Data is rarely clean and often you can have corrupt or missing values.
It is important to identify, mark and handle missing data when developing machine learning models in order to get the very best performance.
In this post you will discover how to handle missing values in your machine learning data using Weka.
After reading this post you will know:
- How to mark missing values in your dataset.
- How to remove data with missing values from your dataset.
- How to impute missing values.
Kick-start your project with my new book Machine Learning Mastery With Weka, including step-by-step tutorials and clear screenshots for all examples.
Let’s get started.

How To Handle Missing Data For Machine Learning in Weka
Photo by Peter Sitte, some rights reserved.
Predict the Onset of Diabetes
The problem used for this example is the Pima Indians onset of diabetes dataset.
It is a classification problem where each instance represents medical details for one patient and the task is to predict whether the patient will have an onset of diabetes within the next five years.
You can learn more about the dataset here:
You can also access this dataset in your Weka installation, under the data/ directory in the file called diabetes.arff.
Need more help with Weka for Machine Learning?
Take my free 14-day email course and discover how to use the platform step-by-step.
Click to sign-up and also get a free PDF Ebook version of the course.
Mark Missing Values
The Pima Indians dataset is a good basis for exploring missing data.
Some attributes such as blood pressure (pres) and Body Mass Index (mass) have values of zero, which are impossible. These are examples of corrupt or missing data that must be marked manually.
You can mark missing values in Weka using the NumericalCleaner filter. The recipe below shows you how to use this filter to mark the 11 missing values on the Body Mass Index (mass) attribute.
1. Open the Weka Explorer.
2. Load the Pima Indians onset of diabetes dataset.
3. Click the “Choose” button for the Filter and select NumericalCleaner, it us under unsupervized.attribute.NumericalCleaner.
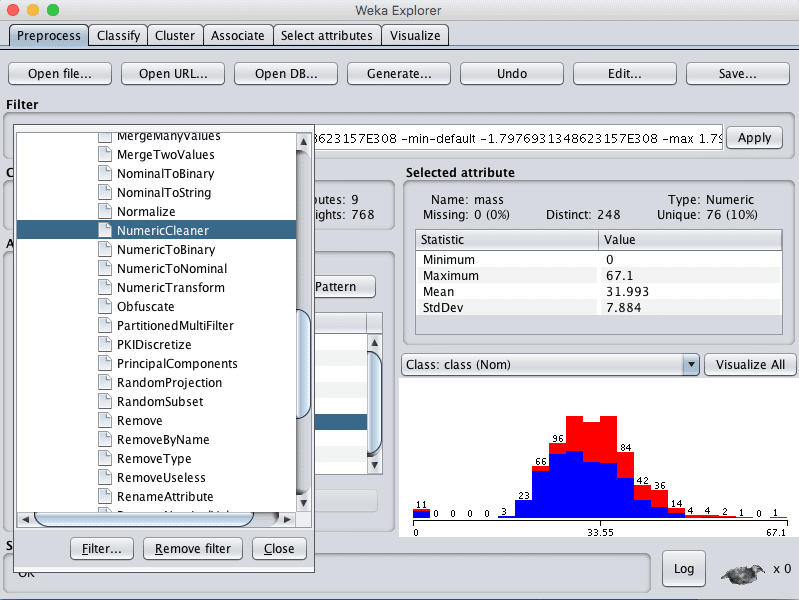
Weka Select NumericCleaner Data Filter
4. Click on the filter to configure it.
5. Set the attributeIndicies to 6, the index of the mass attribute.
6. Set minThreshold to 0.1E-8 (close to zero), which is the minimum value allowed for the attribute.
7. Set minDefault to NaN, which is unknown and will replace values below the threshold.
8. Click the “OK” button on the filter configuration.
9. Click the “Apply” button to apply the filter.
Click “mass” in the “attributes” pane and review the details of the “selected attribute”. Notice that the 11 attribute values that were formally set to 0 are not marked as Missing.
