Often, the input features for a predictive modeling task interact in unexpected and often nonlinear ways.
These interactions can be identified and modeled by a learning algorithm. Another approach is to engineer new features that expose these interactions and see if they improve model performance. Additionally, transforms like raising input variables to a power can help to better expose the important relationships between input variables and the target variable.
These features are called interaction and polynomial features and allow the use of simpler modeling algorithms as some of the complexity of interpreting the input variables and their relationships is pushed back to the data preparation stage. Sometimes these features can result in improved modeling performance, although at the cost of adding thousands or even millions of additional input variables.
In this tutorial, you will discover how to use polynomial feature transforms for feature engineering with numerical input variables.
After completing this tutorial, you will know:
- Some machine learning algorithms prefer or perform better with polynomial input features.
- How to use the polynomial features transform to create new versions of input variables for predictive modeling.
- How the degree of the polynomial impacts the number of input features created by the transform.
Kick-start your project with my new book Data Preparation for Machine Learning, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.

How to Use Polynomial Feature Transforms for Machine Learning
Photo by D Coetzee, some rights reserved.
Tutorial Overview
This tutorial is divided into five parts; they are:
- Polynomial Features
- Polynomial Feature Transform
- Sonar Dataset
- Polynomial Feature Transform Example
- Effect of Polynomial Degree
Polynomial Features
Polynomial features are those features created by raising existing features to an exponent.
For example, if a dataset had one input feature X, then a polynomial feature would be the addition of a new feature (column) where values were calculated by squaring the values in X, e.g. X^2. This process can be repeated for each input variable in the dataset, creating a transformed version of each.
As such, polynomial features are a type of feature engineering, e.g. the creation of new input features based on the existing features.
The “degree” of the polynomial is used to control the number of features added, e.g. a degree of 3 will add two new variables for each input variable. Typically a small degree is used such as 2 or 3.
Generally speaking, it is unusual to use d greater than 3 or 4 because for large values of d, the polynomial curve can become overly flexible and can take on some very strange shapes.
— Page 266, An Introduction to Statistical Learning with Applications in R, 2014.
It is also common to add new variables that represent the interaction between features, e.g a new column that represents one variable multiplied by another. This too can be repeated for each input variable creating a new “interaction” variable for each pair of input variables.
A squared or cubed version of an input variable will change the probability distribution, separating the small and large values, a separation that is increased with the size of the exponent.
This separation can help some machine learning algorithms make better predictions and is common for regression predictive modeling tasks and generally tasks that have numerical input variables.
Typically linear algorithms, such as linear regression and logistic regression, respond well to the use of polynomial input variables.
Linear regression is linear in the model parameters and adding polynomial terms to the model can be an effective way of allowing the model to identify nonlinear patterns.
— Page 11, Feature Engineering and Selection, 2019.
For example, when used as input to a linear regression algorithm, the method is more broadly referred to as polynomial regression.
Polynomial regression extends the linear model by adding extra predictors, obtained by raising each of the original predictors to a power. For example, a cubic regression uses three variables, X, X2, and X3, as predictors. This approach provides a simple way to provide a non-linear fit to data.
— Page 265, An Introduction to Statistical Learning with Applications in R, 2014.
Want to Get Started With Data Preparation?
Take my free 7-day email crash course now (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
Polynomial Feature Transform
The polynomial features transform is available in the scikit-learn Python machine learning library via the PolynomialFeatures class.
The features created include:
- The bias (the value of 1.0)
- Values raised to a power for each degree (e.g. x^1, x^2, x^3, …)
- Interactions between all pairs of features (e.g. x1 * x2, x1 * x3, …)
For example, with two input variables with values 2 and 3 and a degree of 2, the features created would be:
- 1 (the bias)
- 2^1 = 2
- 3^1 = 3
- 2^2 = 4
- 3^2 = 9
- 2 * 3 = 6
We can demonstrate this with an example:
|
1 2 3 4 5 6 7 8 9 10 |
# demonstrate the types of features created from numpy import asarray from sklearn.preprocessing import PolynomialFeatures # define the dataset data = asarray([[2,3],[2,3],[2,3]]) print(data) # perform a polynomial features transform of the dataset trans = PolynomialFeatures(degree=2) data = trans.fit_transform(data) print(data) |
Running the example first reports the raw data with two features (columns) and each feature has the same value, either 2 or 3.
Then the polynomial features are created, resulting in six features, matching what was described above.
|
1 2 3 4 5 6 7 |
[[2 3] [2 3] [2 3]] [[1. 2. 3. 4. 6. 9.] [1. 2. 3. 4. 6. 9.] [1. 2. 3. 4. 6. 9.]] |
The “degree” argument controls the number of features created and defaults to 2.
The “interaction_only” argument means that only the raw values (degree 1) and the interaction (pairs of values multiplied with each other) are included, defaulting to False.
The “include_bias” argument defaults to True to include the bias feature.
