Extreme Gradient Boosting (XGBoost) is an open-source library that provides an efficient and effective implementation of the gradient boosting algorithm.
Although other open-source implementations of the approach existed before XGBoost, the release of XGBoost appeared to unleash the power of the technique and made the applied machine learning community take notice of gradient boosting more generally.
Shortly after its development and initial release, XGBoost became the go-to method and often the key component in winning solutions for classification and regression problems in machine learning competitions.
In this tutorial, you will discover how to develop Extreme Gradient Boosting ensembles for classification and regression.
After completing this tutorial, you will know:
- Extreme Gradient Boosting is an efficient open-source implementation of the stochastic gradient boosting ensemble algorithm.
- How to develop XGBoost ensembles for classification and regression with the scikit-learn API.
- How to explore the effect of XGBoost model hyperparameters on model performance.
Kick-start your project with my new book Ensemble Learning Algorithms With Python, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.

Extreme Gradient Boosting (XGBoost) Ensemble in Python
Photo by Andrés Nieto Porras, some rights reserved.
Tutorial Overview
This tutorial is divided into three parts; they are:
- Extreme Gradient Boosting Algorithm
- XGBoost Scikit-Learn API
- XGBoost Ensemble for Classification
- XGBoost Ensemble for Regression
- XGBoost Hyperparameters
- Explore Number of Trees
- Explore Tree Depth
- Explore Learning Rate
- Explore Number of Samples
- Explore Number of Features
Extreme Gradient Boosting Algorithm
Gradient boosting refers to a class of ensemble machine learning algorithms that can be used for classification or regression predictive modeling problems.
Ensembles are constructed from decision tree models. Trees are added one at a time to the ensemble and fit to correct the prediction errors made by prior models. This is a type of ensemble machine learning model referred to as boosting.
Models are fit using any arbitrary differentiable loss function and gradient descent optimization algorithm. This gives the technique its name, “gradient boosting,” as the loss gradient is minimized as the model is fit, much like a neural network.
For more on gradient boosting, see the tutorial:
Extreme Gradient Boosting, or XGBoost for short is an efficient open-source implementation of the gradient boosting algorithm. As such, XGBoost is an algorithm, an open-source project, and a Python library.
It was initially developed by Tianqi Chen and was described by Chen and Carlos Guestrin in their 2016 paper titled “XGBoost: A Scalable Tree Boosting System.”
It is designed to be both computationally efficient (e.g. fast to execute) and highly effective, perhaps more effective than other open-source implementations.
The name xgboost, though, actually refers to the engineering goal to push the limit of computations resources for boosted tree algorithms. Which is the reason why many people use xgboost.
— Tianqi Chen, in answer to the question “What is the difference between the R gbm (gradient boosting machine) and xgboost (extreme gradient boosting)?” on Quora
The two main reasons to use XGBoost are execution speed and model performance.
Generally, XGBoost is fast when compared to other implementations of gradient boosting. Szilard Pafka performed some objective benchmarks comparing the performance of XGBoost to other implementations of gradient boosting and bagged decision trees. He wrote up his results in May 2015 in the blog post titled “Benchmarking Random Forest Implementations.”
His results showed that XGBoost was almost always faster than the other benchmarked implementations from R, Python Spark, and H2O.
From his experiment, he commented:
I also tried xgboost, a popular library for boosting which is capable of building random forests as well. It is fast, memory efficient and of high accuracy
— Benchmarking Random Forest Implementations, Szilard Pafka, 2015.
XGBoost dominates structured or tabular datasets on classification and regression predictive modeling problems. The evidence is that it is the go-to algorithm for competition winners on the Kaggle competitive data science platform.
Among the 29 challenge winning solutions 3 published at Kaggle’s blog during 2015, 17 solutions used XGBoost. […] The success of the system was also witnessed in KDDCup 2015, where XGBoost was used by every winning team in the top-10.
— XGBoost: A Scalable Tree Boosting System, 2016.
Now that we are familiar with what XGBoost is and why it is important, let’s take a closer look at how we can use it in our predictive modeling projects.
Want to Get Started With Ensemble Learning?
Take my free 7-day email crash course now (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
XGBoost Scikit-Learn API
XGBoost can be installed as a standalone library and an XGBoost model can be developed using the scikit-learn API.
The first step is to install the XGBoost library if it is not already installed. This can be achieved using the pip python package manager on most platforms; for example:
|
1 |
sudo pip install xgboost |
You can then confirm that the XGBoost library was installed correctly and can be used by running the following script.
|
1 2 3 |
# check xgboost version import xgboost print(xgboost.__version__) |
Running the script will print your version of the XGBoost library you have installed.
Your version should be the same or higher. If not, you must upgrade your version of the XGBoost library.
|
1 |
1.1.1 |
It is possible that you may have problems with the latest version of the library. It is not your fault.
Sometimes, the most recent version of the library imposes additional requirements or may be less stable.
If you do have errors when trying to run the above script, I recommend downgrading to version 1.0.1 (or lower). This can be achieved by specifying the version to install to the pip command, as follows:
|
1 |
sudo pip install xgboost==1.0.1 |
If you see a warning message, you can safely ignore it for now. For example, below is an example of a warning message that you may see and can ignore:
|
1 |
FutureWarning: pandas.util.testing is deprecated. Use the functions in the public API at pandas.testing instead. |
If you require specific instructions for your development environment, see the tutorial:
The XGBoost library has its own custom API, although we will use the method via the scikit-learn wrapper classes: XGBRegressor and XGBClassifier. This will allow us to use the full suite of tools from the scikit-learn machine learning library to prepare data and evaluate models.
Both models operate the same way and take the same arguments that influence how the decision trees are created and added to the ensemble.
Randomness is used in the construction of the model. This means that each time the algorithm is run on the same data, it will produce a slightly different model.
When using machine learning algorithms that have a stochastic learning algorithm, it is good practice to evaluate them by averaging their performance across multiple runs or repeats of cross-validation. When fitting a final model, it may be desirable to either increase the number of trees until the variance of the model is reduced across repeated evaluations, or to fit multiple final models and average their predictions.
Let’s take a look at how to develop an XGBoost ensemble for both classification and regression.
XGBoost Ensemble for Classification
In this section, we will look at using XGBoost for a classification problem.
First, we can use the make_classification() function to create a synthetic binary classification problem with 1,000 examples and 20 input features.
