Hi Jason,
Great article as always! I am eager to try out the 5×2 CV on a real world dataset.
Regarding the following line: “Instead of statistical hypothesis tests, estimation statistics can be calculated, such as confidence intervals”
I am wondering how confidence intervals can be calculated for a neural net model. I would normally begin by calculating the standard error for the model’s forecast and then multiple it by the t-value corresponding to the confidence level that I am interested in, so as to arrive at the +/- confidence interval range. But the standard error of the forecast happens to be the RMS of the standard error of the model plus the standard error for each parameter of the model, I can probably get the former easily by just squaring the model’s RMS loss on the training set. But how to get the parameter error? Getting that value requires my knowing the value of each parameter in the final trained model – something that the AI library would not easily expose. I cannot ignore the parametric error either, given how large a component of the forecast error it can potentially be for a neural net model because of the sheer number of parameters involved.
Let me know if I am missing something in my train of thought that you can help me set straight on. Or is this is a real problem with neural nets i.e. not able to use traditional statistical techniques to arrive at the confidence funnel of the forecast made by the model?
Thanks
Great question!
A very simple and effective approach is to use the bootstrap:
https://machinelearningmastery.com/calculate-bootstrap-confidence-intervals-machine-learning-results-python/
Thanks! Will check it out.
Hi Jason,
The tutorial provides a good understanding for selecting appropriate tests. However, which non-parametric test is most suitable when comparing multiple classifiers on a single dataset?
Can we apply Wilcoxon singed ranked test in such this case?
… and running 10-folds CV
Pair-wise tests between classifiers, probably with a modified t-test or wilcoxon.
Hello Jason,
Great article as usual!
I am facing the problem to select the best regression algorithm after obtaining the cross-validated results for several of them (SVR, etc.).
After reading your post I was wondering what approach should I employ in order to do it generic in a function in which given several results (e.g from cross_val_score function) for each algorithm this function returns which is better based on a statistical test (or other approach).
Thanks for your help and congratulations for your blog.
Kind Regards
I’d recommend the 2×5 CV approach with the modified Student’s t-test.
Hi Jason,
In one research paper a classification accuracy is reported for one dataset as 78.34 +/- 1.3 for 25 runs. (This 1.3 can be 95% CI or standard dev.).
I also run my algorithm on the same dataset with 25 runs and calculated as 74.9 +/- 1.05 as 95% CI using student’s t value.
Now only plotting and seeing the overlap can we decide about significance or not. I want to know is there a significant difference b/w their and my results.
What about doing ANOVA and how could we do it as I don’t have 25 runs from the paper. Can I generate fake random data with their mean and std. dev. and then do ANOVA.
What could be the difference (or how to tackle) if they reported std. dev as +/- instead of 95% CI.
Thanks.
Perhaps contact the author and ask what the score represents?
Perhaps reproduce the finding yourself?
Perhaps make a reasonable assumption and proceed.
What I wrote earlier to generate random data to reproduce with similar kind of mean and std. dev and then use ANOVA, does it make sense? The test data set for all 25 runs will be different, so I believe it cannot be a paired test. Thanks.
The described methodology makes me very nervous.
Perhaps talk to a statistician?
I don’t know what feeling to show, happy or sad!!
It’s now confirmed that in a paper the authors use std. dev after +/-. So I generate 25 random numbers from normal distribution with their mean and sigma. Then I make ANOVA table and box plot them. Of course for every different run there’s some difference because of random numbers. I’m not sure whether it could be presented as reasonable justification for the significance?
Thanks again.
Hi!
This post mostly deals with cross-validation when comparing algorithms, Is there any guidance for using paired statistical tests with a single train/test split, where the difference in error for each training example becomes a single sample?
Is the independence assumption here violated because the predictions are all coming from the same model?
Thanks!
Good question, perhaps you could start with a paired Student’s t test:
https://machinelearningmastery.com/parametric-statistical-significance-tests-in-python/
Hi Jason,
I asked participants to learn 9 conversations and then used an NPC classifier to look at 2 brain regions to see if the classifier was able to accurately distinguish between each conversation.
Do I use a paired students t-test (AKA dependent means t-test) or the Wilcoxon signed-rank test?
Thanks!
Probably the Student’s t-test, but check the distribution of the data to confirm it’s Gaussian.
