Data Preparation for Machine Learning Crash Course.
Get on top of data preparation with Python in 7 days.
Data preparation involves transforming raw data into a form that is more appropriate for modeling.
Preparing data may be the most important part of a predictive modeling project and the most time-consuming, although it seems to be the least discussed. Instead, the focus is on machine learning algorithms, whose usage and parameterization has become quite routine.
Practical data preparation requires knowledge of data cleaning, feature selection data transforms, dimensionality reduction, and more.
In this crash course, you will discover how you can get started and confidently prepare data for a predictive modeling project with Python in seven days.
This is a big and important post. You might want to bookmark it.
Kick-start your project with my new book Data Preparation for Machine Learning, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.
- Updated Jun/2020: Changed the target for the horse colic dataset.

Data Preparation for Machine Learning (7-Day Mini-Course)
Photo by Christian Collins, some rights reserved.
Who Is This Crash-Course For?
Before we get started, let’s make sure you are in the right place.
This course is for developers who may know some applied machine learning. Maybe you know how to work through a predictive modeling problem end to end, or at least most of the main steps, with popular tools.
The lessons in this course do assume a few things about you, such as:
- You know your way around basic Python for programming.
- You may know some basic NumPy for array manipulation.
- You may know some basic scikit-learn for modeling.
You do NOT need to be:
- A math wiz!
- A machine learning expert!
This crash course will take you from a developer who knows a little machine learning to a developer who can effectively and competently prepare data for a predictive modeling project.
Note: This crash course assumes you have a working Python 3 SciPy environment with at least NumPy installed. If you need help with your environment, you can follow the step-by-step tutorial here:
Crash-Course Overview
This crash course is broken down into seven lessons.
You could complete one lesson per day (recommended) or complete all of the lessons in one day (hardcore). It really depends on the time you have available and your level of enthusiasm.
Below is a list of the seven lessons that will get you started and productive with data preparation in Python:
- Lesson 01: Importance of Data Preparation
- Lesson 02: Fill Missing Values With Imputation
- Lesson 03: Select Features With RFE
- Lesson 04: Scale Data With Normalization
- Lesson 05: Transform Categories With One-Hot Encoding
- Lesson 06: Transform Numbers to Categories With kBins
- Lesson 07: Dimensionality Reduction with PCA
Each lesson could take you 60 seconds or up to 30 minutes. Take your time and complete the lessons at your own pace. Ask questions and even post results in the comments below.
The lessons might expect you to go off and find out how to do things. I will give you hints, but part of the point of each lesson is to force you to learn where to go to look for help with and about the algorithms and the best-of-breed tools in Python. (Hint: I have all of the answers on this blog; use the search box.)
Post your results in the comments; I’ll cheer you on!
Hang in there; don’t give up.
Want to Get Started With Data Preparation?
Take my free 7-day email crash course now (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
Lesson 01: Importance of Data Preparation
In this lesson, you will discover the importance of data preparation in predictive modeling with machine learning.
Predictive modeling projects involve learning from data.
Data refers to examples or cases from the domain that characterize the problem you want to solve.
On a predictive modeling project, such as classification or regression, raw data typically cannot be used directly.
There are four main reasons why this is the case:
- Data Types: Machine learning algorithms require data to be numbers.
- Data Requirements: Some machine learning algorithms impose requirements on the data.
- Data Errors: Statistical noise and errors in the data may need to be corrected.
- Data Complexity: Complex nonlinear relationships may be teased out of the data.
The raw data must be pre-processed prior to being used to fit and evaluate a machine learning model. This step in a predictive modeling project is referred to as “data preparation.”
There are common or standard tasks that you may use or explore during the data preparation step in a machine learning project.
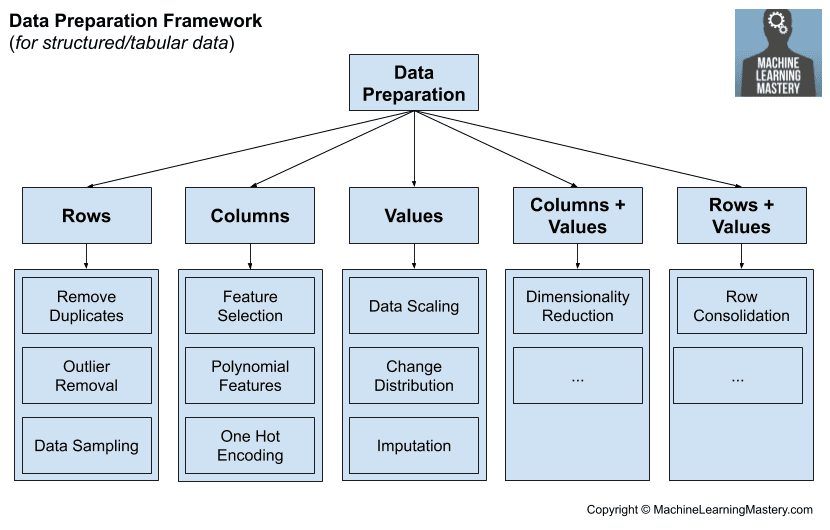
These tasks include:
- Data Cleaning: Identifying and correcting mistakes or errors in the data.
- Feature Selection: Identifying those input variables that are most relevant to the task.
- Data Transforms: Changing the scale or distribution of variables.
- Feature Engineering: Deriving new variables from available data.
- Dimensionality Reduction: Creating compact projections of the data.
Each of these tasks is a whole field of study with specialized algorithms.
Your Task
For this lesson, you must list three data preparation algorithms that you know of or may have used before and give a one-line summary for its purpose.
One example of a data preparation algorithm is data normalization that scales numerical variables to the range between zero and one.
Post your answer in the comments below. I would love to see what you come up with.
In the next lesson, you will discover how to fix data that has missing values, called data imputation.
Lesson 02: Fill Missing Values With Imputation
In this lesson, you will discover how to identify and fill missing values in data.
Real-world data often has missing values.
Data can have missing values for a number of reasons, such as observations that were not recorded and data corruption. Handling missing data is important as many machine learning algorithms do not support data with missing values.
Filling missing values with data is called data imputation and a popular approach for data imputation is to calculate a statistical value for each column (such as a mean) and replace all missing values for that column with the statistic.
The horse colic dataset describes medical characteristics of horses with colic and whether they lived or died. It has missing values marked with a question mark ‘?’. We can load the dataset with the read_csv() function and ensure that question mark values are marked as NaN.
Once loaded, we can use the SimpleImputer class to transform all missing values marked with a NaN value with the mean of the column.
The complete example is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 |
# statistical imputation transform for the horse colic dataset from numpy import isnan from pandas import read_csv from sklearn.impute import SimpleImputer # load dataset url = 'https://raw.githubusercontent.com/jbrownlee/Datasets/master/horse-colic.csv' dataframe = read_csv(url, header=None, na_values='?') # split into input and output elements data = dataframe.values ix = [i for i in range(data.shape[1]) if i != 23] X, y = data[:, ix], data[:, 23] # print total missing print('Missing: %d' % sum(isnan(X).flatten())) # define imputer imputer = SimpleImputer(strategy='mean') # fit on the dataset imputer.fit(X) # transform the dataset Xtrans = imputer.transform(X) # print total missing print('Missing: %d' % sum(isnan(Xtrans).flatten())) |
Your Task
For this lesson, you must run the example and review the number of missing values in the dataset before and after the data imputation transform.
Post your answer in the comments below. I would love to see what you come up with.
In the next lesson, you will discover how to select the most important features in a dataset.
Lesson 03: Select Features With RFE
In this lesson, you will discover how to select the most important features in a dataset.
Feature selection is the process of reducing the number of input variables when developing a predictive model.
It is desirable to reduce the number of input variables to both reduce the computational cost of modeling and, in some cases, to improve the performance of the model.
Recursive Feature Elimination, or RFE for short, is a popular feature selection algorithm.
RFE is popular because it is easy to configure and use and because it is effective at selecting those features (columns) in a training dataset that are more or most relevant in predicting the target variable.
The scikit-learn Python machine learning library provides an implementation of RFE for machine learning. RFE is a transform. To use it, first, the class is configured with the chosen algorithm specified via the “estimator” argument and the number of features to select via the “n_features_to_select” argument.
The example below defines a synthetic classification dataset with five redundant input features. RFE is then used to select five features using the decision tree algorithm.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 |
# report which features were selected by RFE from sklearn.datasets import make_classification from sklearn.feature_selection import RFE from sklearn.tree import DecisionTreeClassifier # define dataset X, y = make_classification(n_samples=1000, n_features=10, n_informative=5, n_redundant=5, random_state=1) # define RFE rfe = RFE(estimator=DecisionTreeClassifier(), n_features_to_select=5) # fit RFE rfe.fit(X, y) # summarize all features for i in range(X.shape[1]): print('Column: %d, Selected=%s, Rank: %d' % (i, rfe.support_[i], rfe.ranking_[i])) |
Your Task
For this lesson, you must run the example and review which features were selected and the relative ranking that each input feature was assigned.
Post your answer in the comments below. I would love to see what you come up with.
In the next lesson, you will discover how to scale numerical data.
Lesson 04: Scale Data With Normalization
In this lesson, you will discover how to scale numerical data for machine learning.
Many machine learning algorithms perform better when numerical input variables are scaled to a standard range.
This includes algorithms that use a weighted sum of the input, like linear regression, and algorithms that use distance measures, like k-nearest neighbors.
One of the most popular techniques for scaling numerical data prior to modeling is normalization. Normalization scales each input variable separately to the range 0-1, which is the range for floating-point values where we have the most precision. It requires that you know or are able to accurately estimate the minimum and maximum observable values for each variable. You may be able to estimate these values from your available data.
You can normalize your dataset using the scikit-learn object MinMaxScaler.
The example below defines a synthetic classification dataset, then uses the MinMaxScaler to normalize the input variables.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 |
# example of normalizing input data from sklearn.datasets import make_classification from sklearn.preprocessing import MinMaxScaler # define dataset X, y = make_classification(n_samples=1000, n_features=5, n_informative=5, n_redundant=0, random_state=1) # summarize data before the transform print(X[:3, :]) # define the scaler trans = MinMaxScaler() # transform the data X_norm = trans.fit_transform(X) # summarize data after the transform print(X_norm[:3, :]) |
Your Task
For this lesson, you must run the example and report the scale of the input variables both prior to and then after the normalization transform.
