XGBoost is a popular implementation of Gradient Boosting because of its speed and performance.
Internally, XGBoost models represent all problems as a regression predictive modeling problem that only takes numerical values as input. If your data is in a different form, it must be prepared into the expected format.
In this post, you will discover how to prepare your data for using with gradient boosting with the XGBoost library in Python.
After reading this post you will know:
- How to encode string output variables for classification.
- How to prepare categorical input variables using one hot encoding.
- How to automatically handle missing data with XGBoost.
Kick-start your project with my new book XGBoost With Python, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.
- Update Sept/2016: I updated a few small typos in the impute example.
- Update Jan/2017: Updated to reflect changes in scikit-learn API version 0.18.1.
- Update Jan/2017: Updated breast cancer example to converted input data to strings.
- Update Oct/2019: Updated usage of OneHotEncoder to suppress warnings.
- Update Dec/2019: Updated example to fix bug in API usage in the multi-class example.
- Update May/2020: Updated to use SimpleImputer in new v0.22 API.

Data Preparation for Gradient Boosting with XGBoost in Python
Photo by Ed Dunens, some rights reserved.
Need help with XGBoost in Python?
Take my free 7-day email course and discover xgboost (with sample code).
Click to sign-up now and also get a free PDF Ebook version of the course.
Label Encode String Class Values
The iris flowers classification problem is an example of a problem that has a string class value.
This is a prediction problem where given measurements of iris flowers in centimeters, the task is to predict to which species a given flower belongs.
Download the dataset and place it in your current working directory with the filename “iris.csv“.
Below is a sample of the raw dataset.
|
1 2 3 4 5 6 |
5.1,3.5,1.4,0.2,Iris-setosa 4.9,3.0,1.4,0.2,Iris-setosa 4.7,3.2,1.3,0.2,Iris-setosa 4.6,3.1,1.5,0.2,Iris-setosa 5.0,3.6,1.4,0.2,Iris-setosa ... |
XGBoost cannot model this problem as-is as it requires that the output variables be numeric.
We can easily convert the string values to integer values using the LabelEncoder. The three class values (Iris-setosa, Iris-versicolor, Iris-virginica) are mapped to the integer values (0, 1, 2).
|
1 2 3 4 |
# encode string class values as integers label_encoder = LabelEncoder() label_encoder = label_encoder.fit(Y) label_encoded_y = label_encoder.transform(Y) |
We save the label encoder as a separate object so that we can transform both the training and later the test and validation datasets using the same encoding scheme.
Below is a complete example demonstrating how to load the iris dataset. Notice that Pandas is used to load the data in order to handle the string class values.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 |
# multiclass classification import pandas import xgboost from sklearn import model_selection from sklearn.metrics import accuracy_score from sklearn.preprocessing import LabelEncoder # load data data = pandas.read_csv('iris.csv', header=None) dataset = data.values # split data into X and y X = dataset[:,0:4] Y = dataset[:,4] # encode string class values as integers label_encoder = LabelEncoder() label_encoder = label_encoder.fit(Y) label_encoded_y = label_encoder.transform(Y) seed = 7 test_size = 0.33 X_train, X_test, y_train, y_test = model_selection.train_test_split(X, label_encoded_y, test_size=test_size, random_state=seed) # fit model no training data model = xgboost.XGBClassifier() model.fit(X_train, y_train) print(model) # make predictions for test data y_pred = model.predict(X_test) predictions = [round(value) for value in y_pred] # evaluate predictions accuracy = accuracy_score(y_test, predictions) print("Accuracy: %.2f%%" % (accuracy * 100.0)) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running the example produces the following output.
|
1 2 3 4 5 6 |
XGBClassifier(base_score=0.5, colsample_bylevel=1, colsample_bytree=1, gamma=0, learning_rate=0.1, max_delta_step=0, max_depth=3, min_child_weight=1, missing=None, n_estimators=100, nthread=-1, objective='multi:softprob', reg_alpha=0, reg_lambda=1, scale_pos_weight=1, seed=0, silent=True, subsample=1) Accuracy: 92.00% |
Notice how the XGBoost model is configured to automatically model the multiclass classification problem using the multi:softprob objective, a variation on the softmax loss function to model class probabilities. This suggests that internally, that the output class is converted into a one hot type encoding automatically.
One Hot Encode Categorical Data
Some datasets only contain categorical data, for example the breast cancer dataset.
This dataset describes the technical details of breast cancer biopsies and the prediction task is to predict whether or not the patient has a recurrence of cancer, or not.
Download the dataset and place it in your current working directory with the filename “breast-cancer.csv“.
Below is a sample of the raw dataset.
|
1 2 3 4 5 6 |
'40-49','premeno','15-19','0-2','yes','3','right','left_up','no','recurrence-events' '50-59','ge40','15-19','0-2','no','1','right','central','no','no-recurrence-events' '50-59','ge40','35-39','0-2','no','2','left','left_low','no','recurrence-events' '40-49','premeno','35-39','0-2','yes','3','right','left_low','yes','no-recurrence-events' '40-49','premeno','30-34','3-5','yes','2','left','right_up','no','recurrence-events' ... |
We can see that all 9 input variables are categorical and described in string format. The problem is a binary classification prediction problem and the output class values are also described in string format.