Hi Dr Brownlee
I have three classification algorithms, and I want to stack them instead of selecting one that performs best among them. I have performed model correlation analysis between my sub-algorithms and stacked those that are weakly correlated. I am not interested in checking which one performs best between the three algorithms, but I want to stack them to get better results.
My question is; in this case, do I need to perform a significance test between my sub-models or between the meta-classifier and the sub-classifiers? Is statistical significance test necessary at all in my scenario?….we disagree with my supervisor
I did 5 experiments. In all the experiments, I use a different combination of predictor variables to predict/classify the same thing, and the 5th experiment produces the best results. In all the 5 scenarios, stacking outperforms the 3 sub-classifiers.
Is it necessary for me to perform a significance test between the models? if yes, at what level do I do the test, and why?
Thank you very much your website is very helpful
I would not mind a link to a scientific paper after your comments.
Great work!
No, sig tests are not required.
Hi Jason,
I have run 10 Xgboost models, each predicting the probability of the customer opening the email for each of these products.
Eg : Model 1 output : Prob(cust opening email of prod 1)
Model 2 output : Prob(cust opening email of prod 2)
All of these models have been built on the same features but different underlying population (training data consists of people who were sent emails in the past). However, the models have been scored on the entire population of customers.
Can i directly compare these probabilities and say cust1 is most likely to open email of prod 5 because his prob is highest for prod5?
If not, can you please throw some light on the approach to transform these probabilities to being comparable?
Models can be compared if they have something in common, e.g. same training data/model/target/etc.
Perhaps take a step back and consider the inputs and outputs of the experiment, what is being compared.
Hey, thanks for this really useful post.
Regarding the use of McNemar’s test for deep learning models:
Dietterich writes about McNemar’s: “it does not directly measure variability due to the choice of the training set or the internal randomness of the learning algorithm. A single training set R is chosen, and the algorithms are compared using that training set only. Hence, McNemar ’s test should be applied only if we believe these sources of variability are small. ”
Internal randomness in deep learning classifiers is high, as the random seed matters a lot. However, you write that McNemar’s test is a good option for deep learning. Why? Simply because we dont have any better alternatives 🙂 ?
Correct. You are trading off the ability to run a model multiple times (expensive) vs the instability of the test under variance (some error).
Hey, me again 🙂
So what test would you recommend for the typical deep learning situation:
– Our dataset is split into two non-overlapping datasets: training and test set.
– We trained a few, say 5, random seeds of each model on the training set and computed the test set performance.
I can’t find an appropriate way of doing it:
– McNemar doesn’t seem appropriate because we have multiple runs / random seeds.
– 5 x 2 CV doesn’t seem appropriate because we haven’t done CV (we only train on the one set and test on the other, not the other way around). (As a consequence, we only have 5 measures of performance for each model, not 10)
– simply comparing the 5 performance measures (one from each run, e.g. mean test set accuracy) between the models with e.g. an unpaired t-test seems very inefficient. This approach doesn’t utilize a) the fact that each measure is actually calculated from potentially thousands of samples (depending on the size of the test set), and b) the fact that these samples are “paired”.
Finally, which is probably a whole new question: The methods you suggested all presume that we know how the test data points were classified by our model. What if we don’t have that knowledge? For example, I currently have the situation that I have the class-scores for each data point (outputs of the final sigmoid/softmax layer of my NN-classifier), but I haven’t determined thresholds for classification. I don’t see any other reason to determine these thresholds since I can report e.g. the AUC-ROC of the models without doing so. Are there any statistical tests I can use to compare the models without having the actual classes for each data point?
I am totally aware that it’s not your job to solve my statistical quarrels, but if you happened to find me questions interesting or think that future readers might benefit from your answer as well, I would be very grateful 🙂
Most don’t do anything, just a single run and report results, or multiple runs and ensemble the models.
That is the default you are looking to improve upon, a low bar.
If possible, create one dataset and do 5×2 CV across a few EC2 instances.
Cross entropy can capture the average difference in distribution between real/predicted, instead of a crisp classification result. It’s why we use it as a loss function. More here:
https://machinelearningmastery.com/how-to-score-probability-predictions-in-python/
Ask away, please. I built this site to have these discussions!
Thanks so much for your answer, your response times are truly insane :-)!
For 5×2 CV I would need to create 5 different, random 50/50 splits of the dataset. I’m in academia, so – as you know probably know – for a number of reasons this seems infeasible:
– Often benchmark datasets have fixed training and test sets, and I can’t reassign them.