For bonus points, calculate the minimum and maximum of each variable before and after the transform to confirm it was applied as expected.
Post your answer in the comments below. I would love to see what you come up with.
In the next lesson, you will discover how to transform categorical variables to numbers.
Lesson 05: Transform Categories With One-Hot Encoding
In this lesson, you will discover how to encode categorical input variables as numbers.
Machine learning models require all input and output variables to be numeric. This means that if your data contains categorical data, you must encode it to numbers before you can fit and evaluate a model.
One of the most popular techniques for transforming categorical variables into numbers is the one-hot encoding.
Categorical data are variables that contain label values rather than numeric values.
Each label for a categorical variable can be mapped to a unique integer, called an ordinal encoding. Then, a one-hot encoding can be applied to the ordinal representation. This is where one new binary variable is added to the dataset for each unique integer value in the variable, and the original categorical variable is removed from the dataset.
For example, imagine we have a “color” variable with three categories (‘red‘, ‘green‘, and ‘blue‘). In this case, three binary variables are needed. A “1” value is placed in the binary variable for the color and “0” values for the other colors.
For example:
|
1 2 3 4 |
red, green, blue 1, 0, 0 0, 1, 0 0, 0, 1 |
This one-hot encoding transform is available in the scikit-learn Python machine learning library via the OneHotEncoder class.
The breast cancer dataset contains only categorical input variables.
The example below loads the dataset and one hot encodes each of the categorical input variables.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 |
# one-hot encode the breast cancer dataset from pandas import read_csv from sklearn.preprocessing import OneHotEncoder # define the location of the dataset url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/breast-cancer.csv" # load the dataset dataset = read_csv(url, header=None) # retrieve the array of data data = dataset.values # separate into input and output columns X = data[:, :-1].astype(str) y = data[:, -1].astype(str) # summarize the raw data print(X[:3, :]) # define the one hot encoding transform encoder = OneHotEncoder(sparse=False) # fit and apply the transform to the input data X_oe = encoder.fit_transform(X) # summarize the transformed data print(X_oe[:3, :]) |
Your Task
For this lesson, you must run the example and report on the raw data before the transform, and the impact on the data after the one-hot encoding was applied.
Post your answer in the comments below. I would love to see what you come up with.
In the next lesson, you will discover how to transform numerical variables into categories.
Lesson 06: Transform Numbers to Categories With kBins
In this lesson, you will discover how to transform numerical variables into categorical variables.
Some machine learning algorithms may prefer or require categorical or ordinal input variables, such as some decision tree and rule-based algorithms.
This could be caused by outliers in the data, multi-modal distributions, highly exponential distributions, and more.
Many machine learning algorithms prefer or perform better when numerical input variables with non-standard distributions are transformed to have a new distribution or an entirely new data type.
One approach is to use the transform of the numerical variable to have a discrete probability distribution where each numerical value is assigned a label and the labels have an ordered (ordinal) relationship.
This is called a discretization transform and can improve the performance of some machine learning models for datasets by making the probability distribution of numerical input variables discrete.
The discretization transform is available in the scikit-learn Python machine learning library via the KBinsDiscretizer class.
It allows you to specify the number of discrete bins to create (n_bins), whether the result of the transform will be an ordinal or one-hot encoding (encode), and the distribution used to divide up the values of the variable (strategy), such as ‘uniform.’
The example below creates a synthetic input variable with 10 numerical input variables, then encodes each into 10 discrete bins with an ordinal encoding.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 |
# discretize numeric input variables from sklearn.datasets import make_classification from sklearn.preprocessing import KBinsDiscretizer # define dataset X, y = make_classification(n_samples=1000, n_features=5, n_informative=5, n_redundant=0, random_state=1) # summarize data before the transform print(X[:3, :]) # define the transform trans = KBinsDiscretizer(n_bins=10, encode='ordinal', strategy='uniform') # transform the data X_discrete = trans.fit_transform(X) # summarize data after the transform print(X_discrete[:3, :]) |
Your Task
For this lesson, you must run the example and report on the raw data before the transform, and then the effect the transform had on the data.
For bonus points, explore alternate configurations of the transform, such as different strategies and number of bins.
Post your answer in the comments below. I would love to see what you come up with.
In the next lesson, you will discover how to reduce the dimensionality of input data.
Lesson 07: Dimensionality Reduction With PCA
In this lesson, you will discover how to use dimensionality reduction to reduce the number of input variables in a dataset.
The number of input variables or features for a dataset is referred to as its dimensionality.
Dimensionality reduction refers to techniques that reduce the number of input variables in a dataset.
More input features often make a predictive modeling task more challenging to model, more generally referred to as the curse of dimensionality.
Although on high-dimensionality statistics, dimensionality reduction techniques are often used for data visualization, these techniques can be used in applied machine learning to simplify a classification or regression dataset in order to better fit a predictive model.
Perhaps the most popular technique for dimensionality reduction in machine learning is Principal Component Analysis, or PCA for short. This is a technique that comes from the field of linear algebra and can be used as a data preparation technique to create a projection of a dataset prior to fitting a model.
The resulting dataset, the projection, can then be used as input to train a machine learning model.
The scikit-learn library provides the PCA class that can be fit on a dataset and used to transform a training dataset and any additional datasets in the future.
The example below creates a synthetic binary classification dataset with 10 input variables then uses PCA to reduce the dimensionality of the dataset to the three most important components.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 |
# example of pca for dimensionality reduction from sklearn.datasets import make_classification from sklearn.decomposition import PCA # define dataset X, y = make_classification(n_samples=1000, n_features=10, n_informative=3, n_redundant=7, random_state=1) # summarize data before the transform print(X[:3, :]) # define the transform trans = PCA(n_components=3) # transform the data X_dim = trans.fit_transform(X) # summarize data after the transform print(X_dim[:3, :]) |
Your Task
For this lesson, you must run the example and report on the structure and form of the raw dataset and the dataset after the transform was applied.
For bonus points, explore transforms with different numbers of selected components.
Post your answer in the comments below. I would love to see what you come up with.
This was the final lesson in the mini-course.
The End!
(Look How Far You Have Come)
You made it. Well done!
Take a moment and look back at how far you have come.
You discovered:
- The importance of data preparation in a predictive modeling machine learning project.
- How to mark missing data and impute the missing values using statistical imputation.
- How to remove redundant input variables using recursive feature elimination.
- How to transform input variables with differing scales to a standard range called normalization.
- How to transform categorical input variables to be numbers called one-hot encoding.
- How to transform numerical variables into discrete categories called discretization.
- How to use PCA to create a projection of a dataset into a lower number of dimensions.
Summary
How did you do with the mini-course?
Did you enjoy this crash course?
Do you have any questions? Were there any sticking points?
Let me know. Leave a comment below.




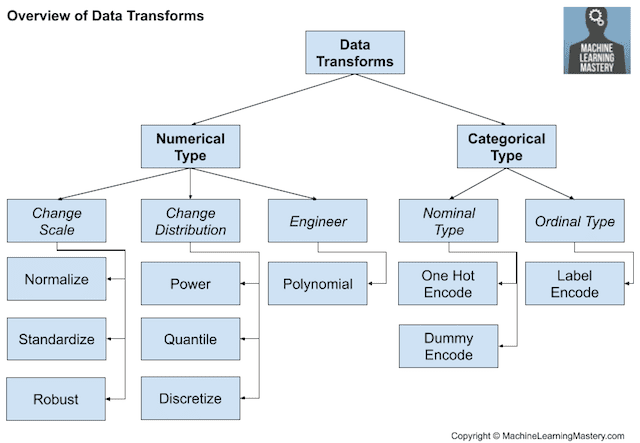

Lesson 3 and Lesson 7 are both dimensionality reduction techniques? Or it is different?
Lesson 3 is about feature selection: choose those features that are statistically meaningful to your model.
Lesson 7 is about dimensionality reduction: you have several features that are meaningful to your model, but those features are too many, or even worst: you’ve got more features than records in your dataset… so you need dimentionality reduction… PCA is based on vector spaces, at the end your new features are eigenvectors and your model will fit doing linear combinations of these new features, but you will not be able to interpret the model.
Great summary!
Great question.
Yes. Technically feature selection does reduce the number of input dimensions.
They are both transforms.
The difference is “dimensionality reduction” really refers to “feature extraction” or methods that create a lower dimensional projection of input data.
Feature selection simply selects columns to keep or delete.
Hello
Lession 3 is Feature Selection
Lession 7 is Feature Extraction
If you are little Math oriented, here is simple example,
y= f(X)= f(x1,x2,x3,x4,x5 ,x6,x7,x8)
that is your response y depends on x1 to x8 predictors(features)
And using some method you come to know that features say x3 and x5 is no way related to the response ie features x3 and x5 does not influence the response y , so you remove them in your modelling process , so now the response becomes
y= f(x1,x2,x4,x6,x7,x8)
and this is called “Feature Selection”
In Feature Extraction , you don’t use any original features directly , you will find a new set of features say z1, z2, z3,z4 (less number of features)and your problem now is
y= f(z1,z2,z3,z4), where did we get these new features from ?
they are derived or calculated from our original features, they can be some linear combinations of original features , for example z1 can be
z1= 2.5×1 + 0.33×2 + 4.00×7
one important property of new features (z’s) calculated by PCA is that all new features are independent of each other , that by no means new feature is some linear combination of other new features.
And most of the times the number of new features(z’s) are less than the number of original features(x’s) that is why it is called “Dimensionality Reduction” and at most they can be same size as original variables but they are better than original predictors at predicting response(y).
Thanks for sharing.
Lesson 02: Fill Missing Values With Imputation
executed the code of this lesson and the results are :
before imputation : Missing: 1605
after imputation : Missing: 0
Nice work!