The complete example is listed below.
|
1 2 3 4 5 6 |
# test classification dataset from sklearn.datasets import make_classification # define dataset X, y = make_classification(n_samples=1000, n_features=20, n_informative=15, n_redundant=5, random_state=7) # summarize the dataset print(X.shape, y.shape) |
Running the example creates the dataset and summarizes the shape of the input and output components.
|
1 |
(1000, 20) (1000,) |
Next, we can evaluate an XGBoost model on this dataset.
We will evaluate the model using repeated stratified k-fold cross-validation, with three repeats and 10 folds. We will report the mean and standard deviation of the accuracy of the model across all repeats and folds.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 |
# evaluate xgboost algorithm for classification from numpy import mean from numpy import std from sklearn.datasets import make_classification from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedStratifiedKFold from xgboost import XGBClassifier # define dataset X, y = make_classification(n_samples=1000, n_features=20, n_informative=15, n_redundant=5, random_state=7) # define the model model = XGBClassifier() # evaluate the model cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1) n_scores = cross_val_score(model, X, y, scoring='accuracy', cv=cv, n_jobs=-1) # report performance print('Accuracy: %.3f (%.3f)' % (mean(n_scores), std(n_scores))) |
Running the example reports the mean and standard deviation accuracy of the model.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see the XGBoost ensemble with default hyperparameters achieves a classification accuracy of about 92.5 percent on this test dataset.
|
1 |
Accuracy: 0.925 (0.028) |
We can also use the XGBoost model as a final model and make predictions for classification.
First, the XGBoost ensemble is fit on all available data, then the predict() function can be called to make predictions on new data. Importantly, this function expects data to always be provided as a NumPy array as a matrix with one row for each input sample.
The example below demonstrates this on our binary classification dataset.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 |
# make predictions using xgboost for classification from numpy import asarray from sklearn.datasets import make_classification from xgboost import XGBClassifier # define dataset X, y = make_classification(n_samples=1000, n_features=20, n_informative=15, n_redundant=5, random_state=7) # define the model model = XGBClassifier() # fit the model on the whole dataset model.fit(X, y) # make a single prediction row = [0.2929949,-4.21223056,-1.288332,-2.17849815,-0.64527665,2.58097719,0.28422388,-7.1827928,-1.91211104,2.73729512,0.81395695,3.96973717,-2.66939799,3.34692332,4.19791821,0.99990998,-0.30201875,-4.43170633,-2.82646737,0.44916808] row = asarray([row]) yhat = model.predict(row) print('Predicted Class: %d' % yhat[0]) |
Running the example fits the XGBoost ensemble model on the entire dataset and is then used to make a prediction on a new row of data, as we might when using the model in an application.
|
1 |
Predicted Class: 1 |
Now that we are familiar with using XGBoost for classification, let’s look at the API for regression.
XGBoost Ensemble for Regression
In this section, we will look at using XGBoost for a regression problem.
First, we can use the make_regression() function to create a synthetic regression problem with 1,000 examples and 20 input features.
The complete example is listed below.
|
1 2 3 4 5 6 |
# test regression dataset from sklearn.datasets import make_regression # define dataset X, y = make_regression(n_samples=1000, n_features=20, n_informative=15, noise=0.1, random_state=7) # summarize the dataset print(X.shape, y.shape) |
Running the example creates the dataset and summarizes the shape of the input and output components.
|
1 |
(1000, 20) (1000,) |
Next, we can evaluate an XGBoost algorithm on this dataset.
As we did with the last section, we will evaluate the model using repeated k-fold cross-validation, with three repeats and 10 folds. We will report the mean absolute error (MAE) of the model across all repeats and folds. The scikit-learn library makes the MAE negative so that it is maximized instead of minimized. This means that larger negative MAE are better and a perfect model has a MAE of 0.
The complete example is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 |
# evaluate xgboost ensemble for regression from numpy import mean from numpy import std from sklearn.datasets import make_regression from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedKFold from xgboost import XGBRegressor # define dataset X, y = make_regression(n_samples=1000, n_features=20, n_informative=15, noise=0.1, random_state=7) # define the model model = XGBRegressor() # evaluate the model cv = RepeatedKFold(n_splits=10, n_repeats=3, random_state=1) n_scores = cross_val_score(model, X, y, scoring='neg_mean_absolute_error', cv=cv, n_jobs=-1, error_score='raise') # report performance print('MAE: %.3f (%.3f)' % (mean(n_scores), std(n_scores))) |
Running the example reports the mean and standard deviation accuracy of the model.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see the XGBoost ensemble with default hyperparameters achieves a MAE of about 76.
|
1 |
MAE: -76.447 (3.859) |
We can also use the XGBoost model as a final model and make predictions for regression.
First, the XGBoost ensemble is fit on all available data, then the predict() function can be called to make predictions on new data. As with classification, the single row of data must be represented as a two-dimensional matrix in NumPy array format.
The example below demonstrates this on our regression dataset.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 |
# gradient xgboost for making predictions for regression from numpy import asarray from sklearn.datasets import make_regression from xgboost import XGBRegressor # define dataset X, y = make_regression(n_samples=1000, n_features=20, n_informative=15, noise=0.1, random_state=7) # define the model model = XGBRegressor() # fit the model on the whole dataset model.fit(X, y) # make a single prediction row = [0.20543991,-0.97049844,-0.81403429,-0.23842689,-0.60704084,-0.48541492,0.53113006,2.01834338,-0.90745243,-1.85859731,-1.02334791,-0.6877744,0.60984819,-0.70630121,-1.29161497,1.32385441,1.42150747,1.26567231,2.56569098,-0.11154792] row = asarray([row]) yhat = model.predict(row) print('Prediction: %d' % yhat[0]) |
Running the example fits the XGBoost ensemble model on the entire dataset and is then used to make a prediction on a new row of data, as we might when using the model in an application.
|
1 |
Prediction: 50 |
Now that we are familiar with using the XGBoost Scikit-Learn API to evaluate and use XGBoost ensembles, let’s look at configuring the model.
XGBoost Hyperparameters
In this section, we will take a closer look at some of the hyperparameters you should consider tuning for the Gradient Boosting ensemble and their effect on model performance.
Explore Number of Trees
An important hyperparameter for the XGBoost ensemble algorithm is the number of decision trees used in the ensemble.