– Only training on 50% of the data set would be pretty bad if I want state-of-the-art performance
– Maybe most importantly, this practice seems pretty much unheard of in academic deep learning research. So reviewers would think that it is a pretty weird thing to do and I might have trouble getting published
Isn’t there anything useful I can do that fits better with the “normal way” things are done in academic deep learning research (i.e. a test that can be done within the given circumstances: with several runs of every model evaluated on a fixed test set)?
The best thing I could come up with so far was to ensemble the multiple runs from every model and use McNemar’s test or simply a Binomial test on the ensembles. But that has obvious problems as well (doesn’t account for internal randomness of deep learning)
(Side-note: I’m coming from a different field (biomedical research), and I am pretty surprised about the lack of statistical hypothesis testing in deep learning research. I looked at around 10 famous papers yesterday: none did any testing, and the vast majority of them did not even report any standard deviations or used multiple runs… There is probably a good reason for it that I don’t understand yet)
Yes. Use the protocol used in all other papers on the topic, or the average thereof. Play the game, even at the expense of reality.
Skip the sig test. The reason – it’s too expensive in terms of data and compute. The real reason, probably a lack of knowledge of good stats. Also a general science-wide trend toward “estimation statistics” (e.g. effect size, intervals, etc.) and away from hypothesis test.
Yes, ensemble multiple runs, it’s a default in most DL+CV papers, even GAN papers these days.
Alright, thanks again. Both for the answers and for the blog-post, which was by far the most easily accessible treatment of this topic that I could find online.
Thanks.
is it possible to use the paired t-test to compare multi-object detection (include regression and classification together) algorithms based on deep learning method (convolutional neural networks)
Not sure I understand what you’re asking, sorry?
I want to compare my algorithm against other muti-object detection algorithms. the performance of the multi-object detection algorithm is defined using the mean average precision metric since a multi-object detection algorithm combines the regression and classification in the same pipeline. how to use a paired t-test to evaluate the robustness of my algorithm against the benchmark algorithms.
Perhaps a good starting point would be to compare samples of mAP scores across the two methods?
Hi Jason,
I want to compare 4 regression keras models each one of them is 5-fold cross validation, Which test method can I use?
Perhaps pair-wise with a modified t-test?
What do you mean by pair-wise with a modified t-test?
Pair-wise means each combination. Modified t-test is described in the above post.
Does the modified t-test is the paired student’s t test that you descibe in this post:
https://machinelearningmastery.com/parametric-statistical-significance-tests-in-python/
It is a modified version of the paired t-test.
Is there is an example that shows how to select a model?
I don’t think so, thanks for the suggestion!
Can I use a statistical test to compare two deep learning models?
Yes.
How I can compare between a deep learning model and a rule based model that try to solve the same problem?
Based on their skill on a hold out dataset.
So, if the rule based model is tested on a dataset using correlation coefficient and the deep learning model is tested on 5- folds using the correlation coefficient.
Then to compare both models I have to :
1- calculate the performance of the deep learning model as the average of its correlation coefficient on the 5-folds.
2- compare the correlation coefficient of the models where the one with higher correlation will be more efficient than the other one.
Is this what do you mean?
That sounds reasonable, although the choice of correlation coefficient as the metric may be odd. Typically we use prediction error for model selection.
If there is a machine learning regression model that uses a train/test split approach to train it.
and a regression deep learning model that uses a 5-fold cross validation approach to train it.
Can I compare between these two models?
if yes, How can I compare them?
Yes. Based on their performance.
How and they are trained using two different approaches, one of them uses (train/test split) while the other uses (k-fold cross validation)?
As I understand, in order to compare between machine learning or deep learning models, I have to:
1- perform statistical test between the two models.
2- based on p-value we can determine whether the two models have equal or different performance.
Here, in this case, the two model are tested using two different approaches (one uses the train/test split while the other uses the 5-fold cross validation).
So, How can I perform the statistical test between them?
I am not sure, but can I make an assumption that the two models have different performance then I can compare them directly based on their correlation degree.
We do the reverse, assume there is no difference and check if there is a difference, and if there is, select the one with the best mean score.
yes, you means the null hypotheses.
But, I talk about comparing the performance of two machine learning models that applies two different testing methodologies.
since I cannot do a statistical test on them, Is it correct to assume first that they have different performance then select the one with the best score?
If No, How I can compare two models applying two different testing methodologies?