Lesson #1: Data Preparation Algorithms
1. PCA :- it’s used to reduce the dimensionality of a large dataset by emphasising variations and revealing the strong patterns in the dataset
2. LASSO :- it involves a penalty factor that determines how many features are retained; while the coefficients of the “less important ” attributes become zero
3. RFE :- it aim at selecting features recursively considering smaller and smaller sets of features
Well done!
sir as i have used PCA based feature selection algo to select optimized features.and after that applied GSO optimization on deep learning …how can i furthur improve my results…any post processing technique
Here are some some suggestions:
https://machinelearningmastery.com/machine-learning-performance-improvement-cheat-sheet/
My results for : Lesson 03: Select Features With RFE
Column: 0, Selected=False, Rank: 4
Column: 1, Selected=False, Rank: 5
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 3
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 2
Great work!
Very well explained….????
Thanks!
Lesson 2
I added a couple of code snippets to make easier to understand the result of “SimpleImputer (strategy = ‘mean’)”
# snipplet
import matplotlib.pyplot as plt
%matplotlib inline
# Just after:
dataframe = read_csv(url, header=None, na_values=’?’)
# snippet
plt.plot(dataframe.values[:,3])
plt.plot(dataframe.values[:,4])
plt.plot(dataframe.values[:,23])
plt.show()
# Just after
Xtrans = imputer.transform(X)
#snipplet
plt.plot(Xtrans[:,3])
plt.plot(Xtrans[:,4])
plt.plot(dataframe.values[:,23])
plt.show()
Great work!
Lesson 1: Three data preparation methods
1) Detrending the data, especially in time series to understand smaller time scale processes and relate them with local forcing
2) Filtering the data if we are interested in phenomenon of particular time or space scale and remove variance contribution from other time scales
3) Replacing non-physical values with some default values to remove bias from the same
Well done!
Lesson 2:
I ran the code and found that the number of NaN values reduced from 1605 to 0.
I was interested in the working of the SimpleImputer. May be I’ll need more time to go through its code or at least have in mind its rough algorithm. But I went through its inputs. For the code you have provided, we give strategy as ‘mean’. Which tells that the Nan values will be replaced by means along the columns. But there are other strategies like ‘median’, ‘most frequent’ or a constant which tells the replacement with each strategy parameter.
There are many more customs to the imputer here. Very nice.
Thank you.
Nice, yes I recommend testing different strategies like mean and median.
using scatter plot between 2 numerical data values. it helps a lot to visualize regression
It sure does.
Lesson 3:
The make_classification function creates a (1000,10) array. The output after running till final line of code is as follows:
Column: 0, Selected=False, Rank: 4
Column: 1, Selected=False, Rank: 5
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 3
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 2
In the make_classification step, we already decide the number of informative parameters=5. I changed the number to 3 and redundant to 7. Then I found the order of above ranking changed.
After reading about RFE in its explanation, it runs the decided estimator with all the features/ columns here and then runs estimator with smaller number of features and then produces the ranks for each column. The estimator is customizable. That is nice.
I tried to change the number of features to select and got a different order of ranking. I will like to look into types of estimator.
Thank you.
Very well done, thank you for sharing your findings!
Thanks for the great article.
Would you recommend using WoE (weight of evidenve) for categorical features or even for binned continuous variables ?
Thanks.
Perhaps try it and compare the performance of resulting models fit on the data to using other encoding techniques.
Ok I’m on Holidays so I’m not a hard worker now!
Lesson 1
In an application I’ve 5 sensors, 4 measuring angles by an angular potenziometer and one linear response by a linear potenziometer. To let homogeneity among the data I used the analog to digital converted data and I normalized every value between -1 and 1 (I’ve negative and positive thresholds alarm to handle). I lowered the noise by integrate 100 samples per seconds (each sensors separately). I used a Bayes Statistical predicted value to detrand data after a collection of 128 data per sensors, mean and standard deviation computed and making the prediction with the last incoming data.
Nice work!
Also, enjoy your break!
Nice work.
How can I use machine learning with arduino or raspberry, thanks
Your mini-course is awesome,
love from Bangladesh.
Thanks!
Lesson 1: Three data preparation methods
1) Numerical data discretization – Transform numeric data into categorical data. This might be useful when ranges could be more effective than exact values in the process of modeling. For example: high-medium-low temperatures might be more interesting than the actual temperature.
2) Outlier detection – By using boxplot it is possible to identify values that can be out of the range we could expect. Outliers can be noises and hence not help the process of finding patterns in datasets.
3) Creation of new attribute – By combining existing attributes it might be interesting to create a new attribute that can help in the process of modeling. For example: temperature range based on minimal and maximal temperature.
Nice work!
Lesson 4:
I am not sure what you meant by scale in this lesson but I am copying the output prior and after the transform.
Prior transform: [[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
After transform: [[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
I see that the values have moved from being both signs to positive sign and also they are now limited within 0-1 (Normalization)
I am adding max and min of the 5 variables here before and after transform.
Max before: [4.10921382, 3.98897142, 4.0536372 , 5.99438395, 5.08933368]
Min before: [-3.55425829, -6.01674626, -4.92105446, -3.89605694, -4.97356645]
If the normalization is correct then the output ahead should be 1s and 0s.
Max after: [1., 1., 1., 1., 1.]
Min after: [0., 0., 0., 0., 0.]
I plotted each variable before and after transform, I see the character of the variable remains the same as desired but the range of the variability has changed.
That is nice. Thank you.
Very well done!
Hi anoop sir can u share ur data base
I see we have 7 lessons here. Among those lessons which ones do refer to “Feature Engineering”? Please reply.
Perhaps you can take lesson 7 as a type of feature extraction or engineering.
Some would say all transforms are a type of feature engineering.
What will be your consideration if you are asked What are the different techniques of feature engineering?
I would ask the questioner what they mean by feature engineering.
I would suggest that polynomial features are a feature engineering method, also this will help:
https://machinelearningmastery.com/discover-feature-engineering-how-to-engineer-features-and-how-to-get-good-at-it/
In Lesson 01 there is a statement like below…
“Data Requirements: Some machine learning algorithms impose requirements on the
data”
What does “machine learning algorithms impose requirements on the data” mean? Please clarify.
E.g. linear regression requires inputs to be numeric and not correlated.
Lesson 5:
In one reading I did not understand the objective of the function. Then I visited following website for more examples – https://scikit-learn.org/stable/modules/preprocessing.html#preprocessing-categorical-features
After going through more examples there I got a slight idea of the code.
Following output is raw data before transform:
[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]
After transform:
[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]
Thank you.
Well done, great work!
Lesson 2:
According to the code, there were 1605 missing values before imputation.
# print total missing before imputation
print(‘Missing: %d’ % sum(isnan(X).flatten()))
Checking after the imputation, one could observe there were no more missing values.
# print total missing after imputation
print(‘Missing: %d’ % sum(isnan(Xtrans).flatten()))
Well done.
Well explained. I always like reading your tutorials.
In Lesson 2 this is how I filled the missing values with mean values:
#read data
import pandas as pd
url = ‘https://raw.githubusercontent.com/jbrownlee/Datasets/master/horse-colic.csv’
df = pd.read_csv(url, header=None, na_values=’?’)
# is there any null values in the data frame
df.isnull().sum()
# fill the nan values with the mean of the column, for one column at a time, for column 3
df[3] = df[3].fillna((df[3].mean()))
# fill all null values with the mean of each column
df_clean = df.apply(lambda x: x.fillna(x.mean()),axis=0)
Well done.
Lesson 1:
Three Algorithms that I have used for Data Preprocessing
Data standardization that standardizes the numeric data using the mean and standard deviation of the column.
I have also used Correlation Plots to identify the correlations and along with that I have used VIF to identify interdependently columns.
Apart from that I have used simple find and replace to replace garbage or null values in the data with mean, mode, or static values.
Nice work!
Sorry, What is VIF, thank you
Lesson 03
My results:
Column: 0, Selected=False, Rank: 4
Column: 1, Selected=False, Rank: 6
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 5
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 3
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 2
As far as I could understand, the five features selected were columns 2,3,4,6, and 8.
The remaining features were ranked in the following order:
Column 9, Column 7, Column 0, Column 5, and Column 1.
Excellent work!
Lesson 6:
Raw data:
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
After Transform:
[[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
By observing the output, I see that the raw data is spread in between lowest value -5.77 to maximum 2.39. Now after transform, it is like binning the data in classes 0-7. So we see that lowest value is put in class 0 and highest value in class 7. These are only 3 rows out of 1000 rows. Now it depends on the size of the bin, which class will each individual raw value fall in. Now this observations also depend on the strategy (above = uniform) inputs in the command line. In the above example, the size of bins is uniform.
When I changed it to kmeans, I see following as output:
[[8. 0. 4. 1. 6.]
[4. 8. 2. 6. 4.]
[8. 4. 4. 4. 4.]]
The higher values are pushed to higher bin numbers. Thus the size of bins are varying.
When I changed the encode to onehot, it generates a sparse matrix instead of numpy array in the earlier matrix. It encodes each entry. I don’t know how this will help in understanding data distribution. Following was the output:
(0, 4) 1.0
(0, 5) 1.0
(0, 12) 1.0
(0, 15) 1.0
(0, 23) 1.0
(1, 1) 1.0
(1, 9) 1.0
(1, 11) 1.0
(1, 18) 1.0
(1, 22) 1.0
This was an interesting exercise. Thank you.
Well done!
Lesson #2: Identifying and Filling Missing values in Data.
I executed the code example and here are the findings before and after imputation
Before: 1605
After: 0
I also tried to play around with the different strategies that can be used on the imputer
Great work!
I have a question on Lesson 05. If I have a category with only 2 classes, should I use the Hot Encoder or simply transform those 2 classes to binary, i.e. class 1: binary 0, class 2: binary 1.
Because, as I understand, the Hot Encoder will encode to (0 1) and (1 0) instead.
Thank you
Fantastic question!
You can, but no, typically we model binary classification as a single variable with a binomial probability distribution.
Thank you.
One thing, is there any notification to the email if my question/thread is answered already on this blog tutorial?
Not at this stage, sorry.
Lesson7:
As I understood, PCA is used to reduce the number of unnecessary variables. Here, with the method we selected 3 variables/ columns out of 10 using PCA. Are these three columns first three PCA axis/ variables?