We will take a closer look at how to use the polynomial feature transforms on a real dataset.
First, let’s introduce a real dataset.
Sonar Dataset
The sonar dataset is a standard machine learning dataset for binary classification.
It involves 60 real-valued inputs and a two-class target variable. There are 208 examples in the dataset and the classes are reasonably balanced.
A baseline classification algorithm can achieve a classification accuracy of about 53.4 percent using repeated stratified 10-fold cross-validation. Top performance on this dataset is about 88 percent using repeated stratified 10-fold cross-validation.
The dataset describes radar returns of rocks or simulated mines.
You can learn more about the dataset from here:
No need to download the dataset; we will download it automatically from our worked examples.
First, let’s load and summarize the dataset. The complete example is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 |
# load and summarize the sonar dataset from pandas import read_csv from pandas.plotting import scatter_matrix from matplotlib import pyplot # Load dataset url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/sonar.csv" dataset = read_csv(url, header=None) # summarize the shape of the dataset print(dataset.shape) # summarize each variable print(dataset.describe()) # histograms of the variables dataset.hist() pyplot.show() |
Running the example first summarizes the shape of the loaded dataset.
This confirms the 60 input variables, one output variable, and 208 rows of data.
A statistical summary of the input variables is provided showing that values are numeric and range approximately from 0 to 1.
|
1 2 3 4 5 6 7 8 9 10 11 12 |
(208, 61) 0 1 2 ... 57 58 59 count 208.000000 208.000000 208.000000 ... 208.000000 208.000000 208.000000 mean 0.029164 0.038437 0.043832 ... 0.007949 0.007941 0.006507 std 0.022991 0.032960 0.038428 ... 0.006470 0.006181 0.005031 min 0.001500 0.000600 0.001500 ... 0.000300 0.000100 0.000600 25% 0.013350 0.016450 0.018950 ... 0.003600 0.003675 0.003100 50% 0.022800 0.030800 0.034300 ... 0.005800 0.006400 0.005300 75% 0.035550 0.047950 0.057950 ... 0.010350 0.010325 0.008525 max 0.137100 0.233900 0.305900 ... 0.044000 0.036400 0.043900 [8 rows x 60 columns] |
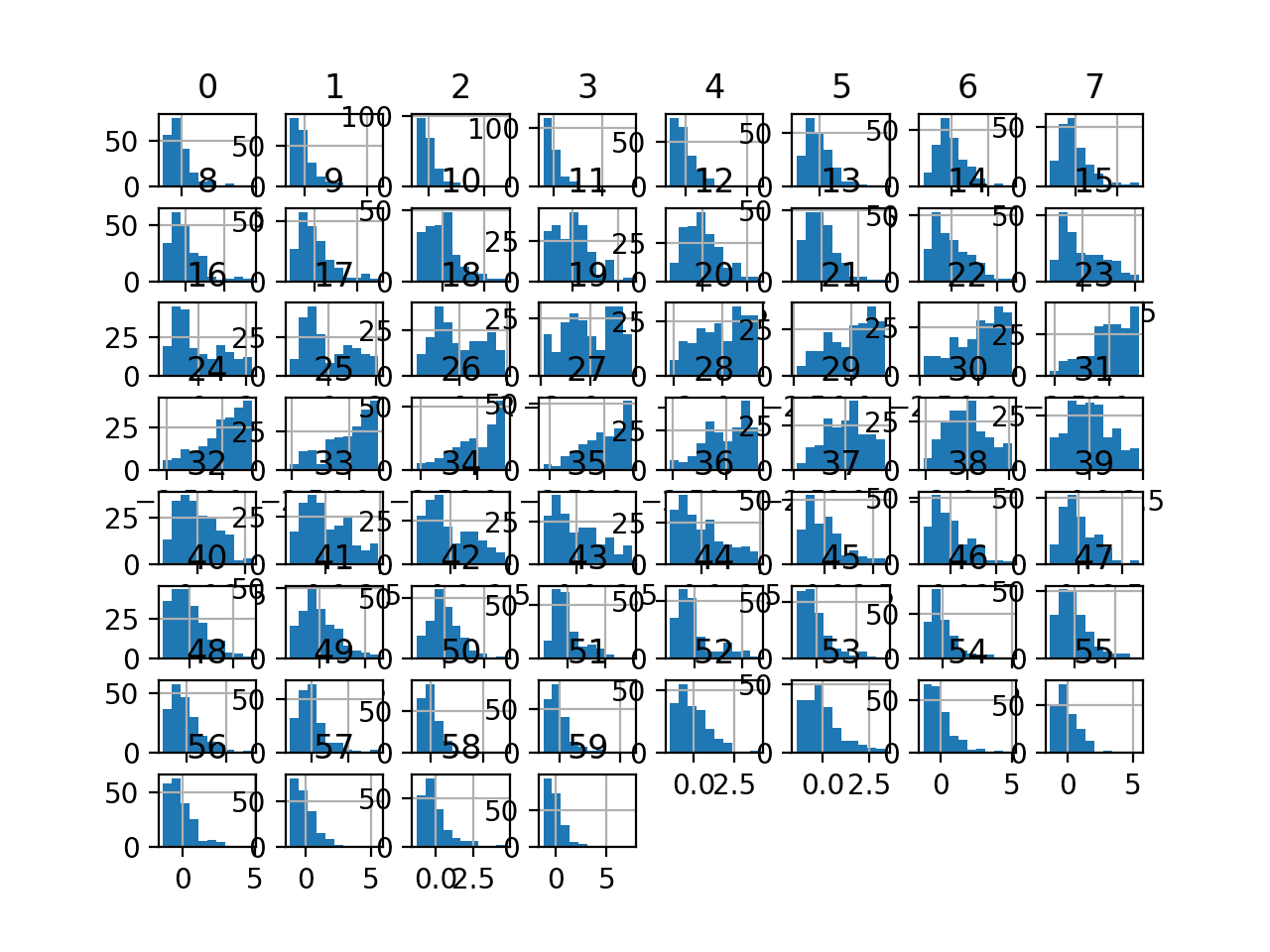
Finally, a histogram is created for each input variable.
