Many machine learning algorithms expect data to be scaled consistently.
There are two popular methods that you should consider when scaling your data for machine learning.
In this tutorial, you will discover how you can rescale your data for machine learning. After reading this tutorial you will know:
- How to normalize your data from scratch.
- How to standardize your data from scratch.
- When to normalize as opposed to standardize data.
Kick-start your project with my new book Machine Learning Algorithms From Scratch, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.
- Update Feb/2018: Fixed minor typo in min/max code example.
- Update Mar/2018: Added alternate link to download the dataset as the original appears to have been taken down.
- Update Aug/2018: Tested and updated to work with Python 3.6.

How To Prepare Machine Learning Data From Scratch With Python
Photo by Ondra Chotovinsky, some rights reserved.
Description
Many machine learning algorithms expect the scale of the input and even the output data to be equivalent.
It can help in methods that weight inputs in order to make a prediction, such as in linear regression and logistic regression.
It is practically required in methods that combine weighted inputs in complex ways such as in artificial neural networks and deep learning.
In this tutorial, we are going to practice rescaling one standard machine learning dataset in CSV format.
Specifically, the Pima Indians dataset. It contains 768 rows and 9 columns. All of the values in the file are numeric, specifically floating point values. We will learn how to load the file first, then later how to convert the loaded strings to numeric values.
Tutorial
This tutorial is divided into 3 parts:
- Normalize Data.
- Standardize Data.
- When to Normalize and Standardize.
These steps will provide the foundations you need to handle scaling your own data.
1. Normalize Data
Normalization can refer to different techniques depending on context.
Here, we use normalization to refer to rescaling an input variable to the range between 0 and 1.
Normalization requires that you know the minimum and maximum values for each attribute.
This can be estimated from training data or specified directly if you have deep knowledge of the problem domain.
You can easily estimate the minimum and maximum values for each attribute in a dataset by enumerating through the values.
The snippet of code below defines the dataset_minmax() function that calculates the min and max value for each attribute in a dataset, then returns an array of these minimum and maximum values.
|
1 2 3 4 5 6 7 8 9 |
# Find the min and max values for each column def dataset_minmax(dataset): minmax = list() for i in range(len(dataset[0])): col_values = [row[i] for row in dataset] value_min = min(col_values) value_max = max(col_values) minmax.append([value_min, value_max]) return minmax |
We can contrive a small dataset for testing as follows:
|
1 2 3 |
x1 x2 50 30 20 90 |
With this contrived dataset, we can test our function for calculating the min and max for each column.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 |
# Find the min and max values for each column def dataset_minmax(dataset): minmax = list() for i in range(len(dataset[0])): col_values = [row[i] for row in dataset] value_min = min(col_values) value_max = max(col_values) minmax.append([value_min, value_max]) return minmax # Contrive small dataset dataset = [[50, 30], [20, 90]] print(dataset) # Calculate min and max for each column minmax = dataset_minmax(dataset) print(minmax) |
Running the example produces the following output.
First, the dataset is printed in a list of lists format, then the min and max for each column is printed in the format column1: min,max and column2: min,max.
For example:
|
1 2 |
[[50, 30], [20, 90]] [[20, 50], [30, 90]] |
Once we have estimates of the maximum and minimum allowed values for each column, we can now normalize the raw data to the range 0 and 1.
The calculation to normalize a single value for a column is:
|
1 |
scaled_value = (value - min) / (max - min) |
Below is an implementation of this in a function called normalize_dataset() that normalizes values in each column of a provided dataset.
|
1 2 3 4 5 |
# Rescale dataset columns to the range 0-1 def normalize_dataset(dataset, minmax): for row in dataset: for i in range(len(row)): row[i] = (row[i] - minmax[i][0]) / (minmax[i][1] - minmax[i][0]) |
We can tie this function together with the dataset_minmax() function and normalize the contrived dataset.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 |
# Find the min and max values for each column def dataset_minmax(dataset): minmax = list() for i in range(len(dataset[0])): col_values = [row[i] for row in dataset] value_min = min(col_values) value_max = max(col_values) minmax.append([value_min, value_max]) return minmax # Rescale dataset columns to the range 0-1 def normalize_dataset(dataset, minmax): for row in dataset: for i in range(len(row)): row[i] = (row[i] - minmax[i][0]) / (minmax[i][1] - minmax[i][0]) # Contrive small dataset dataset = [[50, 30], [20, 90]] print(dataset) # Calculate min and max for each column minmax = dataset_minmax(dataset) print(minmax) # Normalize columns normalize_dataset(dataset, minmax) print(dataset) |
Running this example prints the output below, including the normalized dataset.
|
1 2 3 |
[[50, 30], [20, 90]] [[20, 50], [30, 90]] [[1, 0], [0, 1]] |
We can combine this code with code for loading a CSV dataset and load and normalize the Pima Indians diabetes dataset.