Recall that decision trees are added to the model sequentially in an effort to correct and improve upon the predictions made by prior trees. As such, more trees is often better.
The number of trees can be set via the “n_estimators” argument and defaults to 100.
The example below explores the effect of the number of trees with values between 10 to 5,000.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 |
# explore xgboost number of trees effect on performance from numpy import mean from numpy import std from sklearn.datasets import make_classification from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedStratifiedKFold from xgboost import XGBClassifier from matplotlib import pyplot # get the dataset def get_dataset(): X, y = make_classification(n_samples=1000, n_features=20, n_informative=15, n_redundant=5, random_state=7) return X, y # get a list of models to evaluate def get_models(): models = dict() trees = [10, 50, 100, 500, 1000, 5000] for n in trees: models[str(n)] = XGBClassifier(n_estimators=n) return models # evaluate a give model using cross-validation def evaluate_model(model): cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1) scores = cross_val_score(model, X, y, scoring='accuracy', cv=cv, n_jobs=-1) return scores # define dataset X, y = get_dataset() # get the models to evaluate models = get_models() # evaluate the models and store results results, names = list(), list() for name, model in models.items(): scores = evaluate_model(model) results.append(scores) names.append(name) print('>%s %.3f (%.3f)' % (name, mean(scores), std(scores))) # plot model performance for comparison pyplot.boxplot(results, labels=names, showmeans=True) pyplot.show() |
Running the example first reports the mean accuracy for each configured number of decision trees.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see that that performance improves on this dataset until about 500 trees, after which performance appears to level off or decrease.
|
1 2 3 4 5 6 |
>10 0.885 (0.029) >50 0.915 (0.029) >100 0.925 (0.028) >500 0.927 (0.028) >1000 0.926 (0.028) >5000 0.925 (0.027) |
A box and whisker plot is created for the distribution of accuracy scores for each configured number of trees.
We can see the general trend of increasing model performance and ensemble size.
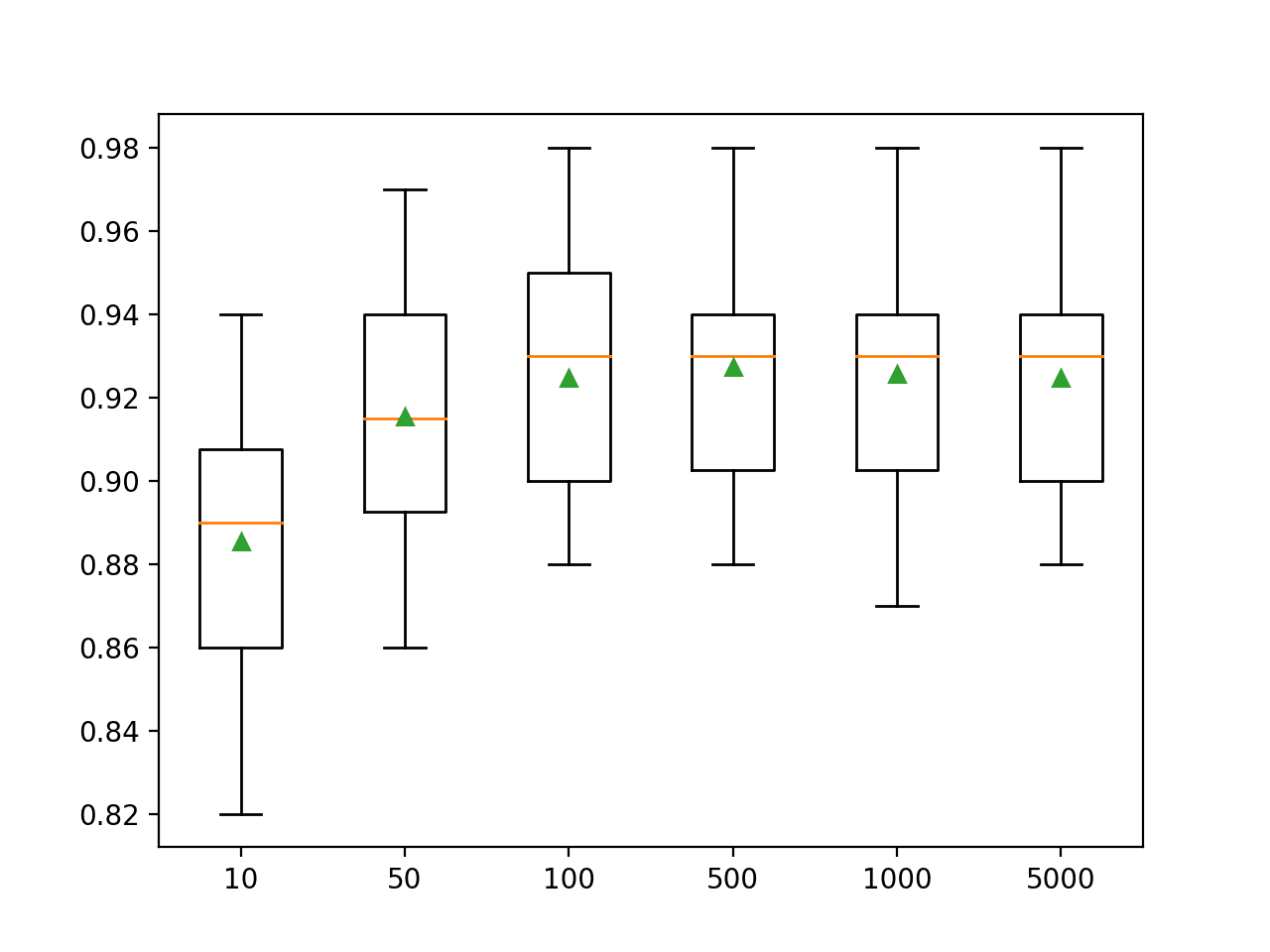
Box Plots of XGBoost Ensemble Size vs. Classification Accuracy
Explore Tree Depth
Varying the depth of each tree added to the ensemble is another important hyperparameter for gradient boosting.
The tree depth controls how specialized each tree is to the training dataset: how general or overfit it might be. Trees are preferred that are not too shallow and general (like AdaBoost) and not too deep and specialized (like bootstrap aggregation).
Gradient boosting generally performs well with trees that have a modest depth, finding a balance between skill and generality.