If we ignore the clutter of the plots and focus on the histograms themselves, we can see that many variables have a skewed distribution.
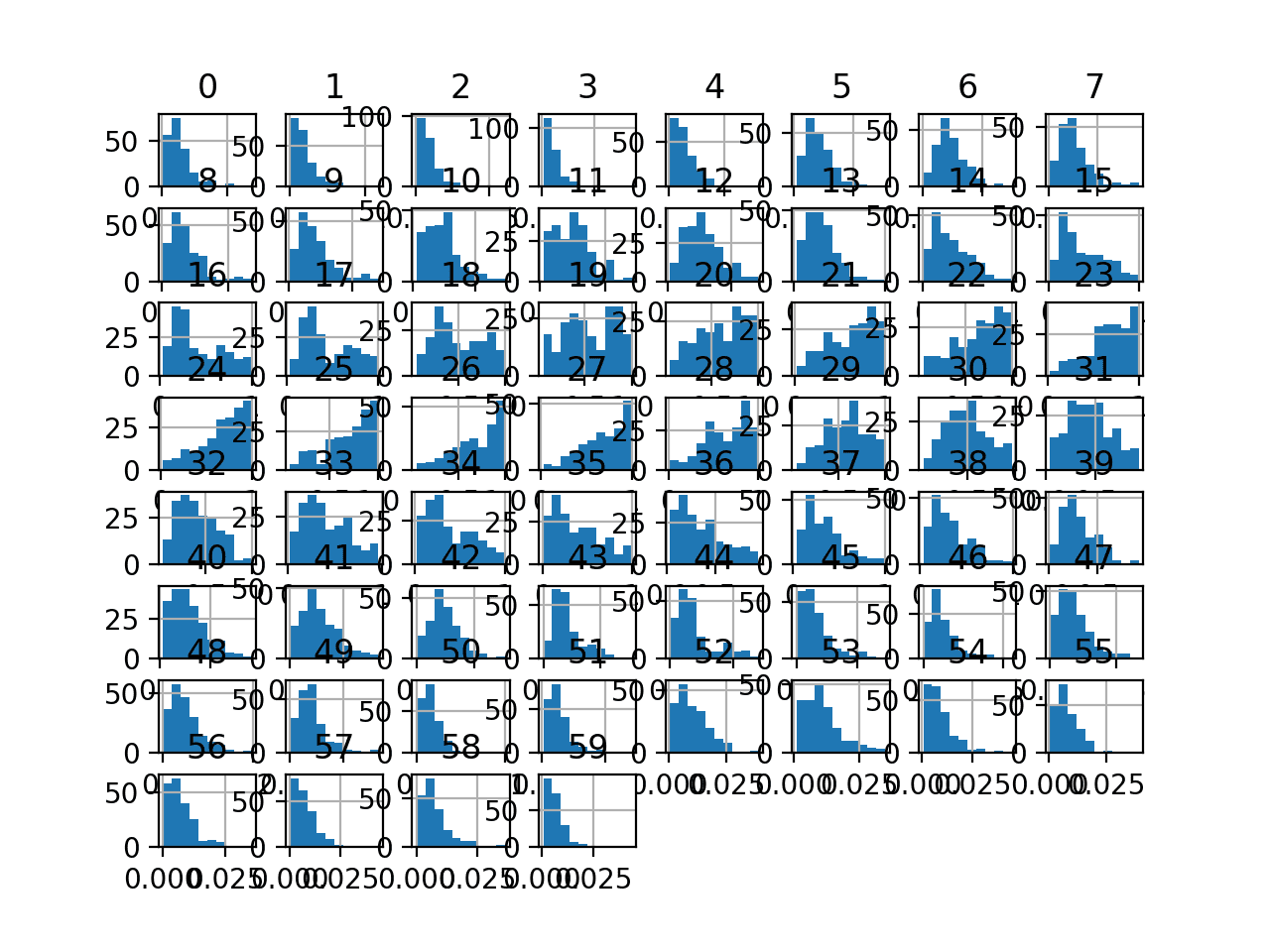
Histogram Plots of Input Variables for the Sonar Binary Classification Dataset
Next, let’s fit and evaluate a machine learning model on the raw dataset.