Following was the result with the given input values:
Before-
[[-0.53448246 0.93837451 0.38969914 0.0926655 1.70876508 1.14351305
-1.47034214 0.11857673 -2.72241741 0.2953565 ]
[-2.42280473 -1.02658758 -2.34792156 -0.82422408 0.59933419 -2.44832253
0.39750207 2.0265065 1.83374105 0.72430365]
[-1.83391794 -1.1946668 -0.73806871 1.50947233 1.78047734 0.58779205
-2.78506977 -0.04163788 -1.25227833 0.99373587]]
After PCA-
[[-1.64710578 -2.11683302 1.98256096]
[ 0.92840209 4.8294997 0.22727043]
[-3.83677757 0.32300714 0.11512801]]
I observed that the numbers are different in the new variables than compared to values in original variable. Is this because we have defined 3 new variables out of 10 variables?
Well done!
No, they are entirely new data constructed from the raw data.
https://github.com/ksrinivasasrao/ATL/blob/master/Untitled3.ipynb
Sir can u please help me with error i am getting
Perhaps you can summarize the problem that you’re having in a sentence or two?
Lesson : Data Preparation and some techniques
Data preparation is the process of cleaning and transforming raw data prior to processing and analysis.
Good data preparation allows for efficient analysis, limit errors, reduces anomalies, and inaccuracies that can occur during data processing and makes all processed data moire accessible to users.
Some other techniques:
a) Data Wrangling/Cleaning – Is the process of cleaning and unifying messy and complex data for easy access and analysis. This involves filling the missing values and getting rid of the outliers in the data set.
b) Data discretization – Part of data reduction but with particular importance especially for numeral data. In this routine the raw values of numeric attribute are placed by interval label(bins) or conceptual labels.
c) Data reduction – Obtain reduced representation in volume but places the same or similar analytical results. Some of those techniques are High Correlation filter, PCA, Random Forest/Decision trees, and Backward/Forward Feature Elimination.
Well done!
Hi Jason,
I have below query.
For Regression, Classification, time series forecasting models we come across terms like Adjusted R Squared, Accuracy_Score, MSE, RMSE, AIC, BIC for evaluating the model performance ( you can let me know if I missed any other metric here)
How many of the above accuracy metrics need to be used for any model? what combination of them is to be used? Is it model dependant?
Pick one metric and optimize it.
Hello Jason,
I saw 1605 missing before imputation and of course 0 missing after.
Thanks for your tutorial.
Nice!
Lesson 1:
Here are the algorithms I came across, I am just starting out in machine learning (smiles )
Independent Component Analysis: used to separate a complex mix of data into their different sources
Principal Component Analysis (PCA): used to reduce the dimensionality of data by creating new features. It does this to increase their chances of being interpret-able while minimising information loss
Forward/Backward Feature Selection: also used to reduce the number of features in a set of data however unlike PCA it does not create new features.
Thanks Jason!!!
Great work!
ps: I couldnt find the way to comment independently so I’ll leave one as a reply to Dalapo since it will also be on lesson 1.
Data preparation methods that I would use before creating machine learning models:
1. Quickly check key statistical value for each feature using Pandas’ DataFrame.describe( ) which shows the minimum, maximum, median, mean, standard deviation and each quartile value for each columns in the data frame. Then from here I can examine the dataset to see if there is any obviously incorrect values and outliers.
2. Check the correlation of each feature relative to each other using the Pandas’ DataFrame.corr( ) function which returns a matrix of correlation between each variable and to visualize the correlation by using seaborn’s heatmap graph. sns.heatmap(data.corr( ))
3. Finally, after gaining a brief understanding of all the features, I will dive into features that I think is important and visualize their frequency distribution, spread and further explore their relationship with other variables.
Thank you!! Your blog is so helpful for someone trying to learn machine learning at home like me!!
Well done!
Lesson #2
I got the following result:
Total missing values before running the code = 1605 and zero after imputation.
I came across other methods of data imputation such as deductive, regression and stochastic regression imputation.
Lesson #3
Result:
Column: 0, Selected=False, Rank: 5
Column: 1, Selected=False, Rank: 4
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 2
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 3
The columns selected were 2, 3, 4, 6 & 8.
NB:My result was different from the above when i used Pycharm (I used Jupyter for the above result).
Could there be an explanation for this?
Well done!
This might help with the different results:
https://machinelearningmastery.com/different-results-each-time-in-machine-learning/
Many thanks!!! This was helpful
You’re welcome!
At first I thought using one- hot encoding would skew the results since it is difficult to decide how much of an influence a categorical feature should have versus numeric ones. I then realized a coefficient would solve the problem of deciding how much exactly any feature, categorical one-hot-encoded or otherwise, affects the output.
3 Data Preparation algorithms – Standardization, Encoding(Label encoding, one hot encoding) & (mean 0, variance 1), dimensionality reduction (PCA, LDA).
Nice work!
Data Preparation Algorithms:
1) Normalization : To normalize numeric columns ranges on scale to reduce difference between ranges.
2) Standardization: To standardize numeric input values with mean and standard deviation to reduce differences between values.
3) NominalToBinary: To convert nominal values into binary values.
I’m just a beginner, correct me if I’m wrong.
Well done!
As asked in first lesson of Data Preparation here is the list of some Data Preparation algorithms:
Data preparation algorithms are
PCA- Mainly used for Dimensionality Reduction
Data Transformation- One-Hot Transform used to encode a categorical variable into binary variables.
Data Mining & Aggregation
Nice work!
Lesson 1 response:
Trying to think of things that have not already been mentioned and are not part of later chapters in this course, so here are my thoughts:
1. Cleaning data to combine multiple instances of what is essentially the same observation, with different spellings, for eg. like customer first name Mike, Michael, or M all relating to the same customer ID. This gives us a truer picture of the values of an attribute for each observation. For eg, if we are measuring customer loyalty by number of purchases by a given customer, we need all those records for Michael combined into one.
2. Pivoting data into long form by removing separate field attributes for what should be one field (eg years 2018, 2019, 2020 as separate attributes rather than pivoted into a single attribute “Year”). This allows for easier analysis of the dataset.
3. Removing features with a constant value across all records. These add no value to the predictive model.
Well done!
Lesson 2 response:
Missing 1605 before imputing values
None after
Lesson 3 response: 5 redundant features at random were requested, so only the remaining non-redundant ones were retained, ranked at 1.
Column: 0, Selected=False, Rank: 4
Column: 1, Selected=False, Rank: 6
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 5
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 3
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 2
Lesson 4 response: keeps the range of values within [0, 1]. While the benefit may not be so obvious with this dataset, when we have sets that could have unrestricted values, this makes analysis more manageable.
Before the transform: [[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
And after: [[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
Lesson 5 response: makes it easier to use the variables for analysis than when they had ranges or descriptions in string format.
Before encoding: [[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]]
And after: [[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]]
Lesson 6 response: floats are now discrete buckets post-transform. When a feature could take on just about any continuous value, grouping ranges of them into buckets makes analysis easier.
Before transform: [[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
And after: [[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
Lesson 7 response: the transform reduced the dimension of the dataset to the requested 3 main features, effecting a transformation as opposed to a mere selection of features as in lesson 3. I need to explore this further to understand it better.
Before the transform: [[-0.53448246 0.93837451 0.38969914 0.0926655 1.70876508 1.14351305
-1.47034214 0.11857673 -2.72241741 0.2953565 ]
[-2.42280473 -1.02658758 -2.34792156 -0.82422408 0.59933419 -2.44832253
0.39750207 2.0265065 1.83374105 0.72430365]
[-1.83391794 -1.1946668 -0.73806871 1.50947233 1.78047734 0.58779205
-2.78506977 -0.04163788 -1.25227833 0.99373587]]
And after: [[-1.64710578 -2.11683302 1.98256096]
[ 0.92840209 4.8294997 0.22727043]
[-3.83677757 0.32300714 0.11512801]]
Working separately on responses to bonus questions.
Well done!
for lesson 2, the printed output as;
Missing: %d 1605
Missing: %d 0
Nive work.
list three data preparation algorithms that you know of or may have used before and give a one-line summary for its purpose.
1. Principal Component Analysis
Reduces the dimensionality of dataset by creating new features that correlate with more than one original feature.
2. Decision Tree Ensembles
Used for feature selection
3. Forward Feature Selection and Backward Feature Selection
Applied to reduce the number of features.
Well done!
Lesson #1
1. Binary Encoding: coverts non-numeric data to numeric values between 0 and 1.
2. Data standardization: converts the structure of disparate datasets into a Common Data Format.
Nice work!
Lesson #2
Missing values before imputation: 1605
Missing values after imputation: 0
Well done!
Lesson #3
Column: 0, Selected=False, Rank: 6
Column: 1, Selected=False, Rank: 4
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 5
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 3
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 2
From the above, features of column 2,3,4,6, and 8 were selected while the remaining were discarded. The selected features were all ranked 1 while columns 9, 7, 1,0, and 5 were ranked 2, 3, 4, 5 and 6, respectively.
Great work!
for day 2
Missing: 1605
Missing: 0
Nice work!
Lesson 02:
Missing: 1605
Missing: 0
Thanks Jason
I´m just begineer.
Can you explain this line to me.
ix = [i for i in range(data.shape[1]) if i != 23]
X, y = data[:, ix], data[:, 23]
Thanks
Well done.
Good question, all columns that are not column 23 are taken as input, and column 23 is taken as the output.
Lesson 5
before
array([[“’50-59′”, “‘ge40′”, “’15-19′”, …, “‘central'”, “‘no'”,
“‘no-recurrence-events'”],
[“’50-59′”, “‘ge40′”, “’35-39′”, …, “‘left_low'”, “‘no'”,
“‘recurrence-events'”],
[“’40-49′”, “‘premeno'”, “’35-39′”, …, “‘left_low'”, “‘yes'”,
“‘no-recurrence-events'”],
…,
[“’30-39′”, “‘premeno'”, “’30-34′”, …, “‘right_up'”, “‘no'”,
“‘no-recurrence-events'”],
[“’50-59′”, “‘premeno'”, “’15-19′”, …, “‘left_low'”, “‘no'”,
“‘no-recurrence-events'”],
[“’50-59′”, “‘ge40′”, “’40-44′”, …, “‘right_up'”, “‘no'”,
“‘no-recurrence-events'”]], dtype=object)
After to apply one hot encode
[[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]
[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 1. 0. 0. 0. 0. 0. 1.]]