You generally do not.
No, the same testing methodology would have to be used for both algorithms, e.g. k-fold cross-validation.
So, I can measure two machine learning models even if they use two different testing methodology.
But, I cannot calculate any statistical test between them?
if the above two statements are correct,
plz, tell me how can I compare two machine learning models that uses two different testing methodology?
Generally, you must use the same testing methodology for both, otherwise the comparison would not be meaningful.
Thank you Jason very much
You’re welcome.
There is a deep learning model that is used for document classification.
This model works as follow:
1- using embedding layer
2- then use bidirectional GRU as the high level document representation which can be used as features for document classification via softmax.
Can I say that the use of the embedding layer then GRU is used for document representation?
Perhaps. It depends on your audience and what you mean by “representation”.
It might be more accurate to describe the input format and the transforms performed before it is processed by the model.
Is there is any way to compare two machine learning algorithms that apply two different testing methodology?
If yes, Can you tell me How?
There may be. You need to talk to a statistician.
Thank you very much for all your effort. I have a question. Paired student t-test should not be used on scores generated with k-fold cross validation because the scores aren’t independent (one observation is used k-1 times). An alternative is to use bootstraping to generate train and test sets and then train and evaluate your model on them. What then, use paired student t-test on these numbers? Aren’t they also dependent since one observation might as well be used k times (if the procedure was repeated k times)?
The scores from the bootstrap wont be independent either as there is overlap across the drawn samples.
Perhaps re-read the above tutorial and note the corrected version of the t-test.
Ok, so this corrected t-test is like the ordinary t-test, but uses different formulas? Now about the bootstrapping – if I were to compare two models by using let’s say 10 iterations of bootstrapping as an alternative to 10-fold cross-validation, then what would I do with these 2 lists of 10 numbers? What mathematical procedure would I need to undertake in order to see whether the models are significantly different?
Correct.
You may be able to use the same corrected version of the Student’s t-test.
Thank you for answering.
You’re welcome.
Hello Jason, I have been reading your blog posts from quite sometime and they have always been concise and to the point. I am thankful to you for taking the effort to writing them. i want to request a follow up post on this where you can demonstrate an example on how these techniques are implemented using python
Will do. Thanks for the request!
Hi Jason, as always this is a very relevant post and plan to use it at my work. We plan to plot AUC trend of various models computed over a large same dataset (say >100K samples) for all of them. If model 2 is consistently performing better over model 1 for the last n days, isn’t that sufficient to say model 2 relatively better than 1? I feel this is a dumb question, but at the same time, dont understand why we need t-test or similar for understanding “relative” performance of models.
You can compare the mean of two samples directly.
Hypothesis tests allow you to see if the difference in the means is real or a statistical fluke.
The blog is very helpful. Can you suggest me procedure to apply t test to compare two or three algorithms evaluated on many dataset?
Yes, use the procedure described in the above tutorial.
Dear Jason, How can we use the t-test for feature selection in the binary classification case.
See this example:
https://machinelearningmastery.com/feature-selection-with-numerical-input-data/
Hi Jason,
I’m trying to evaluate the results of several classification models. The use case of these models require they produce 0% false positives. How do I test if my false positive rate is really 0?
You can achieve zero false positives by forecasting all cases as negative.
Otherwise, all models make errors in prediction, perhaps this will help:
https://machinelearningmastery.com/tour-of-evaluation-metrics-for-imbalanced-classification/
Why do we have to use the same splits of the data? Why can’t we perform model predictions on different splits and then perform a t-test?
If we use the same splits of the data, we can then use a paired test, which has more statistical power.
how to draw some hypothesis testing to proved the developed models using different machine learning techniques are significantly same or different.
Here is an example:
https://machinelearningmastery.com/hypothesis-test-for-comparing-machine-learning-algorithms/
Is that 5×2 cv modified t-test the same as proposed by Alpaydin named combined ftest 5×2 cv?
Reference: https://www.cmpe.boun.edu.tr/~ethem/files/papers/NC110804.PDF
I don’t know, perhaps check the math or ask the author.
I would like to know if statistical significance test is needed even when I use repeated stratified k fold CV on the dataset?
It can help when comparing the results between algorithms (e.g. the content of the above tutorial).