We will use a k-nearest neighbor algorithm with default hyperparameters and evaluate it using repeated stratified k-fold cross-validation. The complete example is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 |
# evaluate knn on the raw sonar dataset from numpy import mean from numpy import std from pandas import read_csv from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedStratifiedKFold from sklearn.neighbors import KNeighborsClassifier from sklearn.preprocessing import LabelEncoder from matplotlib import pyplot # load dataset url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/sonar.csv" dataset = read_csv(url, header=None) data = dataset.values # separate into input and output columns X, y = data[:, :-1], data[:, -1] # ensure inputs are floats and output is an integer label X = X.astype('float32') y = LabelEncoder().fit_transform(y.astype('str')) # define and configure the model model = KNeighborsClassifier() # evaluate the model cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1) n_scores = cross_val_score(model, X, y, scoring='accuracy', cv=cv, n_jobs=-1, error_score='raise') # report model performance print('Accuracy: %.3f (%.3f)' % (mean(n_scores), std(n_scores))) |
Running the example evaluates a KNN model on the raw sonar dataset.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
We can see that the model achieved a mean classification accuracy of about 79.7 percent, showing that it has skill (better than 53.4 percent) and is in the ball-park of good performance (88 percent).
|
1 |
Accuracy: 0.797 (0.073) |
Next, let’s explore a polynomial features transform of the dataset.
Polynomial Feature Transform Example
We can apply the polynomial features transform to the Sonar dataset directly.
In this case, we will use a degree of 3.
|
1 2 3 4 |
... # perform a polynomial features transform of the dataset trans = PolynomialFeatures(degree=3) data = trans.fit_transform(data) |
Let’s try it on our sonar dataset.
The complete example of creating a polynomial features transform of the sonar dataset and summarizing the created features is below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 |
# visualize a polynomial features transform of the sonar dataset from pandas import read_csv from pandas import DataFrame from pandas.plotting import scatter_matrix from sklearn.preprocessing import PolynomialFeatures from matplotlib import pyplot # load dataset url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/sonar.csv" dataset = read_csv(url, header=None) # retrieve just the numeric input values data = dataset.values[:, :-1] # perform a polynomial features transform of the dataset trans = PolynomialFeatures(degree=3) data = trans.fit_transform(data) # convert the array back to a dataframe dataset = DataFrame(data) # summarize print(dataset.shape) |
Running the example performs the polynomial features transform on the sonar dataset.
We can see that our features increased from 61 (60 input features) for the raw dataset to 39,711 features (39,710 input features).
|
1 |
(208, 39711) |
Next, let’s evaluate the same KNN model as the previous section, but in this case on a polynomial features transform of the dataset.
The complete example is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 |
# evaluate knn on the sonar dataset with polynomial features transform from numpy import mean from numpy import std from pandas import read_csv from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedStratifiedKFold from sklearn.neighbors import KNeighborsClassifier from sklearn.preprocessing import LabelEncoder from sklearn.preprocessing import PolynomialFeatures from sklearn.pipeline import Pipeline from matplotlib import pyplot # load dataset url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/sonar.csv" dataset = read_csv(url, header=None) data = dataset.values # separate into input and output columns X, y = data[:, :-1], data[:, -1] # ensure inputs are floats and output is an integer label X = X.astype('float32') y = LabelEncoder().fit_transform(y.astype('str')) # define the pipeline trans = PolynomialFeatures(degree=3) model = KNeighborsClassifier() pipeline = Pipeline(steps=[('t', trans), ('m', model)]) # evaluate the pipeline cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1) n_scores = cross_val_score(pipeline, X, y, scoring='accuracy', cv=cv, n_jobs=-1, error_score='raise') # report pipeline performance print('Accuracy: %.3f (%.3f)' % (mean(n_scores), std(n_scores))) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running the example, we can see that the polynomial features transform results in a lift in performance from 79.7 percent accuracy without the transform to about 80.0 percent with the transform.
|
1 |
Accuracy: 0.800 (0.077) |
Next, let’s explore the effect of different scaling ranges.
Effect of Polynomial Degree
The degree of the polynomial dramatically increases the number of input features.
To get an idea of how much this impacts the number of features, we can perform the transform with a range of different degrees and compare the number of features in the dataset.