We can reuse the same approach from the previous section and convert the string class values to integer values to model the prediction using the LabelEncoder. For example:
|
1 2 3 4 |
# encode string class values as integers label_encoder = LabelEncoder() label_encoder = label_encoder.fit(Y) label_encoded_y = label_encoder.transform(Y) |
We can use this same approach on each input feature in X, but this is only a starting point.
|
1 2 3 4 5 6 7 8 |
# encode string input values as integers features = [] for i in range(0, X.shape[1]): label_encoder = LabelEncoder() feature = label_encoder.fit_transform(X[:,i]) features.append(feature) encoded_x = numpy.array(features) encoded_x = encoded_x.reshape(X.shape[0], X.shape[1]) |
XGBoost may assume that encoded integer values for each input variable have an ordinal relationship. For example that ‘left-up’ encoded as 0 and ‘left-low’ encoded as 1 for the breast-quad variable have a meaningful relationship as integers. In this case, this assumption is untrue.
Instead, we must map these integer values onto new binary variables, one new variable for each categorical value.
For example, the breast-quad variable has the values:
|
1 2 3 4 5 |
left-up left-low right-up right-low central |
We can model this as 5 binary variables as follows:
|
1 2 3 4 5 6 |
left-up, left-low, right-up, right-low, central 1,0,0,0,0 0,1,0,0,0 0,0,1,0,0 0,0,0,1,0 0,0,0,0,1 |
This is called one hot encoding. We can one hot encode all of the categorical input variables using the OneHotEncoder class in scikit-learn.
We can one hot encode each feature after we have label encoded it. First we must transform the feature array into a 2-dimensional NumPy array where each integer value is a feature vector with a length 1.
|
1 |
feature = feature.reshape(X.shape[0], 1) |
We can then create the OneHotEncoder and encode the feature array.
|
1 2 |
onehot_encoder = OneHotEncoder(sparse=False, categories='auto') feature = onehot_encoder.fit_transform(feature) |
Finally, we can build up the input dataset by concatenating the one hot encoded features, one by one, adding them on as new columns (axis=2). We end up with an input vector comprised of 43 binary input variables.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 |
# encode string input values as integers encoded_x = None for i in range(0, X.shape[1]): label_encoder = LabelEncoder() feature = label_encoder.fit_transform(X[:,i]) feature = feature.reshape(X.shape[0], 1) onehot_encoder = OneHotEncoder(sparse=False, categories='auto') feature = onehot_encoder.fit_transform(feature) if encoded_x is None: encoded_x = feature else: encoded_x = numpy.concatenate((encoded_x, feature), axis=1) print("X shape: : ", encoded_x.shape) |
Ideally, we may experiment with not one hot encode some of input attributes as we could encode them with an explicit ordinal relationship, for example the first column age with values like ’40-49′ and ’50-59′. This is left as an exercise, if you are interested in extending this example.
Below is the complete example with label and one hot encoded input variables and label encoded output variable.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 |
# binary classification, breast cancer dataset, label and one hot encoded import numpy from pandas import read_csv from xgboost import XGBClassifier from sklearn.model_selection import train_test_split from sklearn.metrics import accuracy_score from sklearn.preprocessing import LabelEncoder from sklearn.preprocessing import OneHotEncoder # load data data = read_csv('breast-cancer.csv', header=None) dataset = data.values # split data into X and y X = dataset[:,0:9] X = X.astype(str) Y = dataset[:,9] # encode string input values as integers encoded_x = None for i in range(0, X.shape[1]): label_encoder = LabelEncoder() feature = label_encoder.fit_transform(X[:,i]) feature = feature.reshape(X.shape[0], 1) onehot_encoder = OneHotEncoder(sparse=False, categories='auto') feature = onehot_encoder.fit_transform(feature) if encoded_x is None: encoded_x = feature else: encoded_x = numpy.concatenate((encoded_x, feature), axis=1) print("X shape: : ", encoded_x.shape) # encode string class values as integers label_encoder = LabelEncoder() label_encoder = label_encoder.fit(Y) label_encoded_y = label_encoder.transform(Y) # split data into train and test sets seed = 7 test_size = 0.33 X_train, X_test, y_train, y_test = train_test_split(encoded_x, label_encoded_y, test_size=test_size, random_state=seed) # fit model no training data model = XGBClassifier() model.fit(X_train, y_train) print(model) # make predictions for test data y_pred = model.predict(X_test) predictions = [round(value) for value in y_pred] # evaluate predictions accuracy = accuracy_score(y_test, predictions) print("Accuracy: %.2f%%" % (accuracy * 100.0)) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running this example we get the following output.
|
1 2 3 4 5 6 7 |
('X shape: : ', (285, 43)) XGBClassifier(base_score=0.5, colsample_bylevel=1, colsample_bytree=1, gamma=0, learning_rate=0.1, max_delta_step=0, max_depth=3, min_child_weight=1, missing=None, n_estimators=100, nthread=-1, objective='binary:logistic', reg_alpha=0, reg_lambda=1, scale_pos_weight=1, seed=0, silent=True, subsample=1) Accuracy: 71.58% |
Again we can see that the XGBoost framework chose the ‘binary:logistic‘ objective automatically, the right objective for this binary classification problem.