Tree depth is controlled via the “max_depth” argument and defaults to 6.
The example below explores tree depths between 1 and 10 and the effect on model performance.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 |
# explore xgboost tree depth effect on performance from numpy import mean from numpy import std from sklearn.datasets import make_classification from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedStratifiedKFold from xgboost import XGBClassifier from matplotlib import pyplot # get the dataset def get_dataset(): X, y = make_classification(n_samples=1000, n_features=20, n_informative=15, n_redundant=5, random_state=7) return X, y # get a list of models to evaluate def get_models(): models = dict() for i in range(1,11): models[str(i)] = XGBClassifier(max_depth=i) return models # evaluate a give model using cross-validation def evaluate_model(model): cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1) scores = cross_val_score(model, X, y, scoring='accuracy', cv=cv, n_jobs=-1) return scores # define dataset X, y = get_dataset() # get the models to evaluate models = get_models() # evaluate the models and store results results, names = list(), list() for name, model in models.items(): scores = evaluate_model(model) results.append(scores) names.append(name) print('>%s %.3f (%.3f)' % (name, mean(scores), std(scores))) # plot model performance for comparison pyplot.boxplot(results, labels=names, showmeans=True) pyplot.show() |
Running the example first reports the mean accuracy for each configured tree depth.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see that performance improves with tree depth, perhaps peeking around a depth of 3 to 8, after which the deeper, more specialized trees result in worse performance.
|
1 2 3 4 5 6 7 8 9 10 |
>1 0.849 (0.028) >2 0.906 (0.032) >3 0.926 (0.027) >4 0.930 (0.027) >5 0.924 (0.031) >6 0.925 (0.028) >7 0.926 (0.030) >8 0.926 (0.029) >9 0.921 (0.032) >10 0.923 (0.035) |
A box and whisker plot is created for the distribution of accuracy scores for each configured tree depth.
We can see the general trend of increasing model performance with the tree depth to a point, after which performance begins to sit flat or degrade with the over-specialized trees.
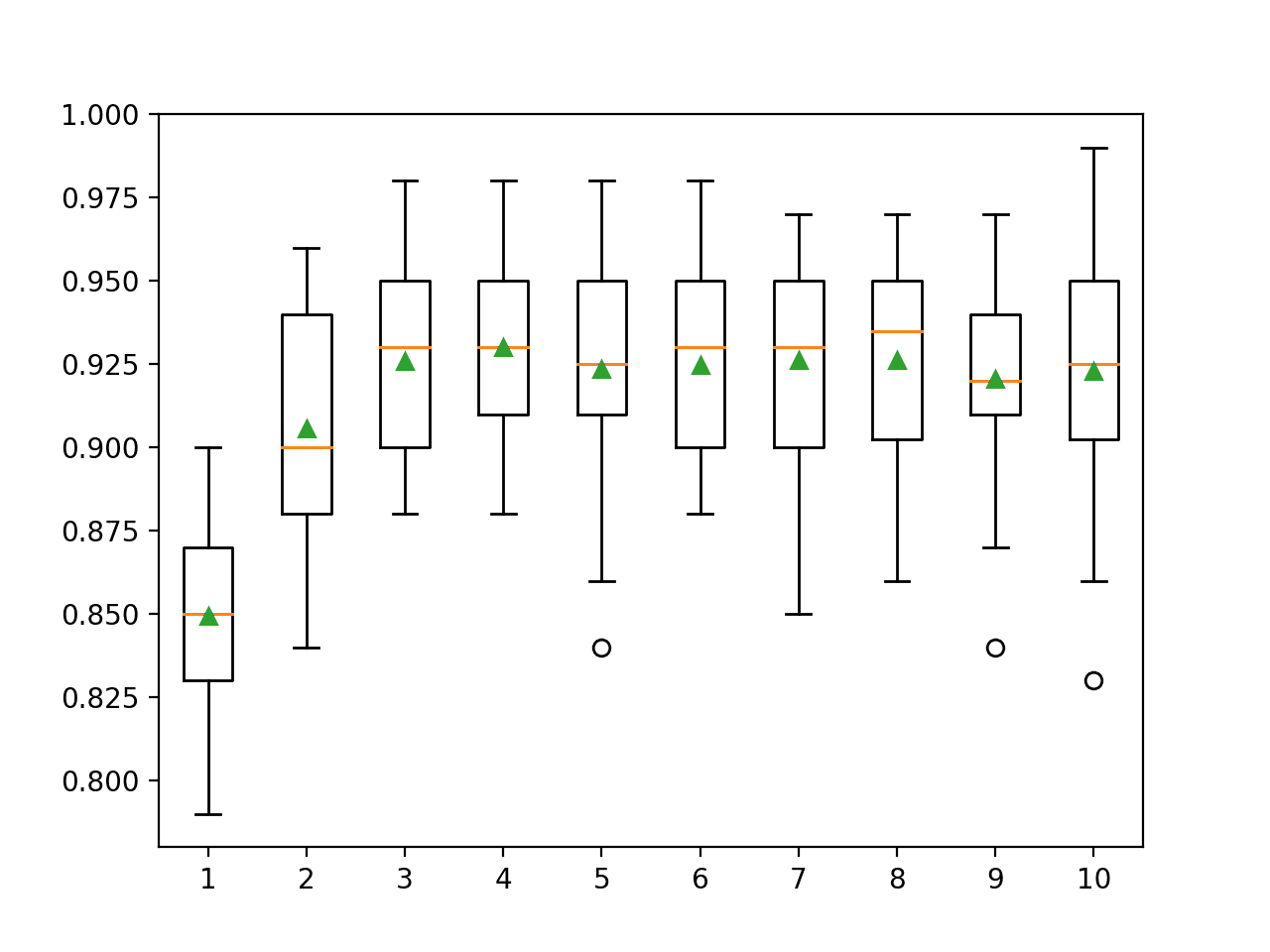
Box Plots of XGBoost Ensemble Tree Depth vs. Classification Accuracy
Explore Learning Rate
Learning rate controls the amount of contribution that each model has on the ensemble prediction.
Smaller rates may require more decision trees in the ensemble.
The learning rate can be controlled via the “eta” argument and defaults to 0.3.