Thank you so much, Sir Jason for sharing this tutorial
Nice work!
Lesson#3 output: So 5,4,6,2 are selected
Column: 0, Selected=False, Rank: 5
Column: 1, Selected=False, Rank: 4
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 3
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 2
Well done!
#Lesson :4
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
Nice work.
# Lesson :5
[[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]]
[[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]]
Nice work!
# lesson 6
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
[[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
Lesson:7
[[-1.64710578 -2.11683302 1.98256096]
[ 0.92840209 4.8294997 0.22727043]
[-3.83677757 0.32300714 0.11512801]]
Before transforming my data
[[-0.53448246 0.93837451 0.38969914 0.0926655 1.70876508 1.14351305
-1.47034214 0.11857673 -2.72241741 0.2953565 ]
[-2.42280473 -1.02658758 -2.34792156 -0.82422408 0.59933419 -2.44832253
0.39750207 2.0265065 1.83374105 0.72430365]
[-1.83391794 -1.1946668 -0.73806871 1.50947233 1.78047734 0.58779205
-2.78506977 -0.04163788 -1.25227833 0.99373587]]
After transforming the data
[[-1.64710578 -2.11683302 1.98256096]
[ 0.92840209 4.8294997 0.22727043]
[-3.83677757 0.32300714 0.11512801]]
here we are reducing to 3 variables.
My question how should I know how many variables to reduce to or maybe I don’t need to reduce or should I always reduce the input variables?
can also reduce the target or dependent variable?
Please Sir Jasom.
Thanks a lot.
Good question, trial and error in order to discover what works best for your dataset.
You only have one target. You can transform it, but not reduce it.
Great “course”!
For the feature selection and feature extraction (lessons 3 and 7), both call for some prior knowledge of picking the right bins or components. Like for the PCA, we chose to have 3 eigenvectors, is there a good process for selecting the right number? I’m sure we can just train a model for 3 or 5 or 7 vectors and find out, but is there a better understanding to be had?
Thanks,
Thanks!
Yes, a grid search over the options is a great way to go.
Lesson 02:
Missing before processing with imputer: 1605, after 0
Well done.
Lesson 04:
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
Great work!
Lesson 05:
# one hot encode the breast cancer dataset
from pandas import read_csv
from sklearn.preprocessing import OneHotEncoder
# define the location of the dataset
url = “https://raw.githubusercontent.com/jbrownlee/Datasets/master/breast-cancer.csv”
# load the dataset
dataset = read_csv(url, header=None)
# retrieve the array of data
data = dataset.values
# separate into input and output columns
X = data[:, :-1].astype(str)
y = data[:, -1].astype(str)
# summarize the raw data
print(X[:3, :])
# define the one hot encoding transform
encoder = OneHotEncoder(sparse=False)
# fit and apply the transform to the input data
X_oe = encoder.fit_transform(X)
# summarize the transformed data
print(X_oe[:3, :])
Excellent.
Lesson 1
Previously I have worked with SQL and Pandas for data cleaning, but I have started studying with your books now and used these for help.
Please, if you could let me know if I understood it right, it would be highly appreciated. Thank you.
Standardisation: Scales the variable to a standard Gaussian probability distribution (mean of zero and standard deviation of one).
Power Transformer: Removes the skew from the probability distribution of a variable and makes it more Gaussian-like, which means that it falls more equally on both sides.
Quantile Transformer: Transforms features to follow a normal distribution and reduces the impact of outliers.
I understand they are all aiming for the Gaussian distribution? Thank you very much for your help.
Nice work!
Lesson 2
1605 before and 0 after. Thank you.
Nice work!
Lesson 3
column: 0, Selected=False, Rank: 4
column: 1, Selected=False, Rank: 6
column: 2, Selected=True, Rank: 1
column: 3, Selected=True, Rank: 1
column: 4, Selected=True, Rank: 1
column: 5, Selected=False, Rank: 5
column: 6, Selected=True, Rank: 1
column: 7, Selected=False, Rank: 2
column: 8, Selected=True, Rank: 1
column: 9, Selected=False, Rank: 3
Everything worked fine, thank you.
Nice work.
Lesson 4: Normalization
Data before the transform:
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
Data after the transform:
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
All in the range 0-1
Nice work!
Lesson 3:RFE
Only 5 features from columns 2, 3, 4, 6, and 8 were selected.
Nice work!
Lesson 5: One Hot Encoding
Raw data:
[[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]]
Transformed data:
[[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]]
That worked fine too.
Just a quick question about y: Are we supposed to transform it too?
# separate into input and output columns
X = data[:, :-1].astype(str)
y = data[:, -1].astype(str)
Thank you and Merry Christmas.
Yes, it is a good idea to transform inputs and outputs separately, so you can invert the transform later separately for predictions.
Nice work!
Lesson 6: KBinsDiscretizer
Before:
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
After Discretization:
[[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
n_bins=20
[[15. 0. 9. 3. 11.]
[ 8. 15. 5. 12. 8.]
[15. 10. 9. 10. 8.]]
strategy=’kmeans’
[[8. 0. 4. 1. 6.]
[4. 8. 2. 6. 4.]
[8. 4. 4. 4. 4.]]
strategy=’quantile’
[[9. 0. 4. 0. 8.]
[4. 8. 1. 8. 4.]
[9. 1. 4. 5. 4.]]
Thank you.
Well done.
Lesson 7: Dimensionality Reduction with PCA
Before:
[[-0.53448246 0.93837451 0.38969914 0.0926655 1.70876508 1.14351305
-1.47034214 0.11857673 -2.72241741 0.2953565 ]
[-2.42280473 -1.02658758 -2.34792156 -0.82422408 0.59933419 -2.44832253
0.39750207 2.0265065 1.83374105 0.72430365]
[-1.83391794 -1.1946668 -0.73806871 1.50947233 1.78047734 0.58779205
-2.78506977 -0.04163788 -1.25227833 0.99373587]]
After:
[[-1.64710578 -2.11683302 1.98256096]
[ 0.92840209 4.8294997 0.22727043]
[-3.83677757 0.32300714 0.11512801]]
With two components:
[[ 0.16205607 0.682448 ]
[-2.73725 -0.90545667]
[-2.86555495 -5.344142 ]]
With four components:
[[-1.64710578e+00 -2.11683302e+00 1.98256096e+00 -3.00364400e-16]
[ 9.28402085e-01 4.82949970e+00 2.27270432e-01 1.95852098e-15]
[-3.83677757e+00 3.23007138e-01 1.15128013e-01 -1.33926993e-16]]
Thank you very much.
Nice work.
Outliers — using a scatter plot in matplotlib and seaborn. Outliers can identify data that is anomalous or does not belong in the datasets and can identify mistakes in data such as mistakes in data entry. Also can reveal values that are out of range and values out of range caused by calculations and data cleaning
Data Standardization: Standards values to ensure there is consistency and to calculate the appropriate and correct number of unique values and value counts. Check for misspellings, values that can be grouped together to make data easier to work with
Filling null values and dropping values – dropping values that will not impact the data and filling in null values as some ML algorithms do not perform with no value or a character like . or , or whitespace.
Well done!
####Lesson 1:
The below three Algorithms I have used for Data Preperation
1. Data standardization that standardizes the numeric data using the mean and standard deviation of the column.
2. Correlation to identify the correlations between the data points.
3. I have used simple find and replace to replace garbage, Nan, blank or null values in the data with mean, mode, or static values of that column.
Well done.
###Lesson 2:
1605 before imputation
0 after imputation
Nice work!
###Lesson 3:
Column: 0, Selected=False, Rank: 6
Column: 1, Selected=False, Rank: 4
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 5
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 3
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 2
From the above, features of column 2,3,4,6, and 8 were selected while the remaining were discarded. The selected features were all ranked 1 while columns 9, 7, 1,0, and 5 were ranked 2, 3, 4, 5 and 6, respectively.
Excellent!
###Lesson 4:
data before the transform:
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
data after the transform:
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
Well done!
###Lesson 5:
the raw data before:
[[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]]
the transformed data after:
[[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]]
Excellent!
###Lesson 6: KBinsDiscretizer
Before:
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
After Discretization:
[[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
n_bins=5
[[3. 0. 2. 0. 2.]
[2. 3. 1. 3. 2.]
[3. 2. 2. 2. 2.]]
n_bins=20
[[15. 0. 9. 3. 11.]
[ 8. 15. 5. 12. 8.]
[15. 10. 9. 10. 8.]]
strategy=’quantile’
[[9. 0. 4. 0. 8.]
[4. 8. 1. 8. 4.]
[9. 1. 4. 5. 4.]]
strategy=’kmeans’
[[8. 0. 4. 1. 6.]
[4. 8. 2. 6. 4.]
[8. 4. 4. 4. 4.]]
encode=’onehot-dense’, strategy=’kmeans’
[[0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.
1. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0.
0. 0.]
[0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 1. 0.
0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0.]
[0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 0.
1. 0. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0.]]
Great work!
Lesson #1 Data Preparation
Ridge Regression: The L2 Regularisation is also known as Ridge Regression or Tikhonov Regularisation. Ridge regression is almost identical to linear regression (sum of squares) except we introduce a small amount of bias.
Genetic Algorithms: This algorithm can be used to find a subset of features.
Linear Discriminant Analysis (LDA): LDA makes assumptions about normally distributed classes and equal class covariances.
Nice work!
Lesson #2 Filling Missing Values
Missing values before Imputation: 408
Missing values after Imputation : 0
Well done!
Lesson # 3
Column: 0, Selected=False, Rank: 4
Column: 1, Selected=False, Rank: 5
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 2
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 3
Selected Features Ranked as 1
Great work!
Lesson 1 – 3 (basic) data cleaning approaches
1. Eliminate variables/fields that have either no values or a single value.
2. Modify variables/fields that have blank values where Null or NaN are appropriate.
3. Standardize responses with same meaning, e.g., CA or Calif or California
Well done!
Lesson1:
-the elimination of null values throught the sustitution for the median or other caracterictic value, or elimination of this rows to the dataset, if you are a big dataset.