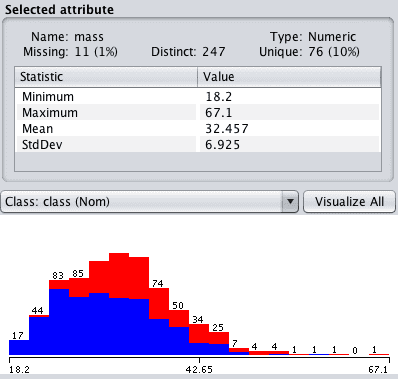
Weka Missing Data Marked
In this example we marked values below a threshold as missing.
You could just as easily mark them with a specific numerical value. You could also mark values missing between a upper and lower range of values.
Next, let’s look at how we can remove instances with missing values from our dataset.
Remove Missing Data
Now that you know how to mark missing values in your data, you need to learn how to handle them.
A simple way to handle missing data is to remove those instances that have one or more missing values.
You can do this in Weka using the RemoveWithValues filter.
Continuing on from the above recipe to mark missing values, you can remove missing values as follows:
1. Click the “Choose” button for the Filter and select RemoveWithValues, it us under unsupervized.instance.RemoveWithValues.
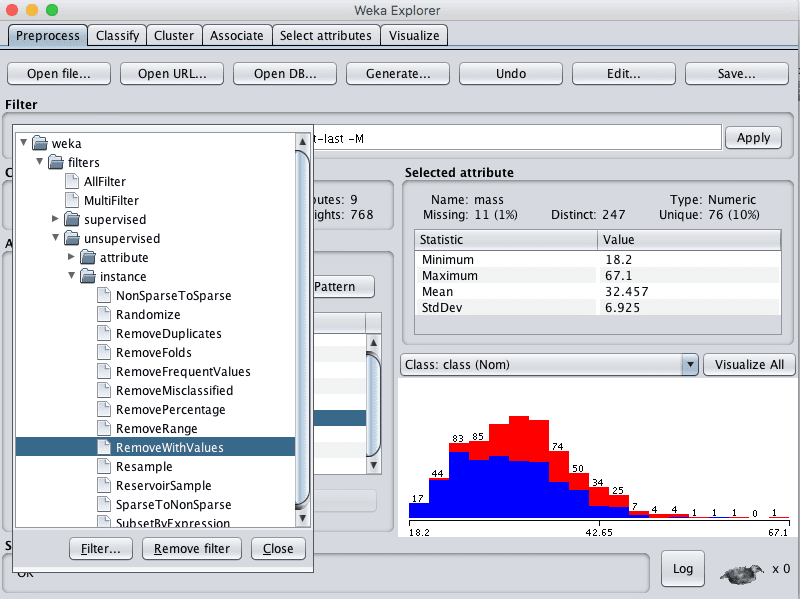
Weka Select RemoveWithValues Data Filter
2. Click on the filter to configure it.
3. Set the attributeIndicies to 6, the index of the mass attribute.
4. Set matchMissingValues to “True”.
5. Click the “OK” button to use the configuration for the filter.
6. Click the “Apply” button to apply the filter.
Click “mass” in the “attributes” section and review the details of the “selected attribute”.
Notice that the 11 attribute values that were marked Missing have been removed from the dataset.
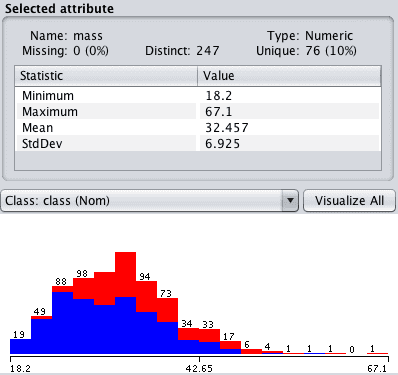
Weka Missing Values Removed
Note, you can undo this operation by clicking the “Undo” button.
Impute Missing Values
Instances with missing values do not have to be removed, you can replace the missing values with some other value.
This is called imputing missing values.
It is common to impute missing values with the mean of the numerical distribution. You can do this easily in Weka using the ReplaceMissingValues filter.