But it is mentioned in another post in the blog:
In his important and widely cited 1998 paper on the use of statistical hypothesis tests to compare classifiers titled “Approximate Statistical Tests for Comparing Supervised Classification Learning Algorithms“, Thomas Dietterich recommends the use of the McNemar’s test.
Specifically, the test is recommended in those cases where the algorithms that are being compared can only be evaluated once, e.g. on one test set, as opposed to repeated evaluations via a resampling technique, such as k-fold cross-validation.
————————–
So, if I have used repeated K fold , McNemars is not appropriate and I heard that the repeated k fold is enough to show the stability as per the following :
Beleites, C. & Salzer, R.: Assessing and improving the stability of chemometric models in small sample size situations Anal Bioanal Chem, 2008, 390, 1261-1271.
DOI: 10.1007/s00216-007-1818-6
I would really appreciate if my query can be clarified as I need to incorporate it in my research article accordingly.Thanks in advance.
Correct, if you use k-fold cv then McNemars is not appropriate.
You would use a modified paired student’s t-test.
We discuss exactly this in the above tutorial.
Thanks a lot for your reply. I would like to ask whether I can justify saying that, statistical significance test is not needed if repeated stratified k fold is used as the evaluation method as per the following research paper:
Beleites, C. & Salzer, R.: Assessing and improving the stability of chemometric models in small sample size situations Anal Bioanal Chem, 2008, 390, 1261-1271.
DOI: 10.1007/s00216-007-1818-6
You can make any claim you like with any support/evidence you like – it’s your research.
Whether your claims are justified based on the evidence is up to your reviewers.
Hi Jason,
I want to compare a deep artificial neural network model with some traditional machine learning algorithms such as logistic regression, support vector machine, k-nearest neighbor and decision tree based on a binary classification problem with a highly imbalanced dataset.
I have used some resampling methods such as SMOTE, ADASYN, ROS and SMOTE-ENN to handle the problem of the imbalanced dataset. I have used various evaluation methods
such as prediction Accuracy, Recall, Precision, and F1-score.
Furthermore, the Random hold-out and stratified 5-fold cross-validation methods are used as model validation techniques.
Which statistical test method can I use?
Thanks in advance!
The above tutorial address this exact question!
You must then choose.
Hi Jason,
In case of having an unbalanced dataset and running 10-fold cross-validation, can I use Wilcoxon signed ranked test to compare two models? although it violated the independence of samples’ constrain?
You can use anything you want on your project.
The above tutorial suggests using a modified student t-test when comparing results from k-fold cross validaiton.
Hi Jason,
First, let me congratulate and thank you for your amazing posts.
I read this post and the https://machinelearningmastery.com/hypothesis-test-for-comparing-machine-learning-algorithms/, as well as several articles on the topic. However, I still confused about the procedure that I’m using.
I have a medical dataset and 12 models. The models were evaluated using 5 repeated 2cv, can I compare the skills of the models using friedman tests(and posthocs), instead of performing 66 paired modified Student’s t-Test (paired_ttest_5x2cv)? I guess that the friedman test could be optimistically biased (as each evaluation of the model is not independent), or perhaps I got it wrong…
The ideal would be to draw several train/test sets from may dataset to ensure independence in each evaluation of the model, but I have limited samples. Also, I have several samples from the same patients, does it poses additional constraints?
Thanks in advance, best
Thanks.
If you used the 5x2cv procedure, you can use a modified version of the pair-wise student t-test as described above.
Perhaps I don’t understand the problem?
Dear Jason,
Thanks! That’s what I supposed… But in this case, I’ll need to perform 66 paired_ttest_5x2cv… well, it could be more! 🙂
Or is there any option to perform a test comparing several models, instead of one-to-one?
Best regards,
Yes.
Having many results to compare may cause a rethink of your model selection process.
Dear Jason, Thanks.
Just to be sure, the Friedman tests are out of the question, right?
As an alternative to the 66 paired_ttest_5x2cv, I was thinking of presenting the average of the models +/- the standard error.
Is it a viable option or do you recommend another?
Thanks,
No, you can use any test you want, the tutorial above are just general suggestions. If you’re unsure, perhaps speak to a statistician.
Hi Jason
Thanks for your help and recomendations!
You’re welcome.
Hi Jason,
what a great article! I was wondering whether you know of the Quade’s test and in what situations you would prefer it to McNemar’s test?
Best,
Clara
Sorry, I do not know that test.
Hi Jason,
Thank you for your informative article, really learnt a lot.