Download the Pima Indians dataset and place it in your current directory with the name pima-indians-diabetes.csv.
Open the file and delete any empty lines at the bottom.
The example first loads the dataset and converts the values for each column from string to floating point values. The minimum and maximum values for each column are estimated from the dataset, and finally, the values in the dataset are normalized.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 |
from csv import reader # Load a CSV file def load_csv(filename): file = open(filename, "rb") lines = reader(file) dataset = list(lines) return dataset # Convert string column to float def str_column_to_float(dataset, column): for row in dataset: row[column] = float(row[column].strip()) # Find the min and max values for each column def dataset_minmax(dataset): minmax = list() for i in range(len(dataset[0])): col_values = [row[i] for row in dataset] value_min = min(col_values) value_max = max(col_values) minmax.append([value_min, value_max]) return minmax # Rescale dataset columns to the range 0-1 def normalize_dataset(dataset, minmax): for row in dataset: for i in range(len(row)): row[i] = (row[i] - minmax[i][0]) / (minmax[i][1] - minmax[i][0]) # Load pima-indians-diabetes dataset filename = 'pima-indians-diabetes.csv' dataset = load_csv(filename) print('Loaded data file {0} with {1} rows and {2} columns').format(filename, len(dataset), len(dataset[0])) # convert string columns to float for i in range(len(dataset[0])): str_column_to_float(dataset, i) print(dataset[0]) # Calculate min and max for each column minmax = dataset_minmax(dataset) # Normalize columns normalize_dataset(dataset, minmax) print(dataset[0]) |
Running the example produces the output below.
The first record from the dataset is printed before and after normalization, showing the effect of the scaling.
|
1 2 3 |
Loaded data file pima-indians-diabetes.csv with 768 rows and 9 columns [6.0, 148.0, 72.0, 35.0, 0.0, 33.6, 0.627, 50.0, 1.0] [0.35294117647058826, 0.7437185929648241, 0.5901639344262295, 0.35353535353535354, 0.0, 0.5007451564828614, 0.23441502988898377, 0.48333333333333334, 1.0] |
2. Standardize Data
Standardization is a rescaling technique that refers to centering the distribution of the data on the value 0 and the standard deviation to the value 1.
Together, the mean and the standard deviation can be used to summarize a normal distribution, also called the Gaussian distribution or bell curve.
It requires that the mean and standard deviation of the values for each column be known prior to scaling. As with normalizing above, we can estimate these values from training data, or use domain knowledge to specify their values.
Let’s start with creating functions to estimate the mean and standard deviation statistics for each column from a dataset.
The mean describes the middle or central tendency for a collection of numbers. The mean for a column is calculated as the sum of all values for a column divided by the total number of values.
|
1 |
mean = sum(values) / total_values |
The function below named column_means() calculates the mean values for each column in the dataset.
|
1 2 3 4 5 6 7 |
# calculate column means def column_means(dataset): means = [0 for i in range(len(dataset[0]))] for i in range(len(dataset[0])): col_values = [row[i] for row in dataset] means[i] = sum(col_values) / float(len(dataset)) return means |
The standard deviation describes the average spread of values from the mean. It can be calculated as the square root of the sum of the squared difference between each value and the mean and dividing by the number of values minus 1.
|
1 |
standard deviation = sqrt( (value_i - mean)^2 / (total_values-1)) |
The function below named column_stdevs() calculates the standard deviation of values for each column in the dataset and assumes the means have already been calculated.
|
1 2 3 4 5 6 7 8 |
# calculate column standard deviations def column_stdevs(dataset, means): stdevs = [0 for i in range(len(dataset[0]))] for i in range(len(dataset[0])): variance = [pow(row[i]-means[i], 2) for row in dataset] stdevs[i] = sum(variance) stdevs = [sqrt(x/(float(len(dataset)-1))) for x in stdevs] return stdevs |
Again, we can contrive a small dataset to demonstrate the estimate of the mean and standard deviation from a dataset.