The complete example is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 |
# compare the effect of the degree on the number of created features from pandas import read_csv from sklearn.preprocessing import LabelEncoder from sklearn.preprocessing import PolynomialFeatures from matplotlib import pyplot # get the dataset def get_dataset(): # load dataset url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/sonar.csv" dataset = read_csv(url, header=None) data = dataset.values # separate into input and output columns X, y = data[:, :-1], data[:, -1] # ensure inputs are floats and output is an integer label X = X.astype('float32') y = LabelEncoder().fit_transform(y.astype('str')) return X, y # define dataset X, y = get_dataset() # calculate change in number of features num_features = list() degress = [i for i in range(1, 6)] for d in degress: # create transform trans = PolynomialFeatures(degree=d) # fit and transform data = trans.fit_transform(X) # record number of features num_features.append(data.shape[1]) # summarize print('Degree: %d, Features: %d' % (d, data.shape[1])) # plot degree vs number of features pyplot.plot(degress, num_features) pyplot.show() |
Running the example first reports the degree from 1 to 5 and the number of features in the dataset.
We can see that a degree of 1 has no effect and that the number of features dramatically increases from 2 through to 5.
This highlights that for anything other than very small datasets, a degree of 2 or 3 should be used to avoid a dramatic increase in input variables.
|
1 2 3 4 5 |
Degree: 1, Features: 61 Degree: 2, Features: 1891 Degree: 3, Features: 39711 Degree: 4, Features: 635376 Degree: 5, Features: 8259888 |
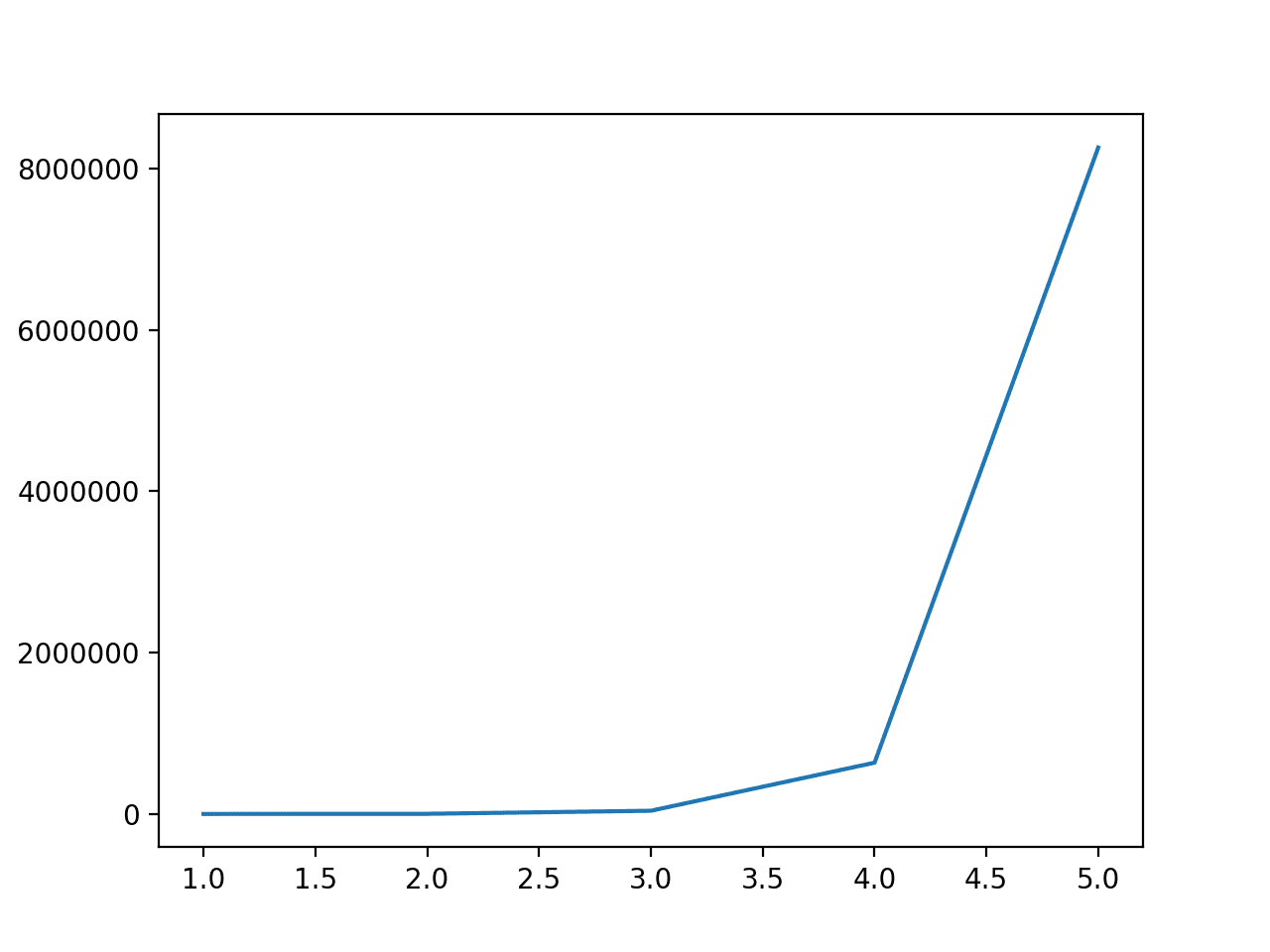
Line Plot of the Degree vs. the Number of Input Features for the Polynomial Feature Transform
More features may result in more overfitting, and in turn, worse results.
