Machine learning algorithms are parameterized so that they can be best adapted for a given problem. A difficulty is that configuring an algorithm for a given problem can be a project in and of itself.
Like selecting ‘the best’ algorithm for a problem you cannot know before hand which algorithm parameters will be best for a problem. The best thing to do is to investigate empirically with controlled experiments.
The caret R package was designed to make finding optimal parameters for an algorithm very easy. It provides a grid search method for searching parameters, combined with various methods for estimating the performance of a given model.
In this post you will discover 5 recipes that you can use to tune machine learning algorithms to find optimal parameters for your problems using the caret R package.
Kick-start your project with my new book Machine Learning Mastery With R, including step-by-step tutorials and the R source code files for all examples.
Let’s get started.
Model Tuning
The caret R package provides a grid search where it or you can specify the parameters to try on your problem. It will trial all combinations and locate the one combination that gives the best results.
The examples in this post will demonstrate how you can use the caret R package to tune a machine learning algorithm.
The Learning Vector Quantization (LVQ) will be used in all examples because of its simplicity. It is like k-nearest neighbors, except the database of samples is smaller and adapted based on training data. It has two parameters to tune, the number of instances (codebooks) in the model called the size, and the number of instances to check when making predictions called k.
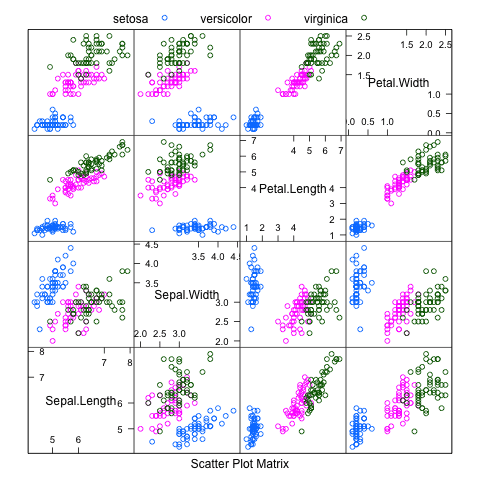
Each example will also use the iris flowers dataset, that comes with R. This classification dataset provides 150 observations for three species of iris flower and their petal and sepal measurements in centimeters.
Each example also assumes that we are interested in the classification accuracy as the metric we are optimizing, although this can be changed. Also, each example estimates the performance of a given model (size and k parameter combination) using repeated n-fold cross validation, with 10 folds and 3 repeats. This too can be changed if you like.
Need more Help with R for Machine Learning?
Take my free 14-day email course and discover how to use R on your project (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
Grid Search: Automatic Grid
There are two ways to tune an algorithm in the Caret R package, the first is by allowing the system to do it automatically. This can be done by setting the tuneLength to indicate the number of different values to try for each algorithm parameter.
This only supports integer and categorical algorithm parameters, and it makes a crude guess as to what values to try, but it can get you up and running very quickly.
The following recipe demonstrates the automatic grid search of the size and k attributes of LVQ with 5 (tuneLength=5) values of each (25 total models).
|
1 2 3 4 5 6 7 8 9 10 11 12 |
# ensure results are repeatable set.seed(7) # load the library library(caret) # load the dataset data(iris) # prepare training scheme control <- trainControl(method="repeatedcv", number=10, repeats=3) # train the model model <- train(Species~., data=iris, method="lvq", trControl=control, tuneLength=5) # summarize the model print(model) |
The final values used for the model were size = 10 and k = 1.
Grid Search: Manual Grid
The second way to search algorithm parameters is to specify a tune grid manually. In the grid, each algorithm parameter can be specified as a vector of possible values. These vectors combine to define all the possible combinations to try.
The recipe below demonstrates the search of a manual tune grid with 4 values for the size parameter and 5 values for the k parameter (20 combinations).
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 |
# ensure results are repeatable set.seed(7) # load the library library(caret) # load the dataset data(iris) # prepare training scheme control <- trainControl(method="repeatedcv", number=10, repeats=3) # design the parameter tuning grid grid <- expand.grid(size=c(5,10,20,50), k=c(1,2,3,4,5)) # train the model model <- train(Species~., data=iris, method="lvq", trControl=control, tuneGrid=grid) # summarize the model print(model) |
The final values used for the model were size = 50 and k = 5.