I am comparing two models, but the models were trained on different datasets, but tested on the same data. Can I still use a Nonparametric Paired Test to assess them?
In general, if the models were trained on different datasets, they cannot be compared directly.
Thank you for your article. It’s direct to the point and easy to grasp.
Same observations with some researchers in the field of deep learning, most papers are not using statistical analyses but just direct comparison of accuracy, F1-score, training/prediction time, and top-1 error. Hence, that is also what I did (1 to 2 train/validate/test runs only for each model & for each parameter setting and compare their performance metrics straightforwardly) until a reviewer told me to perform a significance test. And here I am.
My dataset is only 4000 images of high dimensions (more than 3 channels) and they are unbalanced because they are real data.
I performed hyperparameter setting experiments (7 settings, 1 run each) on one keras deep learning model and chose one best setting to be used to the other keras models that I will be comparing with my proposed model.
(1) In this set-up, if my understanding is correct, I can use the 5×2 CV paired t-test (http://rasbt.github.io/mlxtend/user_guide/evaluate/paired_ttest_5x2cv/) to all combinations of parameter settings, right?
(2) How about in choosing the best parameter/model, is there a post hoc test that can be employed after the 5×2 CV paired t-test? According to this presentation (https://ecs.wgtn.ac.nz/foswiki/pub/Groups/ECRG/StatsGuide/Significance%20Testing%20for%20Classification.pdf) in slide 35 (Summary), it seems that post hoc test is not necessary.
Any help is much appreciated.
Best regards.
(1) seems correct. (2) The t-test tells whether your null hypothesis model is different from your alternative model. So you already confirmed whether there is a “better” model after the test. What is the purpose of post-hoc test? I am not exactly sure it is not necessary but if you can justify the reason, then it might be useful.
Hello, thanks for your wonderful post! I have a new question about the relationship of statistical test and model performance metrics calculation. For instance, I have two algorithm and a dataset. And I conduct the McNemar test and the 5x2CV to compare whether the difference between the two algorithms is statistical significant. Then, I want to report the results. If the McNemar test shows that there is significant difference, can I calculate the performance metrics with the 5-fold cv of each algorithm, and report the result by combining both McNemar test (to provide the significant difference p value) and the performance metrics (with the 5-fold cv) together? In other words, the statistical compare is separate from the model performance metric calculation? Thank you in advance.
I think it is up to your interpretation on how to related the two metrics. Statistical test not always consistent with each other and that’s where the user have to explain or reconcile the result.
Thanks for your reply. In addition, I would like to ask a more specific question. Can I first conduct the statistical comparisons using McNemar test or 5x2CV, then calculate the performance metrics (e.g., accuracy) with the 5-fold cross validation? I think the 5-fold CV can get a good estimate of the model performance as it considers the data partition, compared with McNemar test. Or do I have to calculate the performance metrics with the exact same method as the statistical test (i.e., use the accuracy result from the McNemar test)? Thanks a lot!
I think to be fair, calculate the performance metrics with the same method is better.
Hi,
thank you for all your work. I have a question about a common setup in deep learning:
An unsupervised algorithm is trained with (almost) unlimited amounts of data, but only a tiny labeled set exists for evaluation purposes. I can split the train set in n disjoint sets without problems (apart from the long training times, so n should probably not be much larger than 10) but splitting the test set is impossible. Of course, the choice of test set matters but if I understand this correctly, all described testing procedures would assume that I can split the test set in either independent or slightly overlapping (Bengio’s approach to cross validation) sets.
Do you have any resources for this special case? Any statistical tests that are applicable in this case? Thank you!
You may want to check out semi-supervised learning: https://machinelearningmastery.com/semi-supervised-learning-with-label-propagation/
I was meaning more in regards to statistical testing. It’s complicated to assess whether one algorithm is better than another, when there is very limited labeled data available only during evaluation. I can not split my test data, therefore variation of the test data can not be considered as source of randomness. I was asking myself whether any of the tests above is then still “allowed” or if there are other tests for this case.
Hi GF…You may want to investigate cross-fold validation:
https://machinelearningmastery.com/repeated-k-fold-cross-validation-with-python/
Thanks for the article.
Just a clarifyication, in the 5×2 paired t-test, are we only training 5 times and validating 5 times (once on each split)? Or is it for each of the 5 repeats, we get 2 results (switching the test and train splits)? If the latter (i.e. training 10 total times), wouldn’t the pairs be dependent?