Continuing on from the first recipe above to mark missing values, you can impute the missing values as follows:
1. Click the “Choose” button for the Filter and select ReplaceMissingValues, it us under unsupervized.attribute.ReplaceMissingValues.
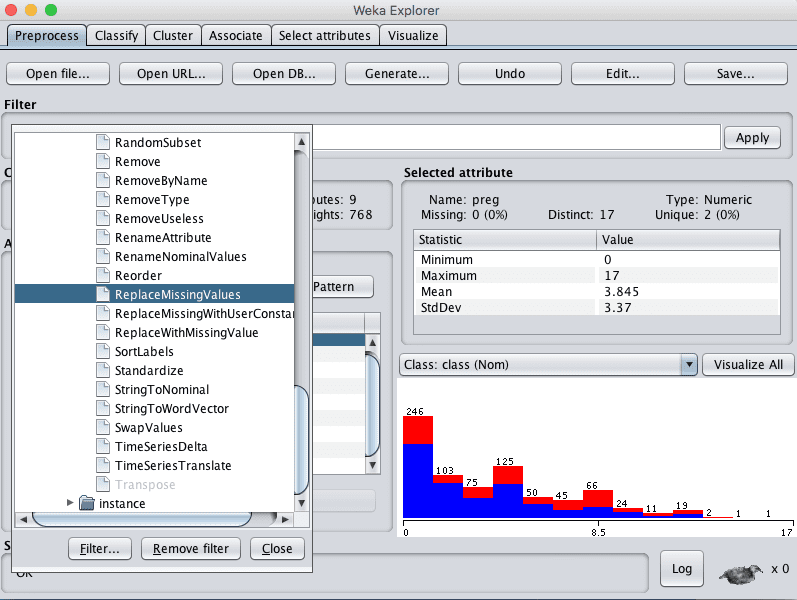
Weka ReplaceMissingValues Data Filter
2. Click the “Apply” button to apply the filter to your dataset.
Click “mass” in the “attributes” section and review the details of the “selected attribute”.
Notice that the 11 attribute values that were marked Missing have been set to the mean value of the distribution.
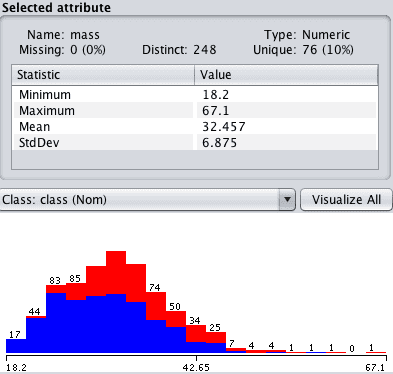
Weka Imputed Values
Summary
In this post you discovered how you can handle missing data in your machine learning dataset using Weka.
Specifically, you learned:
- How to mark corrupt values as missing in your dataset.
- How to remove instances with missing values from your dataset.
- How to impute mean values for missing values in your dataset.
Do you have any questions about missing data or about this tutorial? Ask your questions in the comments below and I will do my best to answer.




I am to ask a question when the data having missing numeric values(blank) is subjected to classification say Multilayer Perceptron, what value is taken for calculation by the WEKA.
Hi Ramesh, you can pre-process your data to impute missing values with a mean or other constant value.
Hi, thank you for your instructions. Now that I have removed the instances with a particular missing attribute value, I would like to train with the remaining data to predict the missing values I just removed, how do I do that?
Hi Enrique, my best advice would be to save a copy of your transformed dataset.
You can then load it in the future and use it in your Exploration and Experiments.
hi sir, thanks for such a fruitful demo. I wanted to ask if we can use KNN imputation with Weka. Or any other imputation based on nearest neighbors
I’m not sure off-hand Shah.
But we are not even sure, which machine learning technique is used by WEKA for imputation or replacing values.
The ReplaceMissingValues filter replaces values with the mean or mode for real and nominal values respectively.