Data Pre-Processing
The dataset can be preprocessed as part of the parameter tuning. It is important to do this within the sample used to evaluate each model, to ensure that the results account for all the variability in the test. If the data set say normalized or standardized before the tuning process, it would have access to additional knowledge (bias) and not give an as accurate estimate of performance on unseen data.
The attributes in the iris dataset are all in the same units and generally the same scale, so normalization and standardization are not really necessary. Nevertheless, the example below demonstrates tuning the size and k parameters of LVQ while normalizing the dataset with preProcess=”scale”.
|
1 2 3 4 5 6 7 8 9 10 11 12 |
# ensure results are repeatable set.seed(7) # load the library library(caret) # load the dataset data(iris) # prepare training scheme control <- trainControl(method="repeatedcv", number=10, repeats=3) # train the model model <- train(Species~., data=iris, method="lvq", preProcess="scale", trControl=control, tuneLength=5) # summarize the model print(model) |
The final values used for the model were size = 8 and k = 6.
Parallel Processing
The caret package supports parallel processing in order to decrease the compute time for a given experiment. It is supported automatically as long as it is configured. In this example we load the doMC package and set the number of cores to 4, making available 4 worker threads to caret when tuning the model. This is used for the loops for the repeats of cross validation for each parameter combination.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 |
# ensure results are repeatable set.seed(7) # configure multicore library(doMC) registerDoMC(cores=4) # load the library library(caret) # load the dataset data(iris) # prepare training scheme control <- trainControl(method="repeatedcv", number=10, repeats=3) # train the model model <- train(Species~., data=iris, method="lvq", trControl=control, tuneLength=5) # summarize the model print(model) |
The results are the same as the first example, only completed faster.
Visualization of Performance
It can be useful to graph the performance of different algorithm parameter combinations to look for trends and the sensitivity of the model. Caret supports graphing the model directly which will compare the accuracy of different algorithm combinations.
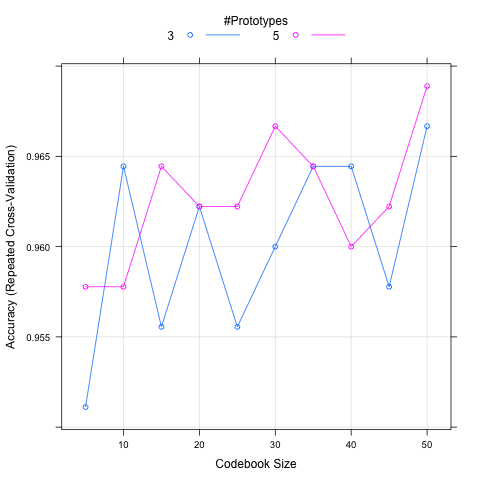
In the recipe below, a larger manual grid of algorithm parameters are defined and the results are graphed. The graph shows the size on the x axis and model accuracy on the y axis. Two lines are drawn, one for each k value. The graph shows the general trends in the increase in performance with size and that the larger value of k is probably preferred.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 |
# ensure results are repeatable set.seed(7) # load the library library(caret) # load the dataset data(iris) # prepare training scheme control <- trainControl(method="repeatedcv", number=10, repeats=3) # design the parameter tuning grid grid <- expand.grid(size=c(5,10,15,20,25,30,35,40,45,50), k=c(3,5)) # train the model model <- train(Species~., data=iris, method="lvq", trControl=control, tuneGrid=grid) # summarize the model print(model) # plot the effect of parameters on accuracy plot(model) |
The final values used for the model were size = 35 and k = 5.
Summary
In this post you have discovered the support in the caret R package for tuning algorithm parameters by using a grid search.
You have seen 5 recipes using the caret R package for tuning the size and k parameters for the LVQ algorithm.
Each recipe in this post is standalone and ready for you to copy-and-paste into your own project and adapt to your problem.




Thanks for great article! Can you give an example for parallel processing in Windows using Caret package, because doMC package is available only for Unix?
doParallel
Thanks, Jason. A great summary of what’s available and how to use them.