Support for Missing Data
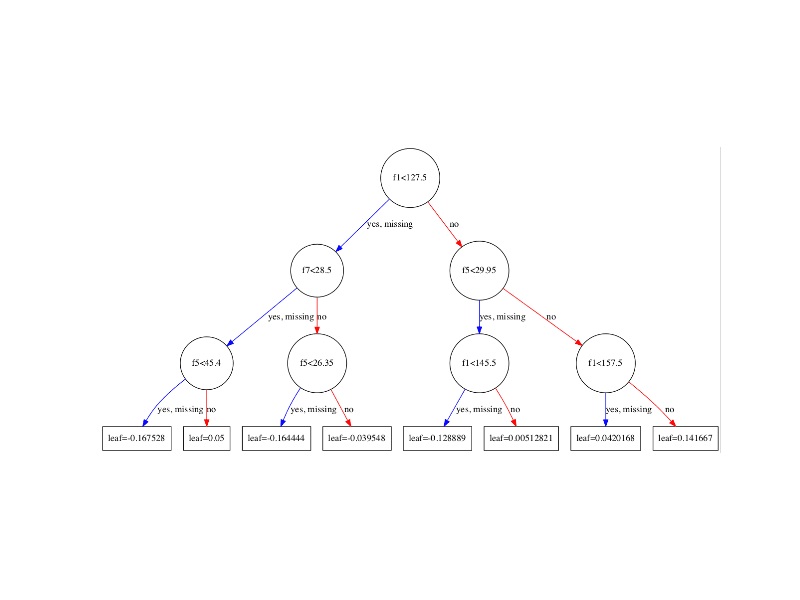
XGBoost can automatically learn how to best handle missing data.
In fact, XGBoost was designed to work with sparse data, like the one hot encoded data from the previous section, and missing data is handled the same way that sparse or zero values are handled, by minimizing the loss function.
For more information on the technical details for how missing values are handled in XGBoost, see Section 3.4 “Sparsity-aware Split Finding” in the paper XGBoost: A Scalable Tree Boosting System.
The Horse Colic dataset is a good example to demonstrate this capability as it contains a large percentage of missing data, approximately 30%.
Download the dataset and place it in your current working directory with the filename “horse-colic.csv“.
Below is a sample of the raw dataset.
|
1 2 3 4 5 6 |
2 1 530101 38.50 66 28 3 3 ? 2 5 4 4 ? ? ? 3 5 45.00 8.40 ? ? 2 2 11300 00000 00000 2 1 1 534817 39.2 88 20 ? ? 4 1 3 4 2 ? ? ? 4 2 50 85 2 2 3 2 02208 00000 00000 2 2 1 530334 38.30 40 24 1 1 3 1 3 3 1 ? ? ? 1 1 33.00 6.70 ? ? 1 2 00000 00000 00000 1 1 9 5290409 39.10 164 84 4 1 6 2 2 4 4 1 2 5.00 3 ? 48.00 7.20 3 5.30 2 1 02208 00000 00000 1 2 1 530255 37.30 104 35 ? ? 6 2 ? ? ? ? ? ? ? ? 74.00 7.40 ? ? 2 2 04300 00000 00000 2 ... |
The values are separated by whitespace and we can easily load it using the Pandas function read_csv.
|
1 |
dataframe = read_csv("horse-colic.csv", delim_whitespace=True, header=None) |
Once loaded, we can see that the missing data is marked with a question mark character (‘?’). We can change these missing values to the sparse value expected by XGBoost which is the value zero (0).
|
1 2 |
# set missing values to 0 X[X == '?'] = 0 |
Because the missing data was marked as strings, those columns with missing data were all loaded as string data types. We can now convert the entire set of input data to numerical values.
|
1 2 |
# convert to numeric X = X.astype('float32') |
Finally, this is a binary classification problem although the class values are marked with the integers 1 and 2. We model binary classification problems in XGBoost as logistic 0 and 1 values. We can easily convert the Y dataset to 0 and 1 integers using the LabelEncoder, as we did in the iris flowers example.