-the conversion of cathegorical variables in numeric variables, througt the creation of dummys variables
Well done!
lesson2:
Missing values in dataframe without imputer: 1605
Missing values in datafram after imputer: 0
Well done!
Lesson # 3
ths output for the code mentioned is:
Column: 0, Selected=False, Rank: 4
Column: 1, Selected=False, Rank: 5
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 2
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 3
Applying the RFE to the horse colic dataset we obtain:
Column: 0, Selected=False, Rank: 20
Column: 1, Selected=False, Rank: 21
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=False, Rank: 6
Column: 5, Selected=False, Rank: 8
Column: 6, Selected=False, Rank: 13
Column: 7, Selected=False, Rank: 10
Column: 8, Selected=False, Rank: 12
Column: 9, Selected=False, Rank: 11
Column: 10, Selected=False, Rank: 14
Column: 11, Selected=False, Rank: 16
Column: 12, Selected=False, Rank: 7
Column: 13, Selected=False, Rank: 19
Column: 14, Selected=False, Rank: 4
Column: 15, Selected=False, Rank: 9
Column: 16, Selected=False, Rank: 17
Column: 17, Selected=False, Rank: 18
Column: 18, Selected=False, Rank: 2
Column: 19, Selected=True, Rank: 1
Column: 20, Selected=True, Rank: 1
Column: 21, Selected=True, Rank: 1
Column: 22, Selected=False, Rank: 5
Column: 23, Selected=False, Rank: 15
Column: 24, Selected=False, Rank: 3
Column: 25, Selected=False, Rank: 22
Well done.
Lesson 4#
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
max for column: [4.10921382 3.98897142 4.0536372 5.99438395 5.08933368]
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
max for column: [1. 1. 1. 1. 1.]
Great work!
Lesson 5#
the output is:
[[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]]
[[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]]
we had 9 column that its converts in 40 column after the one-hot encoding process
Excellent.
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
n_bins=10, encode=’ordinal’, strategy=’uniform’
[[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
n_bins=10, encode=’ordinal’, strategy=’quantile’
[[9. 0. 4. 0. 8.]
[4. 8. 1. 8. 4.]
[9. 1. 4. 5. 4.]]
n_bins=10, encode=’ordinal’, strategy=’kmeans’
[[8. 0. 4. 1. 6.]
[4. 8. 2. 6. 4.]
[8. 4. 4. 4. 4.]]
n_bins=3, encode=’ordinal’, strategy=’uniform’
[[2. 0. 1. 0. 1.]
[1. 2. 0. 1. 1.]
[2. 1. 1. 1. 1.]]
n_bins=3, encode=’ordinal’, strategy=’quantile’
[[2. 0. 1. 0. 2.]
[1. 2. 0. 2. 1.]
[2. 0. 1. 1. 1.]]
n_bins=3, encode=’ordinal’, strategy=’kmeans’
[[2. 0. 1. 0. 2.]
[1. 2. 0. 2. 1.]
[2. 1. 1. 1. 1.]]
as conclusion its can see that if you reduce the number of bins, all strategies has a similar result
Well done!
Lesson 6#
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
n_bins=10, encode=’ordinal’, strategy=’uniform’
[[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
n_bins=10, encode=’ordinal’, strategy=’quantile’
[[9. 0. 4. 0. 8.]
[4. 8. 1. 8. 4.]
[9. 1. 4. 5. 4.]]
n_bins=10, encode=’ordinal’, strategy=’kmeans’
[[8. 0. 4. 1. 6.]
[4. 8. 2. 6. 4.]
[8. 4. 4. 4. 4.]]
n_bins=3, encode=’ordinal’, strategy=’uniform’
[[2. 0. 1. 0. 1.]
[1. 2. 0. 1. 1.]
[2. 1. 1. 1. 1.]]
n_bins=3, encode=’ordinal’, strategy=’quantile’
[[2. 0. 1. 0. 2.]
[1. 2. 0. 2. 1.]
[2. 0. 1. 1. 1.]]
n_bins=3, encode=’ordinal’, strategy=’kmeans’
[[2. 0. 1. 0. 2.]
[1. 2. 0. 2. 1.]
[2. 1. 1. 1. 1.]]
As conclusion its can see that if you reduce the number of bins, all strategies has a similar result
Well done!
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
n_bins=10, encode=’ordinal’, strategy=’uniform’
[[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
n_bins=10, encode=’ordinal’, strategy=’quantile’
[[9. 0. 4. 0. 8.]
[4. 8. 1. 8. 4.]
[9. 1. 4. 5. 4.]]
n_bins=10, encode=’ordinal’, strategy=’kmeans’
[[8. 0. 4. 1. 6.]
[4. 8. 2. 6. 4.]
[8. 4. 4. 4. 4.]]
n_bins=3, encode=’ordinal’, strategy=’uniform’
[[2. 0. 1. 0. 1.]
[1. 2. 0. 1. 1.]
[2. 1. 1. 1. 1.]]
n_bins=3, encode=’ordinal’, strategy=’quantile’
[[2. 0. 1. 0. 2.]
[1. 2. 0. 2. 1.]
[2. 0. 1. 1. 1.]]
n_bins=3, encode=’ordinal’, strategy=’kmeans’
[[2. 0. 1. 0. 2.]
[1. 2. 0. 2. 1.]
[2. 1. 1. 1. 1.]]
As conclusion its can see that if you reduce the number of bins, all strategies has a similar result
Great.
As conclusion its can see that if you reduce the number of bins, all strategies has a similar result
Beform Transformation :
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
After Normalization :
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
Nice work!
Lesson #5
[[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]]
[[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]]
Well done.
Lesson #6
Before Transformation :
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
After Transformation : I have chosen bins as 20
[[15. 0. 9. 3. 11.]
[ 8. 15. 5. 12. 8.]
[15. 10. 9. 10. 8.]]
Great work!
Lesson #7
I have changed the components to 5
[[-0.53448246 0.93837451 0.38969914 0.0926655 1.70876508 1.14351305
-1.47034214 0.11857673 -2.72241741 0.2953565 ]
[-2.42280473 -1.02658758 -2.34792156 -0.82422408 0.59933419 -2.44832253
0.39750207 2.0265065 1.83374105 0.72430365]
[-1.83391794 -1.1946668 -0.73806871 1.50947233 1.78047734 0.58779205
-2.78506977 -0.04163788 -1.25227833 0.99373587]]
[[-1.64710578e+00 -2.11683302e+00 1.98256096e+00 2.05848176e-15
6.57302581e-16]
[ 9.28402085e-01 4.82949970e+00 2.27270432e-01 1.12515298e-15
-5.70714602e-16]
[-3.83677757e+00 3.23007138e-01 1.15128013e-01 -3.85082150e-16
-2.59561787e-16]]
Great work!
Hi Jason!
Can I just do either feature selection or dimensionality reduction? Because if I did both it might result in information loss. In what type of situation that I just need neither feature selection or dimensionality reduction, but not both?
Yes, either but not both.
I am new to the field and still learning about algorithms.
I have learned to search for missing records or data; this helps streamline or clean up the data (excluding missing values.
Another I have used is tree pruning to allow the data to be easier to visualise.
As well as using the best model fit algorithm.
Well done!
Before
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
After
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
Largest element Before Transformation: -5.777320483664414
Smallest element Before Transformation: 2.3932448901303873
Largest element in Transformed array: 0.7959030365356285
Smallest element in Transformed array: 0.023928895614900747
Well done!
Whats your thoughts on using manual dimension reduction techniques, like covariance, pairwise correlation, multi-collinearity and correlation with target, vs more automated techniques. Also from what I gather running the model and getting feature importance from many models that offer it seems more of an accurate approach than the mathematical (as well as non visual) approach mentioned above, but if a huge data set is on hand where this is not possible without distributed computing via cloud systems (at a cost), the former seems more traditional.
If it works well for you, go for it!
Lesson #1
1.Removing null values by substitution for the median or other methods and deleting the rows related to this null value
2. remove columns with single value (zero-variance) with the unique operator
3.Converting categorical variables to numeric variables (numbers), by creating dummies variables
Well done!
Lesson #2
I couldn’t read the csv file directly from the URL so I downloaded it.
number of Missing values before imputation: 1605
number of Missing values after imputation: 0
Well done!
Lesson 1
Principal Component Analysis, or PCA, is a dimensionality-reduction method that is often used to reduce the dimensionality of large data sets, by transforming a large set of variables into a smaller one that still contains most of the information in the large set.
Well done!
PCA
Forward Feature Selection: This one is used to reduce the number of features.
Decision Tree Ensemble
Good! Keep on.
Lesson 2
Missing: 1605
Missing: 0
Column: 0, Selected=False, Rank: 3
Column: 1, Selected=False, Rank: 5
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 2
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 4
Good job!
lesson #4
Before Tranformation
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
After Transformation
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
The scale before transformation (-6 – 3) while scale is (0 – 1) after transformation.
That looks great. Keep on!
Lesson #5
[[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]]
[[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]]
Lesson #6
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
[[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
when the n_bin is set to 6
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
[[4. 0. 2. 1. 3.]
[2. 4. 1. 3. 2.]
[4. 3. 2. 3. 2.]]
when the n_bin is set to 20
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
[[15. 0. 9. 3. 11.]
[ 8. 15. 5. 12. 8.]
[15. 10. 9. 10. 8.]]
Lesson #7
[[-0.53448246 0.93837451 0.38969914 0.0926655 1.70876508 1.14351305
-1.47034214 0.11857673 -2.72241741 0.2953565 ]
[-2.42280473 -1.02658758 -2.34792156 -0.82422408 0.59933419 -2.44832253
0.39750207 2.0265065 1.83374105 0.72430365]
[-1.83391794 -1.1946668 -0.73806871 1.50947233 1.78047734 0.58779205
-2.78506977 -0.04163788 -1.25227833 0.99373587]]
[[-1.64710578 -2.11683302 1.98256096]
[ 0.92840209 4.8294997 0.22727043]
[-3.83677757 0.32300714 0.11512801]]
I see you posted your work on Lessons 5-7 in a day and you completed it. Well done!
Lesson#1
1. Principle Component Analysis (PCA) to reduce the dimensionality of the features. Mitigates model complexity by reducing the dimensionality of the dataset, while retaining the predictive information.