It may be a good idea to treat the degree for the polynomial features transform as a hyperparameter and test different values for your dataset.
The example below explores degree values from 1 to 4 and evaluates their effect on classification accuracy with the chosen model.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 |
# explore the effect of degree on accuracy for the polynomial features transform from numpy import mean from numpy import std from pandas import read_csv from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedStratifiedKFold from sklearn.neighbors import KNeighborsClassifier from sklearn.preprocessing import PolynomialFeatures from sklearn.preprocessing import LabelEncoder from sklearn.pipeline import Pipeline from matplotlib import pyplot # get the dataset def get_dataset(): # load dataset url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/sonar.csv" dataset = read_csv(url, header=None) data = dataset.values # separate into input and output columns X, y = data[:, :-1], data[:, -1] # ensure inputs are floats and output is an integer label X = X.astype('float32') y = LabelEncoder().fit_transform(y.astype('str')) return X, y # get a list of models to evaluate def get_models(): models = dict() for d in range(1,5): # define the pipeline trans = PolynomialFeatures(degree=d) model = KNeighborsClassifier() models[str(d)] = Pipeline(steps=[('t', trans), ('m', model)]) return models # evaluate a give model using cross-validation def evaluate_model(model, X, y): cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1) scores = cross_val_score(model, X, y, scoring='accuracy', cv=cv, n_jobs=-1, error_score='raise') return scores # define dataset X, y = get_dataset() # get the models to evaluate models = get_models() # evaluate the models and store results results, names = list(), list() for name, model in models.items(): scores = evaluate_model(model, X, y) results.append(scores) names.append(name) print('>%s %.3f (%.3f)' % (name, mean(scores), std(scores))) # plot model performance for comparison pyplot.boxplot(results, labels=names, showmeans=True) pyplot.show() |
Running the example reports the mean classification accuracy for each polynomial degree.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see that performance is generally worse than no transform (degree 1) except for a degree 3.
It might be interesting to explore scaling the data before or after performing the transform to see how it impacts model performance.
|
1 2 3 4 |
>1 0.797 (0.073) >2 0.793 (0.085) >3 0.800 (0.077) >4 0.795 (0.079) |
Box and whisker plots are created to summarize the classification accuracy scores for each polynomial degree.
We can see that performance remains flat, perhaps with the first signs of overfitting with a degree of 4.
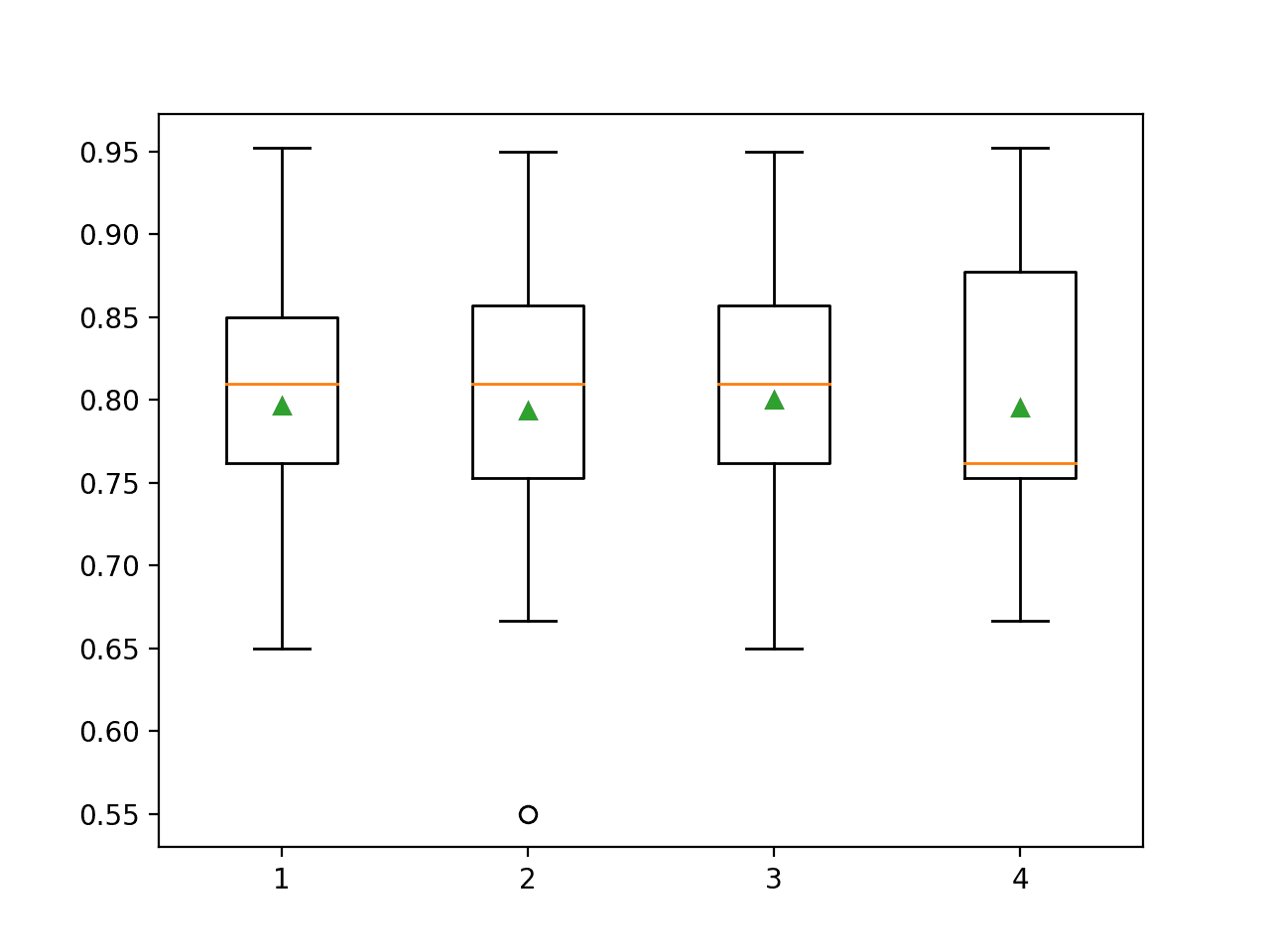
Box Plots of Degree for the Polynomial Feature Transform vs. Classification Accuracy of KNN on the Sonar Dataset
Further Reading
This section provides more resources on the topic if you are looking to go deeper.
Books
- An Introduction to Statistical Learning with Applications in R, 2014.
- Feature Engineering and Selection, 2019.
APIs
Articles
Summary