|
1 2 3 4 |
# encode Y class values as integers label_encoder = LabelEncoder() label_encoder = label_encoder.fit(Y) label_encoded_y = label_encoder.transform(Y) |
The full code listing is provided below for completeness.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 |
# binary classification, missing data from pandas import read_csv from xgboost import XGBClassifier from sklearn.model_selection import train_test_split from sklearn.metrics import accuracy_score from sklearn.preprocessing import LabelEncoder # load data dataframe = read_csv("horse-colic.csv", delim_whitespace=True, header=None) dataset = dataframe.values # split data into X and y X = dataset[:,0:27] Y = dataset[:,27] # set missing values to 0 X[X == '?'] = 0 # convert to numeric X = X.astype('float32') # encode Y class values as integers label_encoder = LabelEncoder() label_encoder = label_encoder.fit(Y) label_encoded_y = label_encoder.transform(Y) # split data into train and test sets seed = 7 test_size = 0.33 X_train, X_test, y_train, y_test = train_test_split(X, label_encoded_y, test_size=test_size, random_state=seed) # fit model no training data model = XGBClassifier() model.fit(X_train, y_train) print(model) # make predictions for test data y_pred = model.predict(X_test) predictions = [round(value) for value in y_pred] # evaluate predictions accuracy = accuracy_score(y_test, predictions) print("Accuracy: %.2f%%" % (accuracy * 100.0)) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running this example produces the following output.
|
1 2 3 4 5 6 |
XGBClassifier(base_score=0.5, colsample_bylevel=1, colsample_bytree=1, gamma=0, learning_rate=0.1, max_delta_step=0, max_depth=3, min_child_weight=1, missing=None, n_estimators=100, nthread=-1, objective='binary:logistic', reg_alpha=0, reg_lambda=1, scale_pos_weight=1, seed=0, silent=True, subsample=1) Accuracy: 83.84% |
We can tease out the effect of XGBoost’s automatic handling of missing values, by marking the missing values with a non-zero value, such as 1.
|
1 |
X[X == '?'] = 1 |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Re-running the example demonstrates a drop in accuracy for the model.
|
1 |
Accuracy: 79.80% |
We can also impute the missing data with a specific value.
It is common to use a mean or a median for the column. We can easily impute the missing data using the scikit-learn SimpleImputer class.
|
1 2 3 |
# impute missing values as the mean imputer = SimpleImputer() imputed_x = imputer.fit_transform(X) |
Below is the full example with missing data imputed with the mean value from each column.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 |
# binary classification, missing data, impute with mean import numpy from pandas import read_csv from xgboost import XGBClassifier from sklearn.model_selection import train_test_split from sklearn.metrics import accuracy_score from sklearn.preprocessing import LabelEncoder from sklearn.impute import SimpleImputer # load data dataframe = read_csv("horse-colic.csv", delim_whitespace=True, header=None) dataset = dataframe.values # split data into X and y X = dataset[:,0:27] Y = dataset[:,27] # set missing values to 0 X[X == '?'] = numpy.nan # convert to numeric X = X.astype('float32') # impute missing values as the mean imputer = SimpleImputer() imputed_x = imputer.fit_transform(X) # encode Y class values as integers label_encoder = LabelEncoder() label_encoder = label_encoder.fit(Y) label_encoded_y = label_encoder.transform(Y) # split data into train and test sets seed = 7 test_size = 0.33 X_train, X_test, y_train, y_test = train_test_split(imputed_x, label_encoded_y, test_size=test_size, random_state=seed) # fit model no training data model = XGBClassifier() model.fit(X_train, y_train) print(model) # make predictions for test data y_pred = model.predict(X_test) predictions = [round(value) for value in y_pred] # evaluate predictions accuracy = accuracy_score(y_test, predictions) print("Accuracy: %.2f%%" % (accuracy * 100.0)) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running this example we see results equivalent to the fixing the value to one (1). This suggests that at least in this case we are better off marking the missing values with a distinct value of zero (0) rather than a valid value (1) or an imputed value.
|
1 |
Accuracy: 79.80% |
It is a good lesson to try both approaches (automatic handling and imputing) on your data when you have missing values.
Summary
In this post you discovered how you can prepare your machine learning data for gradient boosting with XGBoost in Python.
Specifically, you learned:
- How to prepare string class values for binary classification using label encoding.
- How to prepare categorical input variables using a one hot encoding to model them as binary variables.
- How XGBoost automatically handles missing data and how you can mark and impute missing values.
Do you have any questions?
Ask your questions in the comments and I will do my best to answer.





hi Jason, the train data for the last example should be imputed_x, but you use the original X which has missing data. I tried with data imputed_x and get the accuracy 79.8%
Thanks Ralph. Fixed.
Hi Jason,
Thanks for the tutorial with such useful information! I have one question regarding the label encoding and the one hot encoding you applied on the breast cancer dataset.
You perform label encoding and one hot encoding for the whole dataset and then split into train and test set. This way it can be ensured that all the data are transformed with the same encoding configuration.
However, if we have new unseen data with the raw dataset type, how can we ensure that label encoding and one hot encoding is still transforming the unseen data in the same way? Do we need to save the encoders for the sake of processing unseen data?
Thanks in advance!
Great question Qichang.
I would prepare the encodings on the training data, store the mappings (or pickle the objects), then reuse the encodings on the test data.
It means we must be confident that the training data is representative of the data we may need to predict in the future.
how exactly would this be done?
Which part are you having trouble with?
Hi Jason,
I have a question.
how to prepare the encodings on the training data, store the mappings?
Thanks in advance!
You can save the python object directly, e.g. you can use pickle library.