You can learn more here:
http://weka.sourceforge.net/doc.stable/weka/filters/unsupervised/attribute/ReplaceMissingValues.html
I want to predict an attribute which is present in training dataset but absent in test dataset. Even if it is present, I tried using that brand instances as Null or missing values. I am using randomforest and it gats wrapped in inputmapped classifier and although it executes but it predicts only ? marks(i.e,. Null).
the dataset that I am using contains importer names,unit,currency,duty values,brand specs(in text,some in numeric or real). The value that I am trying to predict is a text(nominal). Most of the text data can be easily lablled as binary data type.
Hi
Please send me a notification whenever a new post or newsletter is added.
Regards,
Mohammed
You can access the RSS feed here:
https://machinelearningmastery.com/feed/
Hi Jason,
Nice article, however, I feel like you missed a good point of explaining which step on missing data should be taken and so I am a tad confused.
At what point would I prefer to remove samples that contain missing information over the choice of Imputing them?
The removal of samples also leads me to the question of what is the minimum threshold of samples that you should retain in your dataset for it to still be representative of the objective you are attempting to carry out?
We cannot know which method will be “best” for a given problem.
Try multiple approaches systematically and select the approach that gives the best predictive models on your problem.
The amount of data require is also dependent on the complexity of the problem.
Sorry, no easy answers. Lots of experimentation and trial-and-error required in applied machine learning. Develop a robust test harness!
In weka tool when i click on tree under classification it is not showing J48 How do i add it ?
Hello, in case of nominal attribute, how could I remove missing instances (e.g. flagged with “?”)? I didn’t find a filter. Applying the “Remove with values” should be able but it didn’t work. Thank you.
Are you able to use the pandas functions demonstrated in the tutorial?
I believe the type of the column should not matter for the dataframe functions used.
Hi Jason,
I wanna ask about missing data. I know sometimes missing data are too important for us to completely remove them from analysis. For instance I have a data set that has 30% of he overall data missing. I am trying to do a time series prediction and data are missing consistently for Saturdasy and Sundays for 5 years. I am employing a Kalman smoothing method to fill in the spaces for those two days…
Do you think this method may be relatively okay? or do you think I should remove the rows with missing values?
Try different methods and see what works best for your problem. It’s hard (intractable!) to say what will work best a priori.
could you please mention a few evaluation (if any) techniques to check how well my method of replacing missing data worked? How do I know which is the best method since there’s no way I could ever know what this missing data are.
Thank you in advance.
Yes, focus on the predictive skill of models trained with the data on which you performed the imputation.
Hi Jason,
Good article 🙂
I have some doubts. I generate my model, im missing atributes i insert the symbol “?”
Do you know how Weka works when the value of attributes missing is “?”
Can you give-me one paper that talk about this?
Best Regards
Does this tutorial work for you?
I am so sorry Jason, my Technique is different …
I do not want to fill in the values, I want to leave them blank …
Do you know how the algorithm works when I do this?
Thanks again 🙂
Hello Jason 🙂
I learn 🙂 …
If you use MLP the missing values are ignored by setting the input value to zero.
If you use SMOreg or Linear Regression the missing values are for mean/mode imputation.
Nice.
hi jason i am doing prediction with naive bayes and i have a lot of missing values in my dataset ,for this moment i use replace missing values and then pre-process and predict is it mayby better if i delete the attributes with 95% of missing values?
I would recommend trying a suite of different approaches and see what works best for your specific data.
i don’t quite understand what is the difference between handling the missing values the way you did in the article as opposed to only using the “replace missing values” filter ?
I do exactly that.
i am faced problem
i don have a missing value but the sign (?) is appear when apply in weka
Sorry, I don’t follow, what do you mean exactly?
How do you handle the problem of question marks (?) in your detailed accuracy table when there are no missing values in your data set?
Hi Greg…Additional ideas can be found here:
https://machinelearningmastery.com/handle-missing-data-python/
when we are trying apply unsupervised attribute replacemissingvalues filter in weka ,i am able to get filled up miising value as output
Well done!
Hi, I have trouble in handling missing value.