2. K-Means Clustering. Unsupervised learning technique that groups datasets into clusters based on inherent characteristics. The cluster labels can then reveal further insights / be used for predictive machine learning tasks.
3. Statsmodels deterministic process, to derive time series features (e.g. seasonality)
Great work!
Lesson 2: Imputation
total missing values 1605. Using imputer, the missing values are replaced using the mean along each column of the data;data after imputation has 0 missing values.
Good. Keep on!
Lesson 3:
My results:
Column: 0, Selected=False, Rank: 4
Column: 1, Selected=False, Rank: 6
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 5
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 2
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 3
Varying the random_state of the classifier, the rank for the unselected columns may vary, but the selected columns are consistent.
However, I’m not certain regarding the implications moving forward – translated to another dataset, these columns could refer to one hot encoded features such as ‘blue-car’, ‘green-car’, ‘red-car’, etc. If only ‘blue-car’ and ‘green-car’ columns were selected as relevant columns, does that mean that the ‘red-car’ group is not information with respect to target classification in the first place? Thanks
Yes, that’s correct. Probably that’s too common with not much information. For example, in England, you one-hot encode people who can speak English is probably meaningless, but if you encode people who can speak Korean or Vietnamese, that might mean something.
Thank you!
Hi!
Thank you very much abour the lessons, It’s a very useful course! 🙂
My doubt is: when we have to prepare the data, the correct order is the order of the lessons?
I mean: first we have to sill missing values, then select features, then scale data with normalization…
The data normalization is after the features selection?
Thank you very much 🙂
It is recommended but not necessary. You can always skip lessons!
Here are some data preparation algorithms:
1. drop() – Get rid off features which have nothing to do with label (col(s) that to be predicted)
2. drop_duplicates() – get rid off duplicate data
3.. isnull().isnull() or isnull().any(axis=1) – identify rows/observations which have null values in one or more features
4.. replace() – converts object data type to integers/floats when algorithms demand int/float values
5.. SimpleImputer(missing_values = np.nan, strategy=’???’) – replace null value with (???) mean, median, mode(most frequent) or a specific value (constant)
Note: This is tutorial exercise given to me by Jason. Thanks Jason
hi . don’t be tired ! so active you are ,
I am in these course . I use from these lessons
I had a request . is it possible for you that give a complete example , a typic example , that all of lessons can be done with it? for example we can do prepare , clean , select feature and etc. on it ?
I knowing is difficult , but not for you.
sincerely …hossein
so thank you for your great efforts.
Do you think this 7-day steps is same as what you wanted?
hi
2,3,4,6,8 were true column and others false , so thanks
I run script and the answer is as these file
Thank you for the feedback Hossein!
Regards,
hi
how are you ?
I learn so much , all subject were great
thank you
Thank you Hossein for your feedback and kind words!
Hi,
the three data preparation algorithms/steps that i have used in my previous process are
1) Data Ingest / Data loading
This is the first step after the data is identified, it is loaded into the tables or a BI tool to do some analysis the data.
2) Data Cleanse – In this step we identify if there are any NULL / NaaN and try to fill in or remove those rows depending upon the situation
3) Analyze – Here we do the analysis and identify what data points will be used as features.
Thank you for your feedback Himanshu! Keep up the great work!
Regards,
Hi Jason,
I read the 2nd chapter and saw that in the file horse-colic.csv there are 1605 values which have non numeric data (?). After applying the SimpleImputer method on the dataset there are no values with NaN data.
I had one question why in the code do we have to seperate the data in to train and test in the following line.
X, y = data[:, ix], data[:, 23]
thanks
Hi Jason,
I read the 3rd chapter and the following features were selected.
Column: 2
Column: 3
Column: 4
Column: 6
Column: 8
thanks
Thank you for the feedback, Himanshu!
HELLO, for day2 I run the code and get these:1605, 0.
Thank you.
Thank you for the feedback, Mitra!
Hello.
Result of lesson 3:
Column: 0, Selected=False, Rank: 4
Column: 1, Selected=False, Rank: 5
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 2
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 3
Thank you so much
I have practiced about imputing missing values with simple imputer method in sklearn.
here below is the code for simple imputer.
from numpy import isnan
from pandas import read_csv
from sklearn.impute import SimpleImputer
# load dataset
url = ‘https://raw.githubusercontent.com/jbrownlee/Datasets/master/horse-colic.csv’
dataframe = read_csv(url, header=None, na_values=’?’)
# split into input and output elements
data = dataframe.values
ix = [i for i in range(data.shape[1]) if i != 23]
X, y = data[:, ix], data[:, 23]
# print total missing
print(‘Missing: %d’ % sum(isnan(X).flatten()))
# define imputer
imputer = SimpleImputer(strategy=’mean’)
# fit on the dataset
imputer.fit(X)
# transform the dataset
Xtrans = imputer.transform(X)
# print total missing
print(‘Missing: %d’ % sum(isnan(Xtrans).flatten()))
The number of missing values before imputing are 1605 and after imputing the number of missing values are 0 meaning all missing values are replaced by the mean of each columns of the dataset.
The general outputs when we run the above code snippet are here below.
Missing: 1605
Missing: 0
Hey
Select features with RFE
# report which features were selected by RFE
from sklearn.datasets import make_classification
from sklearn.feature_selection import RFE
from sklearn.tree import DecisionTreeClassifier
# define dataset
X, y = make_classification(n_samples=1000, n_features=10, n_informative=5, n_redundant=5, random_state=1)
# define RFE
rfe = RFE(estimator=DecisionTreeClassifier(), n_features_to_select=5)
# fit RFE
rfe.fit(X, y)
# summarize all features
for i in range(X.shape[1]):
print(‘Column: %d, Selected=%s, Rank: %d’ % (i, rfe.support_[i], rfe.ranking_[i]))
The output is her below
Column: 0, Selected=False, Rank: 3
Column: 1, Selected=False, Rank: 5
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 2
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 4
Hello.
For lesson4 :
before normalization:
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
and after that:
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
I have made a scikit learn pipeline which will prepare the data starting with Data Normalization using Minmax scaler and then Feature selection using RFE. I have used hyperopt to optimize the result by using different kinds of parameters. But I am curious to try kbindiscretizer but I am working with financial data GDP which according to me shouldn’t be imputated and hence cannot use scikit learn PCA as it does not work with nan values.
Thank you for the feedback! Given that you work with financial data, you may find the following of interest:
https://machinelearningmastery.com/using-cnn-for-financial-time-series-prediction/
Hi, is the following statement correct?
“Normalization is required when the ranges among features are too disproportionate,
otherwise the feature with largest range of value would overlaps others in terms of its parameters.”
Hi Adilson…That is a correct statement!
I was looking for some point which help me to become a machine learning engineer. After reading your blog every picture become clear in my mind. Now I can design my career path. Thank you for best career advice. Your blog have fabulous information.
Lesson 1
Outliers treatment
Outliers are abnormal values in a dataset that don’t go with the regular distribution and have the potential to significantly distort a regression model, for example.
Missing value treatment
Occur when you don’t have data stored for certain variables. deletion or Imputation can be used to solve this problems. Imputation is used to replacing a missing value with another value based on a reasonable estimate.
Data Normalization
If one feature has very large values, it will dominate over other features when calculating the distance. So Normalization gives all features the same influence on the distance metric.
Normalization is a scaling technique in which values are shifted and rescaled so that they end up ranging between 0 and 1.
Great feedback Ismar! Keep up the great work!
Lesson 2
Before to aplicate the imputation, it was 1605 missing values as we can see at this line:
print(‘Missing: %d’ % sum(isnan(X).flatten()))
Missing: 1605
After imputation, we don’t have missing values anymore:
# define imputer
imputer = SimpleImputer(strategy=’mean’)
# fit on the dataset
imputer.fit(X)
print(‘Missing: %d’ % sum(isnan(Xtrans).flatten()))
Missing: 0
The parameter (strategy=’mean’) indicates that the missing values were filled with the average of the column values.
Lesson 3
Column: 0, Selected=False, Rank: 4
Column: 1, Selected=False, Rank: 5
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 2
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 3
Selected features:
Column 2, Column 3, Column 4, Column 6, Column 8. (Rank = 1)
Lesson 4
Before the normalization transform:
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
print(X.min())
-6.0167462574529615
print(X.max())
5.994383947517616
After the normalization transform:
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
print(X_norm.min())
0.0
print(X_norm.max())
1.0
for a model to perform better, what is the maximum number of feature is required. e.g. if I have a dataset with 50 features, what is the maximum number of feature will I retain using the feature selection method?
Hi Gabriel…You may find the following of interest:
https://machinelearningmastery.com/feature-selection-with-real-and-categorical-data/
Lesson 1 task: Data preparation algorithms and their purposes
Principal Component Analysis (PCA) discovers or reduces the dimensionality of a dataset
Forward feature selection is a method that reduces the input variable to your model by using only relevant data with an exclusion of noise
Thank you for your support and feedback Sope! Keep up the great work!
I get
Missing: 1605
Missing: 0
in Lesson 02: Fill Missing Values With Imputation
Hi Thinzar…Please specify the exact error message so that we may better assist you.
Day 3: Select Features With RFE
Column: 0, Selected=False, Rank: 5
Column: 1, Selected=False, Rank: 4
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=True, Rank: 1
Column: 4, Selected=True, Rank: 1
Column: 5, Selected=False, Rank: 6
Column: 6, Selected=True, Rank: 1
Column: 7, Selected=False, Rank: 3
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 2
Run the example and features 2, 3, 4, 6 and 8 are selected with Rank =1.