Hi Max…The following should help add clarity:
https://towardsdatascience.com/statistical-tests-for-comparing-machine-learning-and-baseline-performance-4dfc9402e46f
Thanks for the great article!
Here is what I’m trying to compare: datasets using the same algorithm
I have datasetA and datasetB, I have a KPI (let’s call it accuracy), and a model
I want to know if the model performs better on datasetB vs datasetA
both datasets are rather small and cannot be combined because they use different units of measure. I can always train the model on a portion of each dataset and test it on the remaining portion.
In this case
– my datasets are independent of each other
– evaluating the model on each dataset produces a response, and I can produce several responses through resampling
So I have independent samples made of responses that are not independent
What test best fits this?
Hi Brian…This is a challenging question to answer without actually seeing the data, however the following may be of interest:
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3116565/
Thanks for the great article!
Can the statistical test described above be used to compare the LSTM model for the time series case?
I have 3 LSTM models. Model 1 was trained using variables x1, model 2 was trained using variables x1 and x2, and model 3 was trained using variables x1, x2, and x3. The three models also have different hyperparameters. However, all three models predict the same Y value. The three models are also applied 10 folds walk forward validation. My goal was to see if adding a variable would increase the accuracy (lower RMSE) of the prediction. What statistical test is appropriate to use?
Hi Eko…You may find the following helpful. It presents some level of theory, however it also contains sample code for comparing multiple models.
https://par.nsf.gov/servlets/purl/10186554
Thank you for the response and the reference paper.
I’ve read it. In this paper, they use the same feature, namely “Adjusted Close” and then compare the ARIMA, BiLSTM and LSTM models. But in my case, I use different features for each model. To be clear, LSTM1 uses “Adjusted Close”, LSTM2 uses “Adjusted Close” and “Open”, and LSTM3 uses “Adjusted Close”, “Open” and “High”. LSTM1, LSTM2, and LSTM3 have different hyperparameter. Does it make sense for me to use the method in the paper to compare my three models?
Hi Jason, thank you so much for the great article, it’s really helpful!
A question of clarification: As in Dietterich 1998, your article warns against “10-fold cross-validation […] with an unmodified paired Student t-test”. I get that. It also says “Nadeau and Bengio further correction to the test statistic may be used with the 5×2-fold cross validation…”.
So, what about using Nadeau and Bengio’s corrected paired t-test in combination with (unrepeated) k-fold cv? Or is the corrected t-test still only recommended in combination with the 5x2cv set up?
I have been doing 5-fold cv on a fairly small dataset. If possible I would like to avoid using 5x2cv because then I’d be only training on 50% of my already-small dataset. Can I reasonably use Nadeau and Bengin’s corrected paired t-test with 5-fold cv?
Thank you so much again!
Hi Nelly…I see no reason why your idea could not be implemented. Try it and let us know what you find!
Ok, thanks for replying!
Hi I have a very large dataset for a deep learning medical image analysis project that takes a long time to train. I am testing out different combinations of architectures, pre-processing methods etc. Currently I use 5 fold cross validation. However, I recently ran a model 5 times on the same train/test data and it gave me varying results (due to inherent stochasticity of the models I think). I was wondering, is the rationale behind 5 repeats of 2-fold cross-validation to account for the stochasticity of the models ? In most other cases I’ve only heard of N fold CV and not on 5 repeats of the same.
Hi AV…the following may be of interest:
https://machinelearningmastery.com/k-fold-cross-validation/
Hi! I am working on a ML problem for which I am trying to evaluate if a particular classification algorithm is significantly better than a random classifier. I have both a binary classification model and a multi-class classification model. I am unable to use CV since my dataset is already divided into a train and test set. For the binary class model, I am guessing from this article that the McNemar test would be best! For the multi-class classification algorithm, how would you recommend I test whether the classification algorithm is significantly better than a random classifier? Thank you so much for this article and your help! 🙂
Hi Neha…The following may be of interest to you:
https://towardsdatascience.com/statistical-tests-for-comparing-classification-algorithms-ac1804e79bb7
Nice article. Suppose I have two stochastic models, A and B, and I run each N times with different seeds. What is the appropriate statistical test in this situation?
Hi Silvia…I would highly recommend the following resource for comparing such models:
https://machinelearningmastery.com/hypothesis-test-for-comparing-machine-learning-algorithms/
The models are run on the exact same dataset in each run.