Is this applicable to regression problems?
Yes, it could be.
If my response variable is not categorical then what will be the method? Here you use Learning vector quantization method. model <- train(Species~., data=iris, method="lvq", trControl=control, tuneLength=5)??
Is there also a dedicated multi core setting for the python stack widely used in the other tutorials on this site?
I can’t use CUDA.
Yes, I believe the n_jobs argument to most sklearn functions.
what does the codebook size or the size parameter refer to ?
Codebooks refers to the learning capacity of the model.
I have started learning models. Your article is well written. Thanks 🙂
Thanks, I’m glad it helped.
Thanks so much for this! very helpful
Glad to hear it.
Thanks for the nice article! It is very helpful to find articles about Machine Learning so well written.
I have related question: do you know if it’s possible to give different tuning parameters for different classifiers in a multiclass problem, e.g., giving different C values to different SVMs using SVMRadial with the train function in Caret?
Yes, you can provide the parameters directly to the models.
Hi Jason,
thank you for the quick reply. So, how does one specify different cost values for different classifiers in the same train function? I haven’t found an example of this functionality yet.
Thanks in advance and best regards.
You can specify algorithm parameters in the tuneGrid argument when evaluating the model. It does not have to be a grid, it can be one set of params.
Hi Sir, great post!
How do you indicate CARET to use sensitivity insted of accuracy to select the best model?
You can specify the ‘metric’ argument, more details here:
https://machinelearningmastery.com/machine-learning-evaluation-metrics-in-r/
Could you clarify what happens under the hood in the train() function when specifying a grid? I am curious about the output model from this line.
a) Is the model that is the output from this line the BEST model with the lowest error amongst the validation sets in the cv framework? I.e. if the grid search is run with 5 fold CV, then the output model would be ONE of the FIVE possible models?
b) All validations splits are assessed and their error metrics are averaged, the best performing C from the grid is determined and subsequently, this C is run on ALL training data and the resulting model that is the output? I.e. using the above example, for C=1 and C=10 in the grid, the five and five ROC AUC results will be averaged and if C=10 is the winner, C=10 will be used to re-run the model on all training data not using CV and that model is the output?
I am running nested CV and tuning hyperparameters in the inner CV – there is a model output which I am transferring to my test set, but I did not find a clear answer so far about what this model really is.
I’m not sure off hand, perhaps post to the R user group?
Hi, posted the question on the caret GitHub page, but seems like the best parameters are refitted to the whole training set and that model is then output (and can be subsequently used in a test set to predict)
“We recommend using resampling to evaluate each distinct parameter value combination to get good estimates of how well each candidate performs. Once the results are calculated, the “best” tuning parameter combination is chosen and the final model is fit to the entire training set with this value.” from https://bookdown.org/max/FES/model-optimization-and-tuning.html
and here: https://github.com/topepo/caret/issues/995
Nice!
I love to read hard copy of books for proper study
Thanks.
sir, I need your help.
1) This same post can you post for python also please ?
2) How to perform Grid Search: Automatic Grid in python ?
3) Because i find only Grid Search: Manual Grid in pyton and I think there is no concept of Grid Search: Automatic Grid in python if any alternative or same is there please let me know.
Yes, see this post:
https://machinelearningmastery.com/how-to-tune-algorithm-parameters-with-scikit-learn/
Sir, You are absolutly right but i want to run logit model from statsmodel and want to obtain summary through RandomizedSearchCV.
import statsmodels.formula.api as smf
cv = RepeatedStratifiedKFold(n_splits=5,n_repeats=2,
random_state=True)
param_grid = {‘alpha’: sp_rand()}
# create and fit a logit model, testing random alpha values
logit_model=smf.logit(“Target~ ” + all_columns,train)
rsearch = RandomizedSearchCV(estimator=logit_model1, param_distributions=param_grid,cv=cv, n_iter=100) #param_distributions=param_grid
l1=rsearch.fit()
error : fit() missing 1 required positional argument: ‘X’
but here X is not req because we have already fitted a formula above.
I have some suggestions here:
https://machinelearningmastery.com/faq/single-faq/can-you-read-review-or-debug-my-code