The example below explores the learning rate and compares the effect of values between 0.0001 and 1.0.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 |
# explore xgboost learning rate effect on performance from numpy import mean from numpy import std from sklearn.datasets import make_classification from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedStratifiedKFold from xgboost import XGBClassifier from matplotlib import pyplot # get the dataset def get_dataset(): X, y = make_classification(n_samples=1000, n_features=20, n_informative=15, n_redundant=5, random_state=7) return X, y # get a list of models to evaluate def get_models(): models = dict() rates = [0.0001, 0.001, 0.01, 0.1, 1.0] for r in rates: key = '%.4f' % r models[key] = XGBClassifier(eta=r) return models # evaluate a give model using cross-validation def evaluate_model(model): cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1) scores = cross_val_score(model, X, y, scoring='accuracy', cv=cv, n_jobs=-1) return scores # define dataset X, y = get_dataset() # get the models to evaluate models = get_models() # evaluate the models and store results results, names = list(), list() for name, model in models.items(): scores = evaluate_model(model) results.append(scores) names.append(name) print('>%s %.3f (%.3f)' % (name, mean(scores), std(scores))) # plot model performance for comparison pyplot.boxplot(results, labels=names, showmeans=True) pyplot.show() |
Running the example first reports the mean accuracy for each configured learning rate.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see that a larger learning rate results in better performance on this dataset. We would expect that adding more trees to the ensemble for the smaller learning rates would further lift performance.
This highlights the trade-off between the number of trees (speed of training) and learning rate, e.g. we can fit a model faster by using fewer trees and a larger learning rate.
|
1 2 3 4 5 |
>0.0001 0.804 (0.039) >0.0010 0.814 (0.037) >0.0100 0.867 (0.027) >0.1000 0.923 (0.030) >1.0000 0.913 (0.030) |
A box and whisker plot is created for the distribution of accuracy scores for each configured learning rate.
We can see the general trend of increasing model performance with the increase in learning rate of 0.1, after which performance degrades.
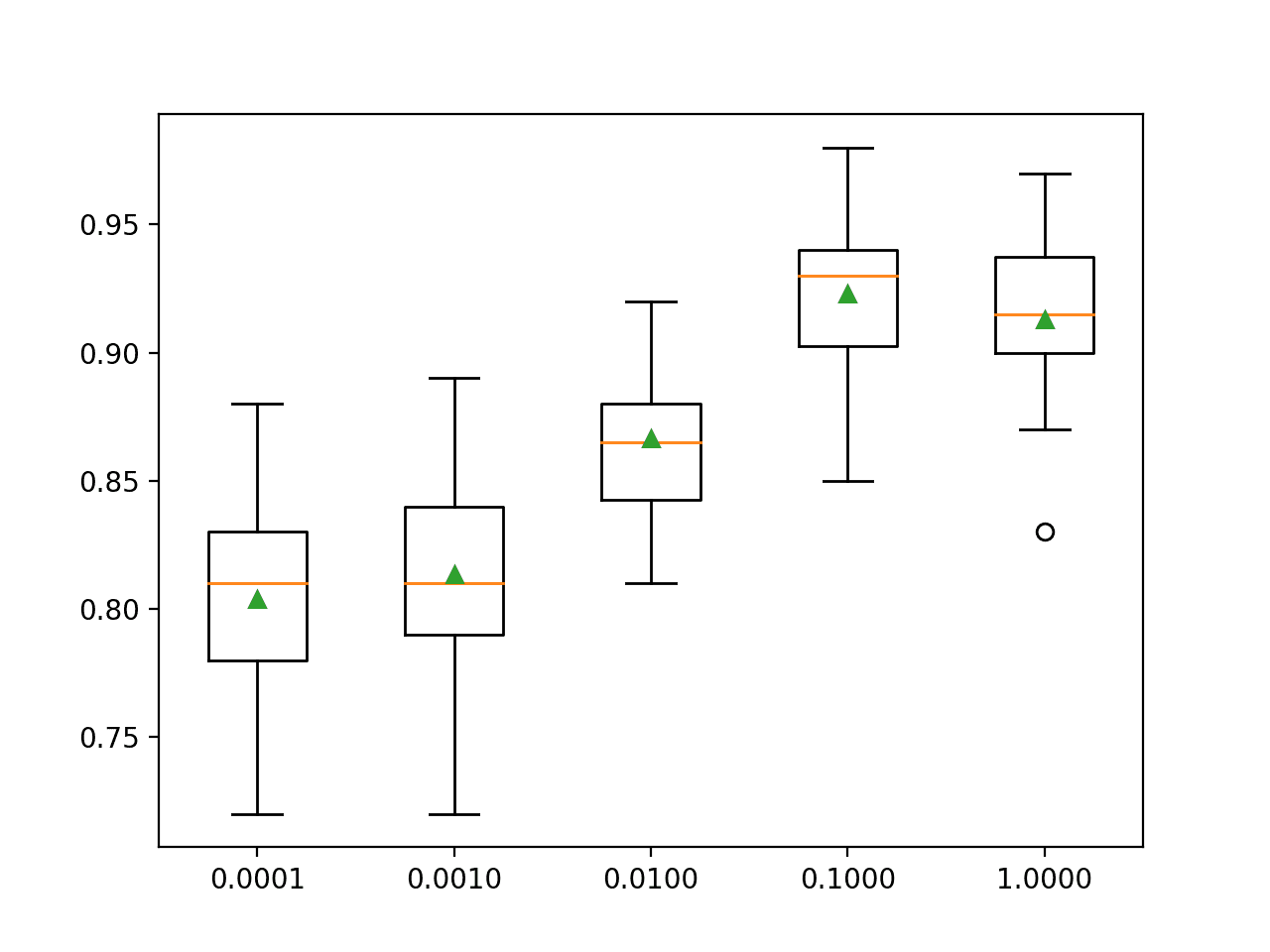
Box Plot of XGBoost Learning Rate vs. Classification Accuracy
Explore Number of Samples
The number of samples used to fit each tree can be varied. This means that each tree is fit on a randomly selected subset of the training dataset.
Using fewer samples introduces more variance for each tree, although it can improve the overall performance of the model.
The number of samples used to fit each tree is specified by the “subsample” argument and can be set to a fraction of the training dataset size. By default, it is set to 1.0 to use the entire training dataset.