The problem is regression, not classification. Is there any way to use Machine Learning to handle missing values?
I am stumped.
Thank you
Yes, you can use the mean value as a good starting point.
Hi Jason. Informative. Thanks. I wanted to ask how does the default settings of Weka (without filters) handle missing values? Are training/testing examples 1) Case-wise excluded (Any example with any missing predictor excluded) 2) Imputed from mean/median of the available feature attributes?
It uses the data as is I believe, e.g. “missing” values are treated as values.
Sorry didnt quite get it. Appreciate more clarity on it. How can “missing” value be treated as value? It is “missing” as implied
In some cases a missing value may be marked with a value, e.g. “none” or “nan”. A method like a decision tree can then make split decisions based on this value.
If the data is actually missing, then the row or value may simply be ignored by the algorithm, although it depends on the algorithm.
Hi!
Im trying to perform imputation to missing data (initially to test the process this missing data is generated by randomly removing data from attributes in a dataSet). I want to do this by using an ANN. So my question is, if I train the ANN in weka setting the class to the actual class, can then use this ANN to predict the value of the missing one by then setting the class of the instance in this missing attribute? I´m trying to do this but I get a “no input instance format defined” error.
If the answer is no, then, Do I have to train the ANN for each missing attribute, setting the data class in this specific attribute to this that is going to be predicted and then using the instance as an input once the ANN (or any other classificator) is arreado trained. For me, this means that there is a need to perform one training (one machine, in fact) for each missing attribute.
I think that what I’m asking is to perform regression, but I do not found how to do this in weka.
The model would be trained to predict the missing value not the class value. E.g. an imputation method.
Yes, a model will be trained for each feature I believe.
Thnks a lot. I have finally decided to train one “Method” for each attribute in a batch single process to later perform a quick imputation.
Really appreciate your answer!
done it successfully
Well done!
How can we know it uses mean or the mode to replace values?
Hi ydil…In Weka, handling missing values is typically done using the
ReplaceMissingValuesfilter, which can replace missing values with the mean (for numeric attributes) or the mode (for nominal attributes).Here’s how you can check or change the default behavior in Weka:
1. **Open Weka Explorer:**
– Start Weka and go to the Explorer.
2. **Load Your Dataset:**
– Load the dataset that you want to preprocess.
3. **Select the Filter:**
– Go to the “Preprocess” tab.
– Click on the “Choose” button under the “Filter” section.
– Select
weka.filters.unsupervised.attribute.ReplaceMissingValues.4. **Check Filter Options:**
– After selecting the filter, you can click on the filter name (e.g.,
ReplaceMissingValues) to view and configure its options. However, this particular filter does not provide options for changing the default behavior (mean for numeric and mode for nominal) since it’s built-in.5. **Apply the Filter:**
– Apply the filter to your dataset by clicking the “Apply” button.
6. **Verify the Changes:**
– Once applied, you can check the transformed dataset to see how the missing values have been handled.
**Note:** If you want more control over how missing values are replaced (for example, using median instead of mean), you might need to preprocess your data manually using other filters or custom scripts.
### Using Weka’s Command Line Interface
If you’re using Weka from the command line, you can apply the
ReplaceMissingValuesfilter with the following command:sh
java -cp weka.jar weka.filters.unsupervised.attribute.ReplaceMissingValues -i input.arff -o output.arff
In this command:
–
-i input.arffspecifies the input file.–
-o output.arffspecifies the output file with missing values replaced.### Example in Weka GUI
1. **Loading Data:**
– Click “Open file…” and select your dataset (e.g.,
data.arff).2. **Applying Filter:**
– Choose the
ReplaceMissingValuesfilter.– Click “Apply”.
3. **Viewing Results:**
– Check the resulting dataset to confirm the changes.
By default, Weka’s
ReplaceMissingValuesfilter uses the mean for numeric attributes and the mode for nominal attributes, and this behavior is generally not configurable directly within the filter settings. If you need a different approach, consider using other preprocessing steps or external data preparation tools before importing the data into Weka.