Hi Jason,
Thanks for your reply. could you give me an example for preparing the encodings on the training data. I still don’t know how to do. For example, there is a category column named color, which have the values ‘red’, ‘blue’, and ‘black’, how to pickle ?
Besides, I have another question. when I trained xgboost model using the xgboost sklearn api, for example,
n_estimators = [50, 100, 150, 200]
max_depth = [2, 4, 6, 8]
k_neighbors = [7]
params = dict(clf__max_depth=max_depth, clf__n_estimators=n_estimators,
over__k_neighbors=k_neighbors)
over = select_smote.selectSmoteType(build_model_util.smote_type)
under = RandomUnderSampler(sampling_strategy=0.5)
if classifier_strategy == ‘gbc’:
clf_estimator = GradientBoostingClassifier()
rfe = RFECV(estimator=clf_estimator)
clf = XGBClassifier(objective=’binary:logistic’, use_label_encoder=False)
pipeline = Pipeline(steps=[(‘over’, over), (‘under’, under), (‘s’, rfe), (‘clf’, clf)])
skf = StratifiedKFold(n_splits=5, shuffle=True, random_state=1)
CV_result = build_model_util.train_estimator(
pipeline, params,
X, y,
skf, scoring=’roc_auc’)
occurred the warning: the default evaluation metric used with the objective ‘binary:logistic’ was changed from ‘error’ to ‘logloss’. Explicitly set eval_metric if you’d like to restore the old behavior. Why?
I am looking forward to your reply. Thanks in advance.
You can safely ignore that warning.
You can also learn more about xgboost loss functions here:
https://machinelearningmastery.com/xgboost-loss-functions/
Thanks Jason for the prompt reply.
Besides this kinds of data transformation, do we need to consider scaling or normalisation of the input variables before passing to XGBoost? We know that it generally yields better result for SVM especially with kernel function.
Generally no scaling. You may see some benefit by spreading out a univariate distribution to highlight specific features (e.g. with a square, log, square root, etc.)
Hi Jason, your post is very helpful. Thanks a lot!
I had a question around how to treat “default” values of continuous predictors for XGBoost. For example, let’s say attributer X may take continuous values as such (say in range of 1 -100). But certain records may have some default value (say 9999) which denotes certain segment of customers for whom that predictor X cannot be calculated or is unavailable. Can we directly use predictor X as input variable for an XGBoost model? Or, should we do some data treatment for X? If so, what would that be?
TIA
Great question JChen.
I would try modeling the data as-is to start with. XGBoost will figure it out.
You could then try some feature engineering (maybe add a new flag variable for such cases) and see if you can further list performance.
Thanks for your reply! This is helpful
I’m glad to hear it JChen.
For your missing data part you replaced ‘?’ with 0. But you have not mentioned while defining XGBClassifier model that in your dataset treat 0 as missing value. And by default ‘missing’ parameter value is none which is equivalent to treating NaN as missing value. So i don’t think your model is handling missing values.
Thanks for the note Sargam.
I get very much better mean accuracy results when source data has missing values as “nan” than putting them to zero (0). Can someone explain that in more detailed? Here and in other sources I have read that it is ok to put nan values to zero but I wonder does “sparsity aware split finding” -algorithm regonize them as nan values?
Yes, xgboost will model nan as a value.
Hi Jason,
I came to this page from another : IRIS ,
source: https://machinelearningmastery.com/multi-class-classification-tutorial-keras-deep-learning-library/#comment-396630
In that webPage, your code classifies IRIS with 96.6% accuracy, which is very good.
In comments section, you told this guy (Abhilash Menon) to use gradient boosting with XGBoost.
Which is this tutorial. Here is also IRIS classification.
Well I do not understand:
Here, the IRIS classification with gradient boosting with XGBoost yields 79% or 83% accuracy (only).
Why should we use gradient boosting with XGBoost then?
Accuracy is too low.
The tutorial is a demonstration of how to use the algorithm.
Your job as the machine learning practitioner is to try many algorithms on your problem and see what works best.
See this post:
https://machinelearningmastery.com/a-data-driven-approach-to-machine-learning/
Hi, Jason. First of all i would like to thank you for the wonderful material. I have been trying the XGBoost algorithm, it seems its acting weird on my pc. The first iris species dataset i got score of 21.2 %. Now this one of the breast cancer my accuracy is 2.1%. I really dont know wats wrong, can you please, please help me
I’m sorry to hear that. Perhaps the library is not installed correctly?
I would recommend posting a question to stackoverflow or even the xgboost users group.
Let me know how you go.
Hello Jason, I know that for regression models we should drop the first dummy variable to avoid the “dummy trap”. I don’t see you doing that in this case. Is is because the dummy variable trap only applies to linear regression models and not gradient boosting algorithms?
I believe the dummy variable trap applies with linear models and when the variables are multicollinear.
Hello Jason.
What happens if I have a string column with NA’s?
Should I first put 0’s instead of the NA’s and then use the OneHotEncounter ?
XGBoost can handle the NAs. No need to do anything.
If you do want to impute or similar, see this post:
https://machinelearningmastery.com/handle-missing-data-python/
Thank you, Jason.