|
1 2 3 4 |
x1 x2 50 30 20 90 30 50 |
Using an excel spreadsheet, we can estimate the mean and standard deviation for each column as follows:
|
1 2 3 |
x1 x2 mean 33.3 56.6 stdev 15.27 30.55 |
Using the contrived dataset, we can estimate the summary statistics.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 |
from math import sqrt # calculate column means def column_means(dataset): means = [0 for i in range(len(dataset[0]))] for i in range(len(dataset[0])): col_values = [row[i] for row in dataset] means[i] = sum(col_values) / float(len(dataset)) return means # calculate column standard deviations def column_stdevs(dataset, means): stdevs = [0 for i in range(len(dataset[0]))] for i in range(len(dataset[0])): variance = [pow(row[i]-means[i], 2) for row in dataset] stdevs[i] = sum(variance) stdevs = [sqrt(x/(float(len(dataset)-1))) for x in stdevs] return stdevs # Standardize dataset dataset = [[50, 30], [20, 90], [30, 50]] print(dataset) # Estimate mean and standard deviation means = column_means(dataset) stdevs = column_stdevs(dataset, means) print(means) print(stdevs) |
Executing the example provides the following output, matching the numbers calculated in the spreadsheet.
|
1 2 3 |
[[50, 30], [20, 90], [30, 50]] [33.333333333333336, 56.666666666666664] [15.275252316519467, 30.550504633038933] |
Once the summary statistics are calculated, we can easily standardize the values in each column.
The calculation to standardize a given value is as follows:
|
1 |
standardized_value = (value - mean) / stdev |
Below is a function named standardize_dataset() that implements this equation
|
1 2 3 4 5 |
# standardize dataset def standardize_dataset(dataset, means, stdevs): for row in dataset: for i in range(len(row)): row[i] = (row[i] - means[i]) / stdevs[i] |
Combining this with the functions to estimate the mean and standard deviation summary statistics, we can standardize our contrived dataset.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 |
from math import sqrt # calculate column means def column_means(dataset): means = [0 for i in range(len(dataset[0]))] for i in range(len(dataset[0])): col_values = [row[i] for row in dataset] means[i] = sum(col_values) / float(len(dataset)) return means # calculate column standard deviations def column_stdevs(dataset, means): stdevs = [0 for i in range(len(dataset[0]))] for i in range(len(dataset[0])): variance = [pow(row[i]-means[i], 2) for row in dataset] stdevs[i] = sum(variance) stdevs = [sqrt(x/(float(len(dataset)-1))) for x in stdevs] return stdevs # standardize dataset def standardize_dataset(dataset, means, stdevs): for row in dataset: for i in range(len(row)): row[i] = (row[i] - means[i]) / stdevs[i] # Standardize dataset dataset = [[50, 30], [20, 90], [30, 50]] print(dataset) # Estimate mean and standard deviation means = column_means(dataset) stdevs = column_stdevs(dataset, means) print(means) print(stdevs) # standardize dataset standardize_dataset(dataset, means, stdevs) print(dataset) |
Executing this example produces the following output, showing standardized values for the contrived dataset.
|
1 2 3 4 |
[[50, 30], [20, 90], [30, 50]] [33.333333333333336, 56.666666666666664] [15.275252316519467, 30.550504633038933] [[1.0910894511799618, -0.8728715609439694], [-0.8728715609439697, 1.091089451179962], [-0.21821789023599253, -0.2182178902359923]] |
Again, we can demonstrate the standardization of a machine learning dataset.