The example below demonstrates the effect of the sample size on model performance with ratios varying from 10 percent to 100 percent in 10 percent increments.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 |
# explore xgboost subsample ratio effect on performance from numpy import arange from numpy import mean from numpy import std from sklearn.datasets import make_classification from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedStratifiedKFold from xgboost import XGBClassifier from matplotlib import pyplot # get the dataset def get_dataset(): X, y = make_classification(n_samples=1000, n_features=20, n_informative=15, n_redundant=5, random_state=7) return X, y # get a list of models to evaluate def get_models(): models = dict() for i in arange(0.1, 1.1, 0.1): key = '%.1f' % i models[key] = XGBClassifier(subsample=i) return models # evaluate a give model using cross-validation def evaluate_model(model): cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1) scores = cross_val_score(model, X, y, scoring='accuracy', cv=cv, n_jobs=-1) return scores # define dataset X, y = get_dataset() # get the models to evaluate models = get_models() # evaluate the models and store results results, names = list(), list() for name, model in models.items(): scores = evaluate_model(model) results.append(scores) names.append(name) print('>%s %.3f (%.3f)' % (name, mean(scores), std(scores))) # plot model performance for comparison pyplot.boxplot(results, labels=names, showmeans=True) pyplot.show() |
Running the example first reports the mean accuracy for each configured sample size.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see that mean performance is probably best for a sample size that covers most of the dataset, such as 80 percent or higher.
|
1 2 3 4 5 6 7 8 9 10 |
>0.1 0.876 (0.027) >0.2 0.912 (0.033) >0.3 0.917 (0.032) >0.4 0.925 (0.026) >0.5 0.928 (0.027) >0.6 0.926 (0.024) >0.7 0.925 (0.031) >0.8 0.928 (0.028) >0.9 0.928 (0.025) >1.0 0.925 (0.028) |
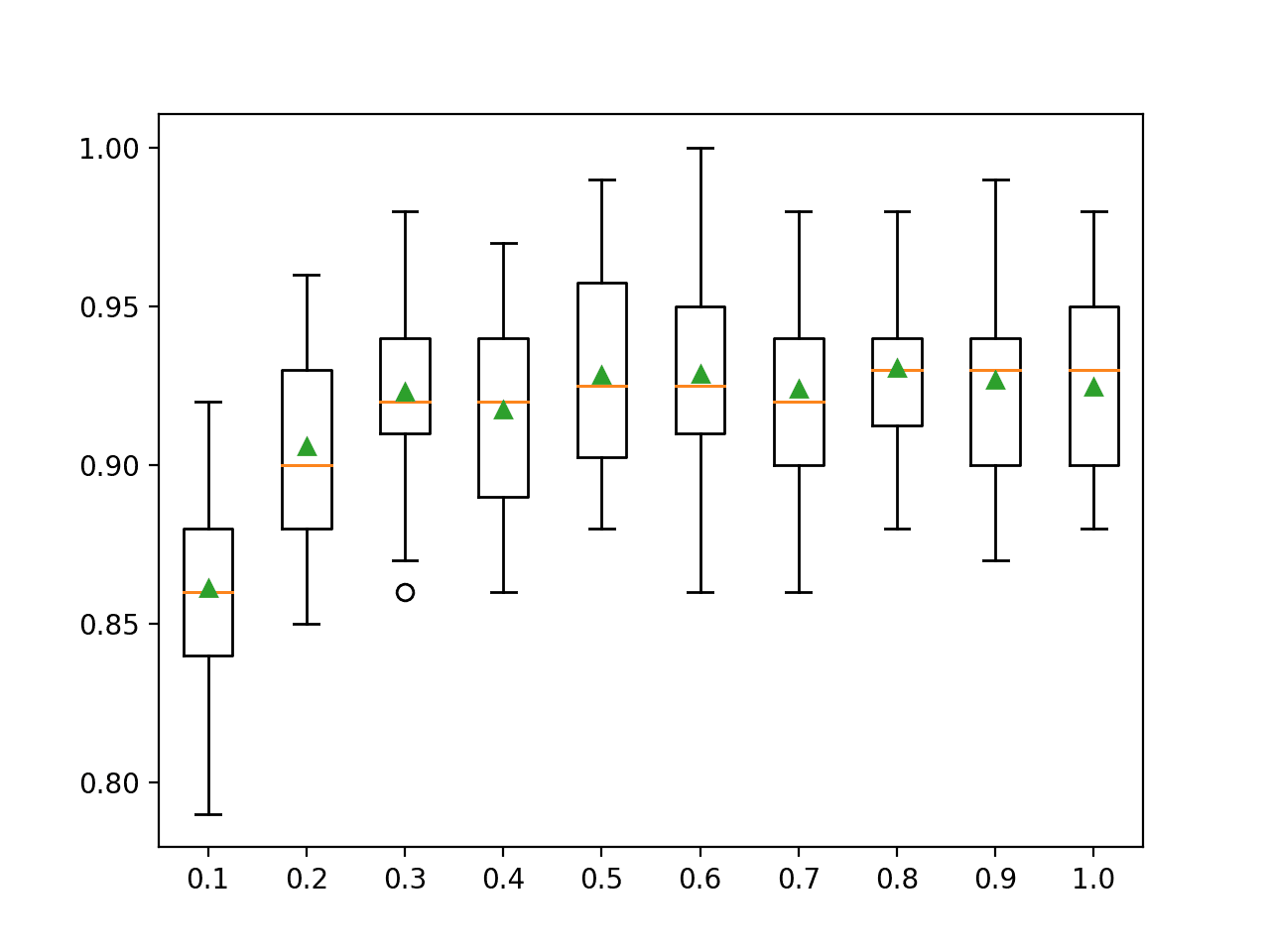
A box and whisker plot is created for the distribution of accuracy scores for each configured sampling ratio.
We can see the general trend of increasing model performance, perhaps peaking around 80 percent and staying somewhat level.