Very useful article.
I met with library CatBoost which has interest options to configure categorical variables. Maybe, this will be interest for your journey.
Thanks, do you have a link?
Link on “parameter tuning” section: https://tech.yandex.com/catboost/doc/dg/concepts/parameter-tuning-docpage/#choosing-cat-features
You can see many interest materials on this site.
+ If it is interest for you, there are other interest libraries which are based on ensembles of trees. I.e., realization of genetic algorithm in “TPOT” (tree based pipeline optimization technique).
Thanks Alexander.
Dear Jason,
Thank you for the great tutorial!
Unfortunately I could not get the “datasets-uci-breast-cancer.csv” – looks like it was removed from the website. I search on the web but could not find a similar file to try out your example.
Is there a possibility you can send me this file?
Greatly appreciated
Dani
Here it is:
https://gist.github.com/jbrownlee/ceb34e3014be83da5f8a255c75b026d7
Thank you very much!
Do we need to deal with outliers before passing features to XGBoost?
Perhaps. Try with and without outlier removal and compare model skill.
Hi Jason,
Thanks for taking the time to map this all out. Prior to reading your tutorial, I used the DataCamp course on XGBoost as a guide, where they use two steps for encoding categorical variables: LabelEncoder followed by OneHotEncoder.
So a categorical variable with 5 levels is converted to values 0-4 and then these are one-hot encoded into columns five columns.
I am interested in the feature importance, so xgb.plot_importance is a great tool. However, the features are two steps removed from their original state.
How would you undo this two-step encoding to get the original variable names?
The one hot encoding can be reversed with an argmax() on each vector.
The label encoding has an inverse_transform() function in sklearn.
Could you discuss this and its’ applicability to XGBoost:
http://roamanalytics.com/2016/10/28/are-categorical-variables-getting-lost-in-your-random-forests/
Thanks for the link.
Jason, Do you have any idea about applying XGBoost on a multilabel classification problem? I just need some help from you on the data preparation part. Some suggestions from stackoverflow I found is massage your data a little by making k copies of every data point that has k correct/positive labels. Then you can hack your way to a simpler multi class problem…. Any suggestions would be highly appreciated!
Sorry, I don’t have any multi-label examples.
it’s been a year, have u got any now ??
I have an example of multi-label classification here:
https://machinelearningmastery.com/how-to-develop-a-convolutional-neural-network-to-classify-satellite-photos-of-the-amazon-rainforest/
Hi,i got a problem.What if the dataset is a mixture of MultiHot category ,OneHot category and continues and string type,which means all columns are of different shape .What should we do?Sklearn machine learning modules like SVM seems unable to handle this,i must reshape all columns into same shape.Is there a better solution ?Thank you!
I recommend transforming all data to numbers. E.g. some one “sample” is one vector of numbers.
Hello 🙂 Great article !!! May I have a question ? How to resolve case if for example new data for prediction will has some categorical value which is not in train data ? I mean for example feature “color”. Train data has values for “color” – white, green, black. So one hot encoder create three columns. But if in new data for prediction will be additional value for “color” – orange – than encoder creates 4 columns. When I use XGboostClassifier I get some error about missmatch.
You must ensure the training set is representative of the problem.
Or, prepare data with prior knowledge of all possible values.
So do you mean to add a column of 0’s for future categorical value (‘orange’) to the training set? And including that zero column wouldn’t change the predictive power for test data that doesn’t have this categorical value? That is, I want to know if this is correct: by adding the extra column just to prepare for an unseen category would make the model more inclusive/general.
You have many options, such as making room in the reprensetation, or marking unseen values as 0, or using different representation, etc. Select an approach that works best and meets the requirements of your project.
Old reply, but for anyone in the future that needs it:
OneHotEncoderhas thecategoriesparameter that will hold all “categories expected”. So, you need to create a list of all expected categories to feed into this (metadata on the column). This pairs well with the Pandas Categories datatype. Define the categories on each of your “object” type columns. There is a good example on this plot_stack_predictors page https://scikit-learn.org/stable/auto_examples/ensemble/plot_stack_predictors.html?highlight=categorical%20encodingThanks!
You can also configure it to ignore labels not seen during training.
Do you know what happens if I have a string column with NA’s?
If the entire column is NA, then you can safely remove it.
I really enjoy reading your hands-on posts Jason. Thanks for sharing them.
What is a better way to encode the numerical values- such as age groups, income range, etc.-to capture the ordinal relationship as well? I feel by one-hot-encoding those attributes we are throwing away some useful information.
Thanks.
Excellent question!
Integer encoding as a first step – see what results you get.
Embedding as a second step – I would expect better results.
Contrast both to a one hot encoding.
What do you mean exactly by embedding? Can you refer me to some references please?
Yes, perhaps start here:
https://machinelearningmastery.com/?s=embedding&post_type=post&submit=Search
similar to the methodology below?
https://medium.com/@krishnakalyan3/a-gentle-introduction-to-embedding-567d8738372b
Yes.
I have IP address field in the dataset. How do I encode it ? Can u please explain with an example ?? or send me links where can I get those type of examples.