The example below demonstrate how to load and standardize the Pima Indians diabetes dataset, assumed to be in the current working directory as in the previous normalization example.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 |
from csv import reader from math import sqrt # Load a CSV file def load_csv(filename): file = open(filename, "rb") lines = reader(file) dataset = list(lines) return dataset # Convert string column to float def str_column_to_float(dataset, column): for row in dataset: row[column] = float(row[column].strip()) # calculate column means def column_means(dataset): means = [0 for i in range(len(dataset[0]))] for i in range(len(dataset[0])): col_values = [row[i] for row in dataset] means[i] = sum(col_values) / float(len(dataset)) return means # calculate column standard deviations def column_stdevs(dataset, means): stdevs = [0 for i in range(len(dataset[0]))] for i in range(len(dataset[0])): variance = [pow(row[i]-means[i], 2) for row in dataset] stdevs[i] = sum(variance) stdevs = [sqrt(x/(float(len(dataset)-1))) for x in stdevs] return stdevs # standardize dataset def standardize_dataset(dataset, means, stdevs): for row in dataset: for i in range(len(row)): row[i] = (row[i] - means[i]) / stdevs[i] # Load pima-indians-diabetes dataset filename = 'pima-indians-diabetes.csv' dataset = load_csv(filename) print('Loaded data file {0} with {1} rows and {2} columns').format(filename, len(dataset), len(dataset[0])) # convert string columns to float for i in range(len(dataset[0])): str_column_to_float(dataset, i) print(dataset[0]) # Estimate mean and standard deviation means = column_means(dataset) stdevs = column_stdevs(dataset, means) # standardize dataset standardize_dataset(dataset, means, stdevs) print(dataset[0]) |
Running the example prints the first row of the dataset, first in a raw format as loaded, and then standardized which allows us to see the difference for comparison.
|
1 2 3 |
Loaded data file pima-indians-diabetes.csv with 768 rows and 9 columns [6.0, 148.0, 72.0, 35.0, 0.0, 33.6, 0.627, 50.0, 1.0] [0.6395304921176576, 0.8477713205896718, 0.14954329852954296, 0.9066790623472505, -0.692439324724129, 0.2038799072674717, 0.468186870229798, 1.4250667195933604, 1.3650063669598067] |
3. When to Normalize and Standardize
Standardization is a scaling technique that assumes your data conforms to a normal distribution.
If a given data attribute is normal or close to normal, this is probably the scaling method to use.
It is good practice to record the summary statistics used in the standardization process, so that you can apply them when standardizing data in the future that you may want to use with your model.
Normalization is a scaling technique that does not assume any specific distribution.
If your data is not normally distributed, consider normalizing it prior to applying your machine learning algorithm.
It is good practice to record the minimum and maximum values for each column used in the normalization process, again, in case you need to normalize new data in the future to be used with your model.
Extensions
There are many other data transforms you could apply.
The idea of data transforms is to best expose the structure of your problem in your data to the learning algorithm.
It may not be clear what transforms are required upfront. A combination of trial and error and exploratory data analysis (plots and stats) can help tease out what may work.
Below are some additional transforms you may want to consider researching and implementing:
- Normalization that permits a configurable range, such as -1 to 1 and more.
- Standardization that permits a configurable spread, such as 1, 2 or more standard deviations from the mean.
- Exponential transforms such as logarithm, square root and exponents.
- Power transforms such as box-cox for fixing the skew in normally distributed data.
Review
In this tutorial, you discovered how to rescale your data for machine learning from scratch.
Specifically, you learned:
- How to normalize data from scratch.
- How to standardize data from scratch.
- When to use normalization or standardization on your data.
Do you have any questions about scaling your data or about this post?
Ask your question in the comments below and I will do my best to answer.