I also select 10 features from original dataset for horse colic dataset. I get
Column: 0, Selected=True, Rank: 1
Column: 1, Selected=False, Rank: 8
Column: 2, Selected=True, Rank: 1
Column: 3, Selected=False, Rank: 2
Column: 4, Selected=False, Rank: 9
Column: 5, Selected=True, Rank: 1
Column: 6, Selected=False, Rank: 7
Column: 7, Selected=False, Rank: 6
Column: 8, Selected=True, Rank: 1
Column: 9, Selected=False, Rank: 11
Column: 10, Selected=False, Rank: 10
Column: 11, Selected=False, Rank: 15
Column: 12, Selected=False, Rank: 16
Column: 13, Selected=False, Rank: 18
Column: 14, Selected=False, Rank: 14
Column: 15, Selected=True, Rank: 1
Column: 16, Selected=False, Rank: 5
Column: 17, Selected=False, Rank: 4
Column: 18, Selected=False, Rank: 3
Column: 19, Selected=True, Rank: 1
Column: 20, Selected=True, Rank: 1
Column: 21, Selected=True, Rank: 1
Column: 22, Selected=True, Rank: 1
Column: 23, Selected=True, Rank: 1
Column: 24, Selected=False, Rank: 12
Column: 25, Selected=False, Rank: 13
Column: 26, Selected=False, Rank: 17
Feature 0, 2, 5, 8, 15, 19, 20, 21, 22, 23 are selected with Rank = 1.
May I know this RFE algorithm is used for stock data prediction.
Hi Thinzar…Sorry, I cannot help you with machine learning for predicting the stock market, foreign exchange, or bitcoin prices.
I do not have a background or interest in finance.
I’m really skeptical.
I understand that unless you are operating at the highest level, that you will be eaten for lunch by the fees, by other algorithms, or by people that are operating at the highest level.
To get an idea of how brilliant some of these mathematicians are that apply machine learning to the stock market, I recommend reading this book:
The Man Who Solved the Market, 2019.
I love this quote from a recent Freakonomics podcast, asking about people picking stocks:
It’s a tax on smart people who don’t realize their propensity for doing stupid things.
— Barry Ritholtz, The Stupidest Thing You Can Do With Your Money, 2017.
I also understand that short-range movements of security prices (stocks) are a random walk and that the best that you can do is to use a persistence model.
I love this quote from the book “A Random Walk Down Wall Street“:
A random walk is one in which future steps or directions cannot be predicted on the basis of past history. When the term is applied to the stock market, it means that short-run changes in stock prices are unpredictable.
— Page 26, A Random Walk down Wall Street: The Time-tested Strategy for Successful Investing, 2016.
You can discover more about random walks here:
A Gentle Introduction to the Random Walk for Times Series Forecasting with Python
But we can be rich!?!
I remain really skeptical.
Maybe you know more about forecasting in finance than I do, and I wish you the best of luck.
What about finance data for self-study?
There is a wealth of financial data available.
If you are thinking of using this data to learn machine learning, rather than making money, then this sounds like an excellent idea.
Much of the data in finance is in the form of a time series. I recommend getting started with time series forecasting here:
Get Started With Time Series Forecasting
Permalink
Thanks for your response, suggestion and eBook’s links.!!
Day 4: I got
Before normalization:
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
Min: -6.0167462574529615 Max: 5.994383947517616
After normalization:
[[0.77608466 0.0239289 0.48251588 0.18352101 0.59830036]
[0.40400165 0.79590304 0.27369632 0.6331332 0.42104156]
[0.77065362 0.50132629 0.48207176 0.5076991 0.4293882 ]]
Min: 0.0 Max: 1.0
Day 5: Transformation Categories
I have got the result by following:
Before Transform:
[[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]]
After Transform:
[[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]]
Min: 0.0 Max: 1.0
Day 7: Dimensionality Reduction With PCA
The result is
Before Transform with PCS:
[[-0.53448246 0.93837451 0.38969914 0.0926655 1.70876508 1.14351305
-1.47034214 0.11857673 -2.72241741 0.2953565 ]
[-2.42280473 -1.02658758 -2.34792156 -0.82422408 0.59933419 -2.44832253
0.39750207 2.0265065 1.83374105 0.72430365]
[-1.83391794 -1.1946668 -0.73806871 1.50947233 1.78047734 0.58779205
-2.78506977 -0.04163788 -1.25227833 0.99373587]]
Min: -5.050910565749583 Max: 4.563133330755685
After Transform with PCA:
[[-1.64710578 -2.11683302 1.98256096]
[ 0.92840209 4.8294997 0.22727043]
[-3.83677757 0.32300714 0.11512801]]
Min: -7.732443010616129 Max: 9.958316592563877
Can I use PCA in Stock Predictive Analysis such as AAPL data from Yahoo Finance? I use this method for feature reduction. How can I use? Please give me a valuable suggestion and with sample code with these data.
Thanks for your response, suggestion and eBook’s links.!!
Lesson No.1
Three types of Data cleaning algorithms are:
1. Data Cleaning – This is establishing and making right mistakes or errors in data.
2. Feature Engineering – This is acquiring new variables from accessible data
3. Dimensionality Reduction – This is producing consolidated projections of the data
Thank you for your feedback Princess Leja!
Lesson 2 – Fill in missing values
Before imputation missing is 1605
After imputation missing is 0
Lesson 3
Column 0, Selected=False, Rank:5
Column 1, Selected=False, Rank:4
Column 2, Selected=True, Rank:1
Column 3, Selected=True, Rank:1
Column 4, Selected=True, Rank:1
Column 5, Selected=False, Rank:6
Column 6, Selected=True, Rank:1
Column 7, Selected=False, Rank:2
Column 8, Selected=True, Rank:1
Column 9, Selected=False, Rank:3
I am using Google Colab and it does not have copy, paste feature and had to key everything. Anyone there who can help on how to transfer my inputs here by copy, pasting?
Jason
Lesson 4
I posted Lesson 4 answers but did not see it uploaded. After posting it a second time, I get a reply that I have posted it. Please help.
Lesson 5
When I run the data it gives me this error “HTTPError: HTTP Error 404: Not Found”
That notwithstanding, I am going ahead with Lesson 6.
Many thanks for these tutorials.
Lesson 5
I have been able to run Lesson 5 and here it is:
Data before transform
[[“’40-49′” “‘premeno'” “’15-19′” “‘0-2′” “‘yes'” “‘3′” “‘right'”
“‘left_up'” “‘no'”]
[“’50-59′” “‘ge40′” “’15-19′” “‘0-2′” “‘no'” “‘1′” “‘right'” “‘central'”
“‘no'”]
[“’50-59′” “‘ge40′” “’35-39′” “‘0-2′” “‘no'” “‘2′” “‘left'” “‘left_low'”
“‘no'”]]
[ ]
# Data after transform through OneHotEncoding
[9]
0s
# define the one hot encoding transform
encoder = OneHotEncoder(sparse=False)
# fit and apply the transform to the input data
X_oe = encoder.fit_transform(X)
# summarize the transformed data
print(X_oe[:3, :])
output
[[0. 0. 1. 0. 0. 0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 1. 1. 0. 0. 0. 0. 0. 1. 0.]
[0. 0. 0. 1. 0. 0. 1. 0. 0. 0. 0. 0. 0. 0. 0. 1. 0. 0. 0. 0. 1. 0. 0. 0.
0. 0. 0. 1. 0. 0. 0. 1. 0. 1. 0. 0. 1. 0. 0. 0. 0. 1. 0.]]
Thanks.
Lesson 6
Raw Data before transform
[[ 2.39324489 -5.77732048 -0.59062319 -2.08095322 1.04707034]
[-0.45820294 1.94683482 -2.46471441 2.36590955 -0.73666725]
[ 2.35162422 -1.00061698 -0.5946091 1.12531096 -0.65267587]]
Data after transform
[[7. 0. 4. 1. 5.]
[4. 7. 2. 6. 4.]
[7. 5. 4. 5. 4.]]
set bins to 20 and strategy to Quantile
[[15. 0. 9. 3. 11.]
[ 8. 15. 5. 12. 8.]
[15. 10. 9. 10. 8.]]
I noticed changing the strategy did not affect the bins. setting bins to 20 was the same in the ordinal and quantile strategies.
Lesson 7
Raw data before transform
[[-0.53448246 0.93837451 0.38969914 0.0926655 1.70876508 1.14351305
-1.47034214 0.11857673 -2.72241741 0.2953565 ]
[-2.42280473 -1.02658758 -2.34792156 -0.82422408 0.59933419 -2.44832253
0.39750207 2.0265065 1.83374105 0.72430365]
[-1.83391794 -1.1946668 -0.73806871 1.50947233 1.78047734 0.58779205
-2.78506977 -0.04163788 -1.25227833 0.99373587]]
Data after transform
[[-1.64710578 -2.11683302 1.98256096]
[ 0.92840209 4.8294997 0.22727043]
[-3.83677757 0.32300714 0.11512801]]
Data after transform setting component to 4
[[-1.64710578e+00 -2.11683302e+00 1.98256096e+00 3.04107353e-15]
[ 9.28402085e-01 4.82949970e+00 2.27270432e-01 -8.63760114e-16]
[-3.83677757e+00 3.23007138e-01 1.15128013e-01 -1.29857567e-15]]
Data transform setting component to 5
output
[[-1.64710578e+00 -2.11683302e+00 1.98256096e+00 -5.20131106e-16
5.30225518e-16]
[ 9.28402085e-01 4.82949970e+00 2.27270432e-01 2.42512258e-15
-6.10368989e-16]
[-3.83677757e+00 3.23007138e-01 1.15128013e-01 2.27724993e-15
1.25985961e-15]]
Different data has been arrived at through PCA transform.
Jason,
This course has been a great course, an eye opener to what I have read before in text books and other materials. I longed everyday to tackle the lessons and here I have finished. With more practice, I am sure I will achieve my objective in data preparation.
My observations were very weak at the beginning of the lessons but they kept improving as I continued to the very end.
Thanks once again!
Exciting crash course! Data preparation is indeed the cornerstone of effective predictive modeling. Can’t wait to dive into the lessons and level up my skills. Thanks for sharing this valuable resource!
Exciting crash course! Data preparation is indeed the cornerstone of effective predictive modeling. Can’t wait to dive into the lessons and level up my skills.
Thank you stefan for your feedback and support! We greatly appreciat it!
I have a question ❓
In the second lesson, why did you use that method to split instead of the normal train_test_split() everyone uses?
And can you explain it in a sentence or two
The method used in the second lesson splits the data based on a specific condition, such as time or sequence order, which is crucial for tasks like time series analysis or ensuring data integrity when the order matters. This is different from train_test_split() which performs random sampling and may break temporal or contextual dependencies in the data.