In this tutorial, you discovered how to use polynomial feature transforms for feature engineering with numerical input variables.
Specifically, you learned:
- Some machine learning algorithms prefer or perform better with polynomial input features.
- How to use the polynomial features transform to create new versions of input variables for predictive modeling.
- How the degree of the polynomial impacts the number of input features created by the transform.
Do you have any questions?
Ask your questions in the comments below and I will do my best to answer.


Hello, again 🙂
I am using feedforward neural network as a regression by using (5 input features/ sensors readings).
I want to try Polynomial Feature Transform for this project to create more input features
My question is:
Can I do (RobustScaler) or (StandardScaler) after doing PolynomialFeatures?
Thank you very much
Yes.
One approach would be to apply the transforms as part of a pipeline to your dataset, then provide the data to the model.
This case study is not convincing regarding the benefits of polynomial models.
You started with 200 datapoints and 60 variables. After adding 1000+ variables you raised model accuracy by 0.3%.
1: This level of increase can be achieved by chance, i.e. just retraining the original model.
2: There is no holdout set cv. Realistically you’d have 150 observations to model with and most models would overfit without feature selection.
3: It demonstrates well how OO nature of python modules is cumbersome for statistical workflows. The same can be done in R using half the LOC and twice the readability.
Yes, I could make a better case. Thanks for the feedback.
Hello Jason,
Is it possible to perform polynomial regression with Keras?
Do you have any easy example?
Thanks
Marco
Yes, you can generate the features then pass them to your keras model.
No, I don’t have an example but it should be straightforward.
Thanks for your sharing,
If we want to make predictions, should we also implement the Polynomial Feature Transform for the test data?
Thanks again
Yes, the same transform object must be used for the train and test datasets. E.g. it is fit on the training set and applied to the train and test sets.
Hello Jason,
your blog posts are incredibly interesting for a practitioner like me! This time I have a question: does the polynomial feature transform not introduce multicollinearity (e.g. correlation between x and x^2), which in turn could lead to model performance degradation?
Many thanks
Thanks.
It may which could impact some linear models.
Dear Dr Jason,
I have to say that I didn’t understand the patterns produced by the transforms of PolynomialFeatures.
I did the following with interactions and polynomials and interactions only using the array [1,2,3,4]
With the assistance of the 2nd answer of https://stackoverflow.com/questions/51906274/cannot-understand-with-sklearns-polynomialfeatures you get the following pattern.
Then fit_transform(data) produces the form
If you exclude the the squares of a^2, b^2, c^2 and d^2.
Which is of the type:
What of degree 3 and 2 values, eg data = [a,b]
Number of transformed outputs = (n + d)!/(n! * d!) where n = number of numbers, d = degree of polynomial.