Thank you in advance.
I don’t have an example.
Perhaps check the literature for common encoding schemes for ip addresses?
Hi,Jason
In the part of one hot encode categorical data, I used just the labelencoder encoding method for input x and then trained the model, I got the same performance as using the one hot encoding method.
Is it that the labelencoder method happened to be good here?
Yes, it is just fine for single variables. It does the same thing.
Hi Jason,
Is accuracy score in machine learning model for classification only ? Can I use Mean Square Error or Mean Absolute Error or R2 Error for classification also ?
Thank you for your advice.
Yes. Accuracy is only for classification.
MSE, MAE, R2 are for regression only.
Thanks for the reply.
Why is it in classification we look for Accuracy. Whereas in regression we look for Error ?
See this:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-classification-and-regression
Hi Jason,
For the binary classification, breast cancer example, find it very hard to understand why you need to do the following:
” … we must transform the feature array into a 2-dimensional NumPy array where each integer value is a feature vector with a length 1.”
feature = label_encoder.fit_transform(X[:,i])
feature = feature.reshape(X.shape[0], 1)
Hope you enlighten by more simpler layman term. Thank you.
In that example we encode each feature separately.
It would have been easier to use the new OrdinalEncoder.
Why after encode each feature separately it still needs to reshape it ?
To meet the expectations of the API.
How do you know each feature needs 2-dimensional NumPy array of the API ?
Thanks Jason.
One dimension for rows, one for columns.
Hi Jason,
After reading data from dataframe pandas, data must be converted to numpy array before fit to a model, is it ?
If after reading data from dataframe, doing fit and transform in scaling features, does it mean that data is automatically convert to array ?
Please pardon me as I am a novice in ML. Thank you for your advice.
Yes.
Yes.
Hi, I tested the code in your another post(https://machinelearningmastery.com/xgboost-for-regression/), xgboost can fit on pandas dataframe directly without convertig to numpy array:
I wrote:
X, y = dataframe.iloc[:,:-1], dataframe.iloc[:,-1:]
instead of:
data = dataframe.values
X, y = data[:, :-1], data[:, -1]
I don’t know if it is converted implicitly or not? Thank you.
Correct me if I’m wrong:
1. As a tree-based algorithm, XGBoost doesn’t require scaling.
2. I fitted gradient boosting decision trees on data with original categorical variables and on the same data with the dummy variables corresponding to each categorical value and got similar results. Is it safe to use just the categorical version (whether it has numeric 0, 1, 99, or string values ‘A0’, ‘A1’, ‘A99’) without one-hot encoding?
One hot encoding is not scaling.
You may be required to prepare categorical inputs before using xgboost using an ordinal or one hot encoding.
I mean scaling as in minmaxscaler() and standardscaler().
If scaling doesn’t matter, why is it important to break categories into dummy variables? I did see better results with just the raw categorical variables. It seemed to me tree based classifiers can map just any values fine, whether they are string or numeric values. Looking for more opinions about this topic.
Because the model cannot operate on labels directly, it requires inputs to be numbers.
Is it fair to say try mapping a categorical variable into numeric categories or to as many 0/1 dummy variables as there are categories and see which version of the categorical variable achieves better results?
You can try that.
how to predict the accuracy when target is continuous?
You cannot, see this:
https://machinelearningmastery.com/faq/single-faq/how-do-i-calculate-accuracy-for-regression
Hi Jason, for the Horse Colic Dataset example, why didn’t you one hot encode some of the inputs such as surgery or capillary refill time.
That sounds like a great extension! Try it and see if it helps.
When a model like in the article is developed, and I’d like to deploy and predict on new incoming data, we know that incoming data has not gone through the same embedding as training/testing data. How can I persist that embedding process and apply it to new incoming data? Maybe Using pipeline, could you point to a detailed example if possible? Thank you.
New data must be prepared in an identical manner to training data.
It is often easy to achieve by saving the data prep objects along with the model, or save the whole pipeline object.
Are there dataset size limitations with XGBoost? What is the minimum number of rows of data you would need in order to use XGBoost? Is there also a rule of thumb for ratio of number of rows and number of features. Of course you need more rows than features but is there a rule of thumb that you follow?
Good question, not really.
More data is generally better:
https://machinelearningmastery.com/faq/single-faq/how-much-training-data-do-i-need
Hi Jason. Your tutorials are great and super helpful.
I have a data set with several columns having mixed integer and categorical type data. Most of the values are integers and some are codes like ‘Q’ or ‘X’. What is the best way to preprocess these columns before feeding to an XGBoost?
Thanks.
Thanks!
Perhaps map to a new sequence of integers (integer/ordinal encode).
Is there a risk of data leakage when you did mean imputation on the whole dataset rather than the training sets?
Yes, ideally we would fit the transform on the training set and apply it to the train and test sets.
More details here:
https://machinelearningmastery.com/data-preparation-without-data-leakage/
I’ve never seen anyone label encode, then one-hot encode. Could you perhaps explain or share some supplemental materials to understand the pros/cons of this type of feature engineering method? specifically potential memory efficiencies?