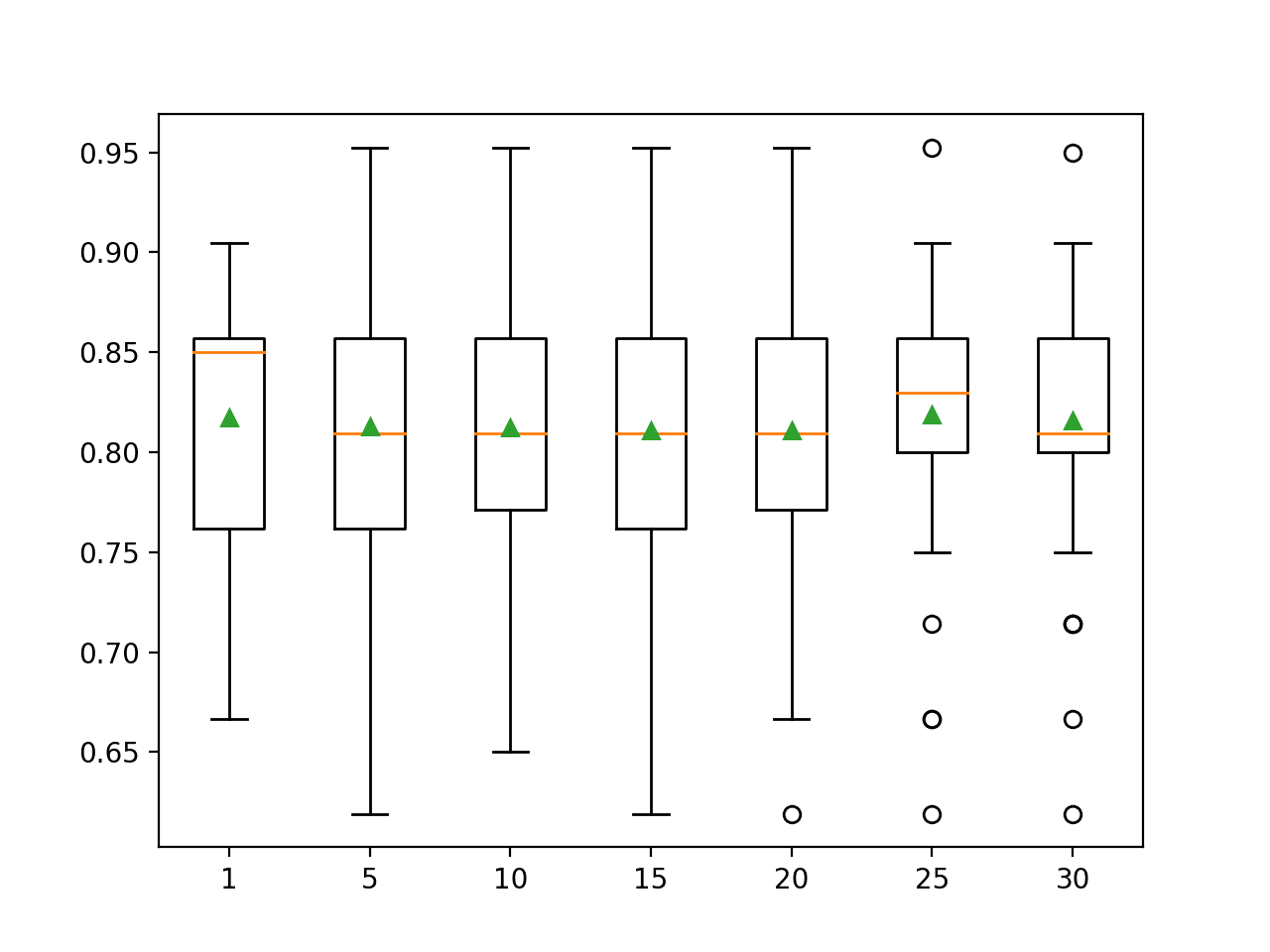

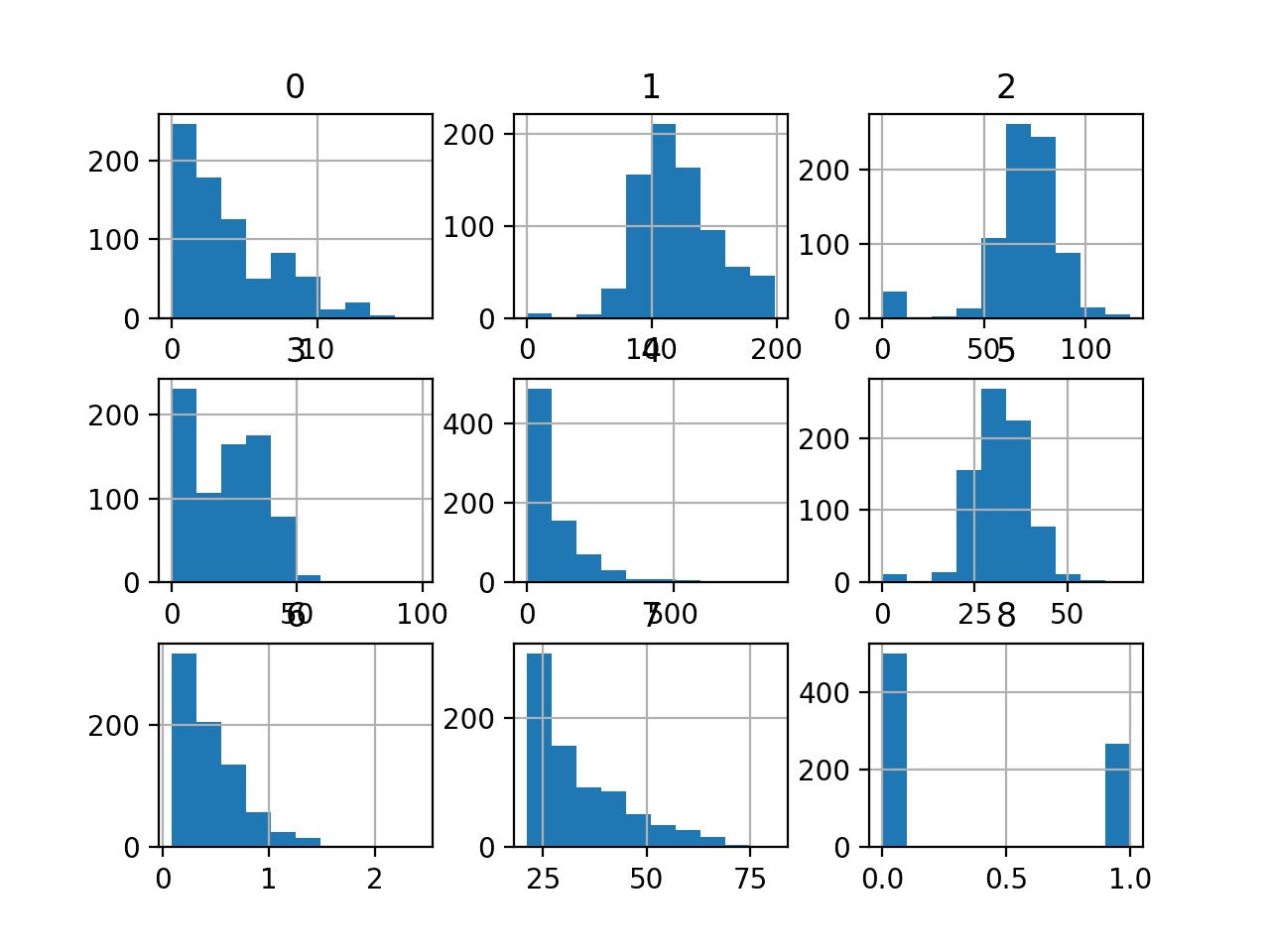


I am wanting to use transaction totals by month as a feature. I plan to use Xgboost and from what I have read it is better to use binary variables. Will normalizing work or should bin the totals into deciles so I can make them binary?
Hi Daniel,
I think you could try a model with transaction totals as-is, binned values, and with binary values (above a threshold or something).
I don’t know the problem you’re working on, but generally it is good practice to try a few different framings of a problem and see which works best.
Thank you for your time
Hi, it’s a very good tutorial. But I still have a doubt. If I wanted to use scikit function to normalize my data and then print it in order to verufy if it really worked, how should I proceed?
I’ve tried something like
scaler = MinMaxScaler(feature_range=(0, 1))
dataset = scaler.fit_transform(dataset)
plt.plot(dataset)
plt.show()
but it doesn’t work. It doesn’t effect the data at all
The code looks good.
Try printing the transformed array rather than plotting.
Hi Jason,
If I have two model (classification and regression), and each of them output log loss and absolute error.
So, if I want to combine the output errors, do I have to normalize both errors first before performing the addition? If yes, can normalization formula above be used to perform normalization on both errors?
Log loss on a regression problem does not make sense.
Sorry, my previous post might confused you. Actually, in my case classification problem outputs log loss error function while regression problem outputs absolute error function (MSE, MAE, R2, etc).
So I want to sum up both errors (from classification and regression problem), and need to normalize them first. scikit-learn provides Normalized parameter in log loss function which it will return the mean loss per sample. However regression loss function such as RMSE does not have Normalization parameter which the mean loss output needs to be normalized manually.
Can you suggest how can I normalized the final output of loss error in regression problem?
Your opinion on this matter is highly appreciated.
Why would you want to normalize the error?
It might be better to interpet the error score than to transform it. For example RMSE is in the units of the output variable directly.
What is the scale formula for -1 -> 1?