Eg number of numbers = 3, degree of polynomial = 2,
number of transformed outputs = (n + d)! / (n! * d!) = 5!/(3! * 2!) =10
Eg number of numbers = 2, degree of polynomial = 3
number of transformed outputs = 5!/(2!*3!) = 10
Thank you,
Anthony of Sydney
Thanks for sharing your findings.
Dear Dr Jason,
I was experimenting with the code under the subheading “Polynomial Feature Transform Example” which is also in Listing 23.10, page 310 (327 of 398) of your book.
The particular lines of interest are:
I ask the question “,,,why didn’t the author transform by
and feed that into the cross_val_score?
Note I did not use the pipeline since BUT I used the transformed polynomial features X2.
I still got the same result.
The question is: In the original code the pipeline seemed to have performed the PolynomialFeatures function of degree 3 without putting the transformed(X) = X2 into the cross_val_score function.
Put it another way, in the original code, how is that pipeline managed to calculate the transformed featurs without using the transformed features in the cross_val_score. Yet I got the same result.
Thank you,
Anthony of Sydney
The pipeline performs the transform to the input data, and the transformed data is then passed to the model.
You can learn more about how pipelines work here:
https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html
Dear Dr Jason,
“The pipeline performs the transform to the input data, and the transformed data is then passed to the model.”
The issue was clarified
Thank you.
Anthony of Sydney.
Happy to hear that.
Hi Jason,
Thank you for the article, it’s very informative. My question is in regards to the application of this technique. Should the conversion of features to a polynomial be used for regression problems or can they be used for classification problems as well?
Thank you,
Bill
Polynomial features can help on regression and classification tasks, perhaps try and compare to results of the same model without polynomial features.
Hi Jason,
I have a question. What is the correct order to make in the Pipeline: StandardScale and Polynomial Features or Polynomial Features and StandardScale? I’m confused about that.
My advice would be to test to see if it makes a difference.
My gut says in this case it probably does not matter.
Got it! Thank you so much! Your website is awesome!
You’re welcome.
Thanks!
Hi, I have only 1 independent feature which exhibit non linear relationship with the target, should I create polynomial features from that independent variable for tree based model?
Perhaps try it and compare results to other methods.
I created polynomial features upto degree 4 and they improved my linear regression model R2 score significantly (validated by Cross Validation). However my question is that the newly created polynomial features are highly correlated to the original feature from which they are generated with pearson correlation values above 0.80. Isn’t it lead to multicollinearity problem? Or is it acceptable to accept the results of the model?
It may be.
Focus on model performance. If the model performs better, then keep the new features.
Dear Jason thanks a lot for your advice. I have another question please. You mentioned with a book reference that “for large values of d (degree) the polynomial curves becomes overly flexible and can take very strange shapes”. My question is for large value of d say 5, if the linear model performance is increasing plus i do not see any abnormality in the regression line, can we keep those features to build the model?
Actually in my case i have 13 features, i find out the most important feature, created 5 degree polynomial features from that single most important feature, observed considerable improvement in model performance till d=5 , and it started reducing at d=6.. I hope you understand my question. thanks in advance.
Sure, use whatever works best for your data and test harness. Ensure your test harness is robust so that you don’t trick yourself.
Thanks a lot for your replies. Highly appreciate your awesome articles always.
Thanks.
Hi Jason, I have a question please. What if I created a polynomial feature having VIF=5.6, p-value=0.7 facilitated to improve the Adj. R2 by 2.5%. How do I can interpret such feature?
Thanks please.
Sorry, I try to avoid interpreting results for readers.
Perhaps compare results to other methods and use what works best or looks more reliable for your specific application.
Thanks for your quick reply. I appreciate it. Your comments are valuable for learners like me.
You’re welcome.
Your articles are really fantastic and full of details and explanations and examples. you are the Jedi Master! 😀
Thanks!
with 2 input variables with values 2,3,4 and a degree of 2. Which is the order output variables?
Maybe this?
1 (the bias)
2^1
3^1
4^1
2^2
3^2
4^2
2*3
3*4
Try to replace
data = asarray([[2,3,4],[2,3,4]])
and you will see it is:
1, 2^1, 3^1, 4^1, 2^2, 2*3, 2*4, 3^2, 3*4, 4^2
but Jason Brownlee wrote above:
“For example, with two input variables with values 2 and 3 and a degree of 2, the features created would be:
1 (the bias)
2^1 = 2
3^1 = 3
2^2 = 4
3^2 = 9
2 * 3 = 6”
So, it seems the interactions are the last.
You look at the output of the code, not the bullet points, you will see it was in the order [2,3,4,6,9] instead.
I have generated new features through polynomial degree =2, now should I replace/discard original features or should keep all features and their poly features for regression
Hi Salaf…The following may be of interest to you for understanding transformations of datasets in general:
https://machinelearningmastery.com/how-to-improve-neural-network-stability-and-modeling-performance-with-data-scaling/