My understanding is that this will cause label encoding will inherently force a ordinal relationship and cause xgboost to try to group observations together based on their respective cardinality (group .9-1 together and 0.01-0.1, due to their close proximity). Does this constraint correct itself if it is subsequently onehot encoded?
I’m currently working with an xgboost model with a very large dataset. after one-hot encoding, my dataset is 400kx2k and a HP grid search causes a complete failure due to memory allocation errors (machine has 48gb RAM).
You can often one hot encode directly these days. In the olden days (a years back) one had to integer/label encode then one hot encode.
One hot encoding can become inefficient with large numbers of categories and rows, a sparse representation can be used instead, or an alternate encoding such as embedding, hashing, etc. can be used:
https://machinelearningmastery.com/faq/single-faq/how-do-i-handle-a-large-number-of-categories
Thanks for this post Jason, it’s really helpful!
I have one regression case on which some of the features are continuous and take variables for example from 1 to 2’000.000 and others which are either dichotomous (1/0) or ordinal which take very specific values (0,1,2) and I’m observing that XGBoost is giving the latter two a lot less use vs the first type of features.
Is there a way to leverage better the latter in order to strengthen their use in the model please?
Appreciate your response!
You’re welcome.
The model will chose what is most useful to make predictions, no need to force the model to use variables that don’t help.
You can try providing copies of the variables with different transforms to see if the model can find a better way to use them.
just have a question on object saving, so, if there is a category column named color, which have the values ‘red’, ‘blue’, and ‘black’, how to pickle that? Thank you for reading my question
Perhaps this will help you save your data:
https://machinelearningmastery.com/how-to-save-a-numpy-array-to-file-for-machine-learning/
Hello Jason, thank you for your wonderful work, it has been of great help to me.
I have data with many nominal variables and missing variables.
I want to harvest XGboost own missing value feature, but Sklearns Onehot encoder can not handle missing values.
– A hack would be to keep track of the row number of missing values and then force the columns to nan’s.
– I have also thought of imputing missing values as “-1” and giving them their own category. i.e. a binary variable will have three levels then. but this will increase the variabel count tremendously.
How would you go about this?
You’re welcome.
Perhaps you can use the XGBoost API directly?
Perhaps you can mark missing values as a special value (-1) as per you suggested? You have 3 values already when one is missing.
Hi Jason,
Thanks for this example. I tried to use XGBoost without one-hot encoding (or label encoding) and it worked for Iris dataset. This is where I’m confused now. You mentioned that “XGBoost cannot model this problem as-is as it requires that the output variables be numeric.”. But it is working for me. Am I missing something here? Please clarify.
Perhaps your data had a numeric output?
Perhaps the API was changed?
Perhaps the model requires numeric input instead of numeric output?
I am training my model using XG boost and storing in blob, I am trying to predict it for another dataset and getting the following error.
ValueError: feature_names mismatch
Though my features are the same. Can you please suggest
Sorry, I have not seen that error. Perhaps try posting your code and error to stackoverflow.com
Hi Jason,
Can GBDT automatically handle missing data(nan value)?
I am looking forward to your reply. Thanks in advance.
XGBoost implementation can include missing data directly.
Hi, Jason,
I’m always looking forward your very helpful blog.
The last line in the 7th code box in “One Hot Encode Categorical Data” section says below.
# encode string input values as integers
:
encoded_x = encoded_x.reshape(X.shape[0], X.shape[1])
I’m wondering weather it is following.
encoded_x = encoded_x.T
Thank you,
I believe the code was correct. Why you want to do encodedd_x.T?
Hey I have a question about xg boosting.
Asymptotically (if we had a very large sample), does it
matter whether we fit prices or log p rices if we fit a linear model by OLS? And
if we fit a very flexible model by XGBoost?
OLS matters a lot between prices and log(prices), because log is nonlinear.
I believe XGBoost does not care it is log or not because it is a decision tree and log is a monotonic function.
how does a very large vs small sample size affect predictions with log vs not log prices in the OLS regression?
Probably it doesn’t matter for log vs not log, but more on how many parameters in your model. The usual ballpark is to get 30 times the number of parameters in your model.
Thanks so much for an outstanding tutorial. My dataset currently has categorical variables that are one-hot encoded (since the variables have numerous values associated with them), as well as binary variables with values that are contained in a single column (i.e., a column containing no missing values where 0=No and 1=Yes). When using XGBoost, is it OK for me to represent binary variables using a single column, rather than use one-hot encoding? If so, does XGBoost handle the zeros in that column as missing values?
Hi Erin…The following discussion may be of interest:
https://www.quora.com/How-does-XGBoost-treat-missing-values-during-training-and-prediction
Hi Jason
I have some data which are in 2d (latitude, longitude, var1, var2,…). I want to predict variable values on specific grid defined points. Is it possible with xgboost? Do you have example?
Thanks
Hi Dzevad…You may find the following of interest:
https://towardsdatascience.com/geopandas-hands-on-building-geospatial-machine-learning-pipeline-9ea8ae276a15