Good question, you can use:
Here’s a snippet to demo the function:
Hello Jason,
In your post you recommend using standardization when the data is normally distributed and normalization when the data is not normally distributed. Do you know why that is? I think I’ve seen standardization described as a z-score elsewhere, but I don’t understand why computing this value for non-normally distributed data isn’t recommended as inputs to machine learning algorithms. Couldn’t a machine learning algorithm still derive value from these inputs even with non-normally distributed data?
Standardization shifts data to have a zero mean and unit standard deviation. For data that is not Gaussian, this transform would not make sense – the data would not be centered and there is no standard deviation for non Gaussian data.
Good morning Jason,
In the context of data transformation I wonder how to quickly check which method is suitable for which ML algorithm. Some algorithms require inputs in the range [0.0,1.0] , others [-1.0, 1.0] or the standardization. When I read the description of algorithms of sklearn at: https://scikit-learn.org/stable/supervised_learning.html#supervised-learning I find this information missing.
Do you recommend some assembly source for this topic? Or rather data transform method shall be seeked in research articles or spot-checked each time new data/problem is approached?
Regards!
And worse, sometimes you get best results by violating the rules/assumptions/expectations.
The best approach is to test different transforms for an algorithm. That’s my recommendation.
This is the interesting feedback.
Thank you for sharing!
Glad it helped.
It is the basis of the “spot-check” approach that I recommend:
https://machinelearningmastery.com/start-here/#process
Hello Jason, If I want to get the real value, how to denormalize?
Perhaps use the sklearn scale objects, then afterward use the inverse transform.
See an example here:
https://machinelearningmastery.com/machine-learning-data-transforms-for-time-series-forecasting/
Hello Jason, Thanks for such great tutorials. I learnt a lot from them.
Currently I am working on time- series forecast for energy consumption with LSTM network. I am facing a problem. As I start the training, sometimes I get the right results, and I can see my loss getting low epoch by epoch. But I run the same model again and sometimes I get nan loss as soon as my training starts or sometimes nan loss comes after the code has run for a few epochs. Strange thing is that I have also got nan loss all of a sudden even after my loss starts converging to a very low value. Please give me some advice on what I should do.
Thanks.
Thanks!
Yes, this is to be expected. See this:
https://machinelearningmastery.com/faq/single-faq/why-do-i-get-different-results-each-time-i-run-the-code
Thanks. Please suggest me what should I do to get rid of nan loss while training my LSTM model.
Perhaps try relu?
Perhaps try scaling the data before fitting the model?
Hello Jason,
I would like to ask you a question about standardization.you can see the question in the link below, Is it nessary to use the mean and std of training set to scale our Validation/Test set ?
https://stats.stackexchange.com/questions/202287/why-standardization-of-the-testing-set-has-to-be-performed-with-the-mean-and-sd
thank you in advance
Sorry, I don’t have the capacity read/answer the link for you. Perhaps you can summarize your question in a sentence or two?
Hi Jason,
I have seen one student comment stating that we have to do train test split first, then apply normalization / standardization on X_train and X_test. Is that the proper and right way of doing it instead of applying the transformation on whole dataset?
Awaiting your reply. Thanks.
Yes, data preparation coefficients are calculated on train, then the transform is applied to train, test and any other datasets (val, new, etc.)
The reason is to avoid data leakage which results in a biased estimate of model performance.
Is there is a difference between zero-mean normalization and z-score normalization?
Probably not:
https://en.wikipedia.org/wiki/Standard_score
Hello Jason,
thanks for this tutorial. I have a question about scaling approach for a dataset containing nearly 40 features. Some features have range from 0 to 1e+10 and some have range from 0 to 10 or even less, and with different distributions. The Min and Max of each feature is specified directly by researching the problem domain (instead of using the Min and Max of the training data). Therefore this scaler is specifically for this problem.
Question is can I use different scaling function for some features and another for remaining features? Like power transformers for extremely large ranges and MinMaxScaler for the other?
If I only use MinMaxScaler for range (0,1) I think features having large range scales down to very small values close to zero.
I would like to know if there is some efficient way to do scaling so that all the features are scaled appropriately.
Thanks for your time
The MinMaxScaler class allows you to specify the range you prefer, but it applies to all input variables.
If you need something else, you might need to write some custom code.
Hello Jason,
thanks for this tutorial. I get this error when I loaded my data, how can I solve it ?
Error: iterator should return strings, not bytes (did you open the file in text mode?)