Thank you for the feedback Silvia! Let us know if you have any additional questions.
Hi Jason – thanks for a great site. This tutorial is a couple of years old now, but hope you look at the questions once in a while. My situation is as follows:
I am investigating a classification problem to predict late payment of customer invoices, and regenerate a slightly larger data set every week including old data plus any new data that was acquired in the interrim. After train-test-split, I train 5 algorithms – Random Forest, Gradient Boost, Light Gradient Boost, XGBoost and Catboost. All of them score between 78 and 81% accuracy and I have tended to take the average prediction on unseen data (new invoices that are not yet late and for which we don’t know whether they will be paid on-time or late). However, when I apply the trained models to predict whether the new, not-late invoices will be paid late, I’ve noticed that the 4 boosted models predict nearly 100% to be late, while RF predicts a much closer mix of positive/negative results to the mix in the known data – roughly 59%. Am I making a mistake by taking the average prediction vs taking the prediction of the highest accuracy model (usually Catboost) or RF which appears to “fit” the proportion of late payments in the general data sets or making a leap of faith and accepting the boosted models’ predictions that almost all invoices in the most recent sets will be paid late? Any suggestions on how to resolve this?
Hi Jim…Please narrow your query to one question so that we may better assist you.
Hi Jason,
Thanks a lot for the tutorial,
I have the following question, what if we want to compare the performance of the same model on two datasets? McNemar’s test was my choice since I’m training a deep learning model ( train/ split approach, evaluated once ) however, from what I understand now, it does not measure the variability caused by the choice of the training set.
Hi Sara…You may find the following resource of interest:
https://neptune.ai/blog/how-to-compare-machine-learning-models-and-algorithms
hi, your tutorial is great, but I have a doubt regarding the statistical significance in this scenario — if I do train data on Lang0 language and generate a model. afterward using the Lang0 model I do testing on other languages like Lang1, Lan2…Lang5 used different algorithms like AlgoA, AlgoB, and AlgoC and got the accuracy. so in that case is it possible to do the statistical significance test? no cross-validation is done while training.
Say I have
Lang Algo1 Algo2 Algo3 Algo4 Algo5
Lang1 80 32 95 93 96.67
Lang2 88 11 98 97 92.51
Lang3 49 12 76 80 72.75
Lang4 81 2 95 94 77.7
Lang5 81 43 95 96 94.95
Hi! Thank you for the great post!
Just a question.. I am assuming the modified paired t-test would have the same assumptions as the paired t-test. Is there a non-parametric equivalent for the modified version for comparing models?
Thanks in advance!
Hi Eve…You may find the following resource of interest:
https://machinelearningmastery.com/parametric-and-nonparametric-machine-learning-algorithms/
Hi, thank you for your so quick reply! I mean if the difference of the pairs of errors between models is not normally distributed, then we would need to use a Wilcoxon signed rank test (as the non-parametric equivalent of a paired t-test). However, the Wilcoxon signed rank test again assumes independence of observations. How would we compare the models in this case?
Hi! Thank you for the great post and guidance. I also have a question about using a significance test to compare two classifiers. In my case, I have ten different datasets, and for each dataset, I trained each classifier three times.
If we do the significance test for each dataset, I am afraid the observations might be insufficient, since there are only three pairs of accuracy values. I would appreciate your opinion on this.
Besides that, I am also thinking if it is reasonable to conduct the test between two classifiers over all these ten datasets. So I get the averaged accuracy for each classifier for each dataset, and in the end, we have ten pairs of accuracy values. I am considering to do the wilcoxon signed-rank test. I would like to know if it sounds feasible for you?
Thanks in advance!
Hi Nico…You are very welcome! The following resource may provide some insight:
https://machinelearningmastery.com/compare-machine-learning-algorithms-python-scikit-learn/
Hi! Thank you for the great post and guidance.
I conducted an experiment with 10 stacked ensemble models for classification, evaluating them using 7 performance metrics (accuracy, precision, recall, f1-score, exact match ratio, hamming loss, and the zero-one-loss). The proposed model showed the best performance. To determine the significance of this result, I’ve been advised to use statistical significance measures like a t-test. Should I use a t-test for this comparison, or are there alternative methods better suited to assess the performance significance among these models?
Specifically, I want to answer this question “In the results section, in comparing the different models, you are to adopt statistical significance measures such as a t-test to ensure performance significance.”