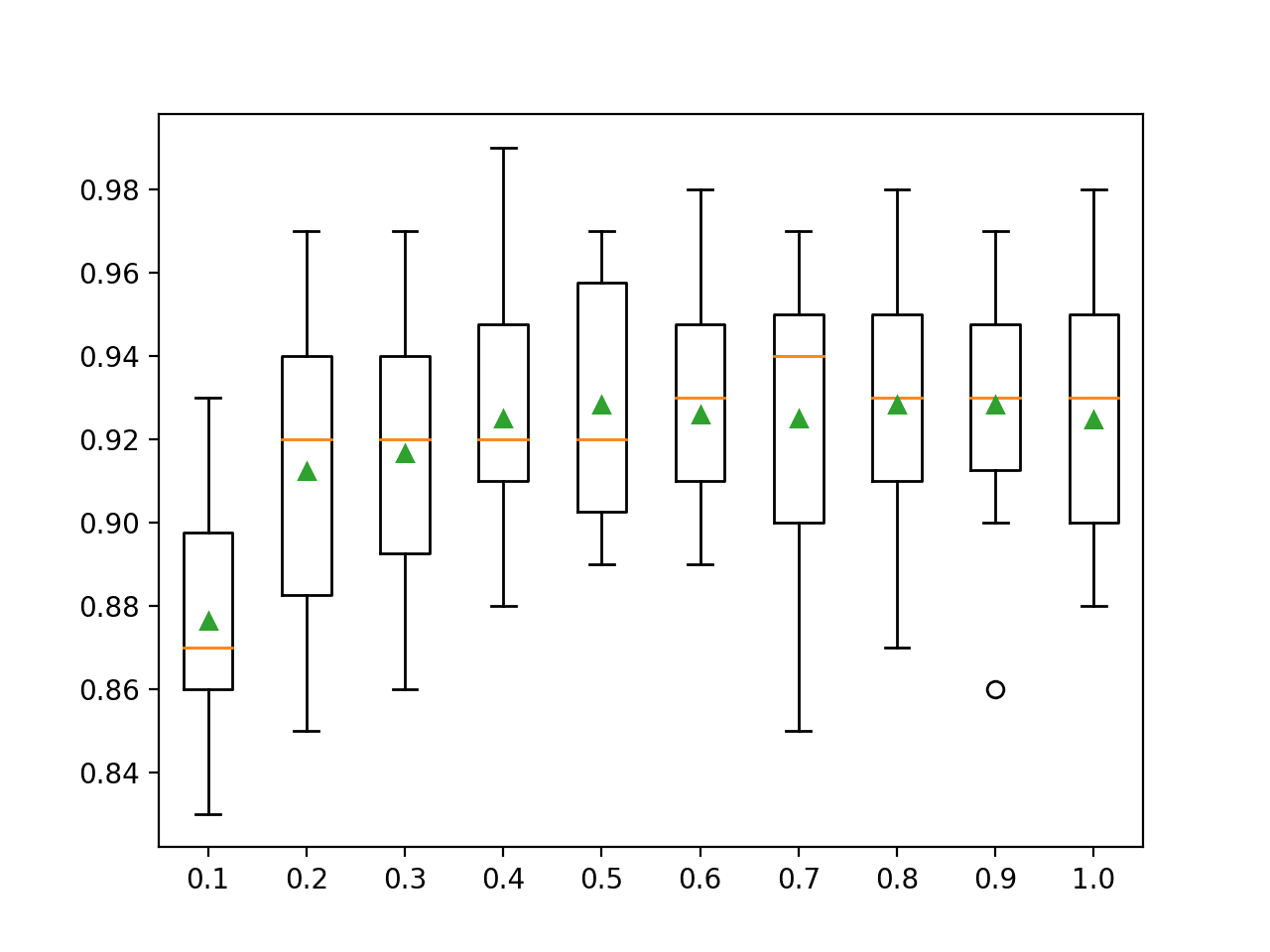
Box Plots of XGBoost Ensemble Sample Ratio vs. Classification Accuracy
Explore Number of Features
The number of features used to fit each decision tree can be varied.
Like changing the number of samples, changing the number of features introduces additional variance into the model, which may improve performance, although it might require an increase in the number of trees.
The number of features used by each tree is taken as a random sample and is specified by the “colsample_bytree” argument and defaults to all features in the training dataset, e.g. 100 percent or a value of 1.0. You can also sample columns for each split, and this is controlled by the “colsample_bylevel” argument, but we will not look at this hyperparameter here.
The example below explores the effect of the number of features on model performance with ratios varying from 10 percent to 100 percent in 10 percent increments.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 |
# explore xgboost column ratio per tree effect on performance from numpy import arange from numpy import mean from numpy import std from sklearn.datasets import make_classification from sklearn.model_selection import cross_val_score from sklearn.model_selection import RepeatedStratifiedKFold from xgboost import XGBClassifier from matplotlib import pyplot # get the dataset def get_dataset(): X, y = make_classification(n_samples=1000, n_features=20, n_informative=15, n_redundant=5, random_state=7) return X, y # get a list of models to evaluate def get_models(): models = dict() for i in arange(0.1, 1.1, 0.1): key = '%.1f' % i models[key] = XGBClassifier(colsample_bytree=i) return models # evaluate a give model using cross-validation def evaluate_model(model): cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1) scores = cross_val_score(model, X, y, scoring='accuracy', cv=cv, n_jobs=-1) return scores # define dataset X, y = get_dataset() # get the models to evaluate models = get_models() # evaluate the models and store results results, names = list(), list() for name, model in models.items(): scores = evaluate_model(model) results.append(scores) names.append(name) print('>%s %.3f (%.3f)' % (name, mean(scores), std(scores))) # plot model performance for comparison pyplot.boxplot(results, labels=names, showmeans=True) pyplot.show() |
Running the example first reports the mean accuracy for each configured ratio of columns.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see that mean performance increases to about half the number of features (50 percent) and stays somewhat level after that. It’s surprising that removing half of the input variables per tree has so little effect.
|
1 2 3 4 5 6 7 8 9 10 |
>0.1 0.861 (0.033) >0.2 0.906 (0.027) >0.3 0.923 (0.029) >0.4 0.917 (0.029) >0.5 0.928 (0.030) >0.6 0.929 (0.031) >0.7 0.924 (0.027) >0.8 0.931 (0.025) >0.9 0.927 (0.033) >1.0 0.925 (0.028) |
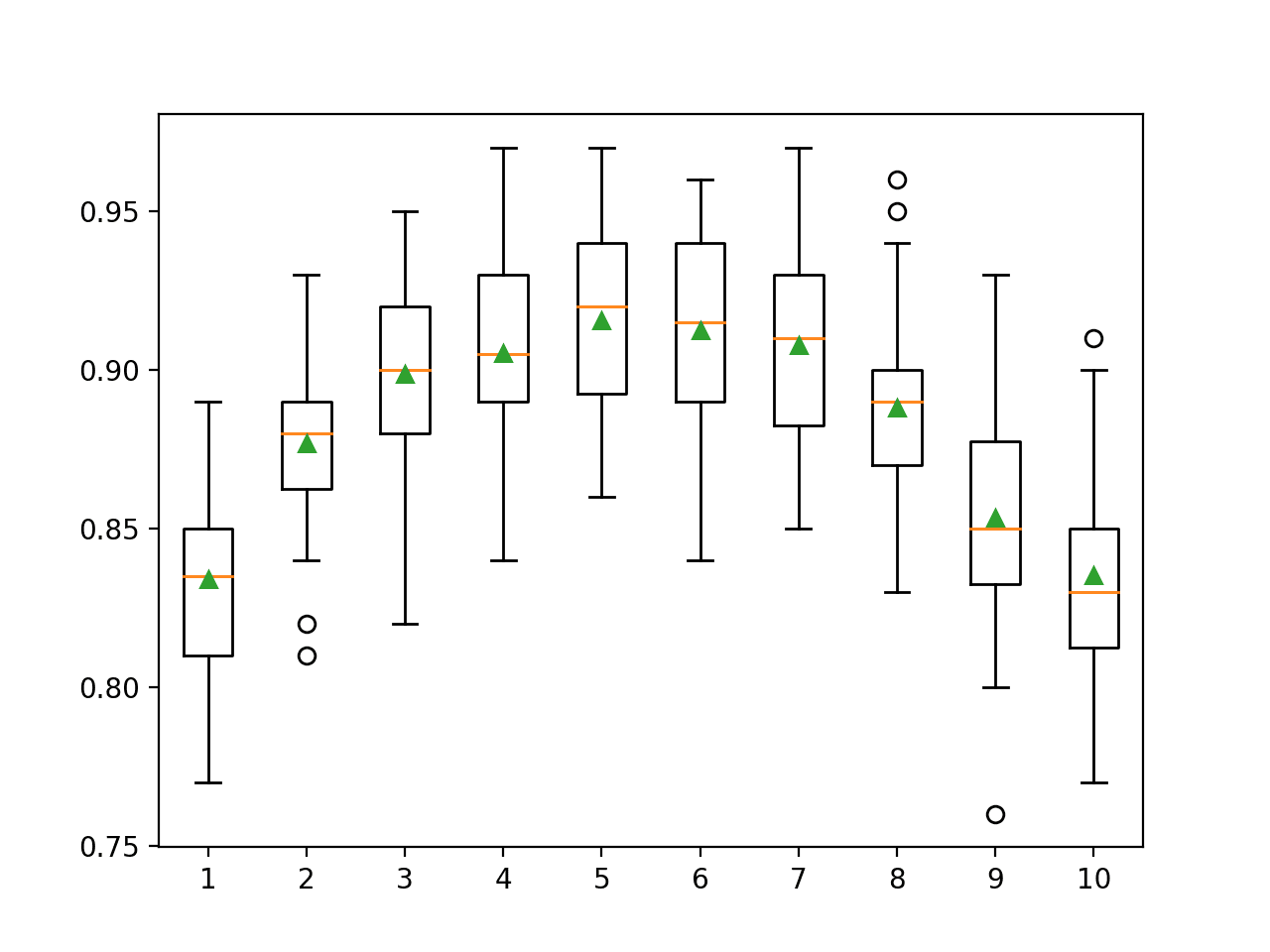
A box and whisker plot is created for the distribution of accuracy scores for each configured column ratio.
We can see the general trend of increasing model performance perhaps peaking with a ratio of 60 percent and staying somewhat level.
Box Plots of XGBoost Ensemble Column Ratio vs. Classification Accuracy
Further Reading
This section provides more resources on the topic if you are looking to go deeper.
Tutorials
- A Gentle Introduction to the Gradient Boosting Algorithm for Machine Learning
- Gradient Boosting with Scikit-Learn, XGBoost, LightGBM, and CatBoost
- A Gentle Introduction to XGBoost for Applied Machine Learning
Papers
Project
APIs
Articles
Summary
In this tutorial, you discovered how to develop Extreme Gradient Boosting ensembles for classification and regression.
Specifically, you learned:
- Extreme Gradient Boosting is an efficient open-source implementation of the stochastic gradient boosting ensemble algorithm.
- How to develop XGBoost ensembles for classification and regression with the scikit-learn API.
- How to explore the effect of XGBoost model hyperparameters on model performance.
Do you have any questions?
Ask your questions in the comments below and I will do my best to answer.





Jason, I’m wondering if my results might vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision ? Sorry, just teasin. I notice you’ve used that phrase here and in other artciles. Top notich material in any case and thanks for putting together these artciles which always pack a lot of information inside a little space.
Thanks.
Yes, I use a wordpress shortcode so the same text disclaimer follows any results in all recent tutorials. It stops the tens of daily emails asking “why are my results slightly different to your results?”
Can I use Gradient Boosting (XGBoost) for multi output regression
I believe so, perhaps try it and see.
Hi Jason,
Thanks for the wonderful tutorial, it was very helpful to me.
I have a question. I want to implement a structure consist of CNN, deep auto encoder and XGB. I have defined a functional Model that consist of CNN and deep auto encoder. In the second layer, the latent vector of deep auto encoder should fed into an extreme gradient boost based classifier.
I used StackingClassifier, but I get the error: The estimator Functional should be classifier
Could you please help me that how I can do this?
Perhaps you can use the neural net as a pre-processing step, save the data, then use it to train another model like XGB.
Hey Jason,
Do you think you can explain what you mean by using Neural Net as a preprocessing step then using the data to train the XGB. By preprocessing what do you mean. I am working in extreme XGB models don came across your code, pretty interesting. great work by the way . You make our lives easier . Thank you
If you can give me clarity or direct me to where I can see an example of what you mean by preprocessing, I will
Be most happy
Hi Amobi…The following resource may be of interest to you:
https://machinelearningmastery.com/data-preparation-gradient-boosting-xgboost-python/
Thank you so much. I have gone through it. Now makes sense to me.
I have a quick question. While studying your post on ensemble Neural Net (sticking Ensemble for Deep learning Neural Network) , I wondered if I can use XGBoost to as a meta learner while solving Neural network regression problem