Ensembles can give you a boost in accuracy on your dataset.
In this post you will discover how you can create some of the most powerful types of ensembles in Python using scikit-learn.
This case study will step you through Boosting, Bagging and Majority Voting and show you how you can continue to ratchet up the accuracy of the models on your own datasets.
Kick-start your project with my new book Machine Learning Mastery With Python, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.
- Update Jan/2017: Updated to reflect changes to the scikit-learn API in version 0.18.
- Update Mar/2018: Added alternate link to download the dataset as the original appears to have been taken down.

Ensemble Machine Learning Algorithms in Python with scikit-learn
Photo by The United States Army Band, some rights reserved.
Combine Model Predictions Into Ensemble Predictions
The three most popular methods for combining the predictions from different models are:
- Bagging. Building multiple models (typically of the same type) from different subsamples of the training dataset.
- Boosting. Building multiple models (typically of the same type) each of which learns to fix the prediction errors of a prior model in the chain.
- Voting. Building multiple models (typically of differing types) and simple statistics (like calculating the mean) are used to combine predictions.
This post will not explain each of these methods.
It assumes you are generally familiar with machine learning algorithms and ensemble methods and that you are looking for information on how to create ensembles in Python.
Need help with Machine Learning in Python?
Take my free 2-week email course and discover data prep, algorithms and more (with code).
Click to sign-up now and also get a free PDF Ebook version of the course.
About the Recipes
Each recipe in this post was designed to be standalone. This is so that you can copy-and-paste it into your project and start using it immediately.
A standard classification problem used to demonstrate each ensemble algorithm is the Pima Indians onset of diabetes dataset. It is a binary classification problem where all of the input variables are numeric and have differing scales.
You can learn more about the dataset here:
Each ensemble algorithm is demonstrated using 10 fold cross validation, a standard technique used to estimate the performance of any machine learning algorithm on unseen data.
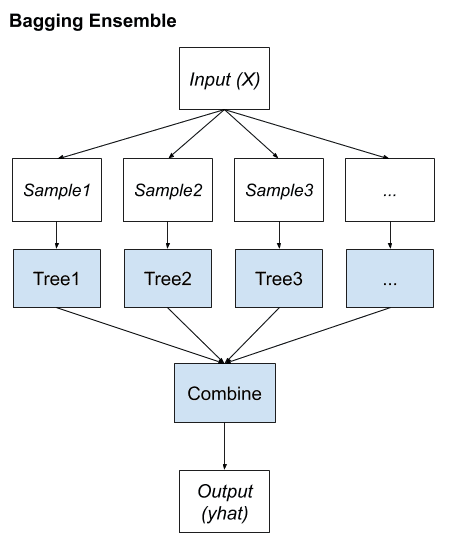
Bagging Algorithms
Bootstrap Aggregation or bagging involves taking multiple samples from your training dataset (with replacement) and training a model for each sample.
The final output prediction is averaged across the predictions of all of the sub-models.
The three bagging models covered in this section are as follows:
- Bagged Decision Trees
- Random Forest
- Extra Trees
1. Bagged Decision Trees
Bagging performs best with algorithms that have high variance. A popular example are decision trees, often constructed without pruning.
In the example below see an example of using the BaggingClassifier with the Classification and Regression Trees algorithm (DecisionTreeClassifier). A total of 100 trees are created.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 |
# Bagged Decision Trees for Classification import pandas from sklearn import model_selection from sklearn.ensemble import BaggingClassifier from sklearn.tree import DecisionTreeClassifier url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/pima-indians-diabetes.data.csv" names = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class'] dataframe = pandas.read_csv(url, names=names) array = dataframe.values X = array[:,0:8] Y = array[:,8] seed = 7 kfold = model_selection.KFold(n_splits=10, random_state=seed) cart = DecisionTreeClassifier() num_trees = 100 model = BaggingClassifier(base_estimator=cart, n_estimators=num_trees, random_state=seed) results = model_selection.cross_val_score(model, X, Y, cv=kfold) print(results.mean()) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running the example, we get a robust estimate of model accuracy.
|
1 |
0.770745044429 |
2. Random Forest
Random forest is an extension of bagged decision trees.
Samples of the training dataset are taken with replacement, but the trees are constructed in a way that reduces the correlation between individual classifiers. Specifically, rather than greedily choosing the best split point in the construction of the tree, only a random subset of features are considered for each split.
You can construct a Random Forest model for classification using the RandomForestClassifier class.
The example below provides an example of Random Forest for classification with 100 trees and split points chosen from a random selection of 3 features.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 |
# Random Forest Classification import pandas from sklearn import model_selection from sklearn.ensemble import RandomForestClassifier url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/pima-indians-diabetes.data.csv" names = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class'] dataframe = pandas.read_csv(url, names=names) array = dataframe.values X = array[:,0:8] Y = array[:,8] seed = 7 num_trees = 100 max_features = 3 kfold = model_selection.KFold(n_splits=10, random_state=seed) model = RandomForestClassifier(n_estimators=num_trees, max_features=max_features) results = model_selection.cross_val_score(model, X, Y, cv=kfold) print(results.mean()) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running the example provides a mean estimate of classification accuracy.
|
1 |
0.770727956254 |
3. Extra Trees
Extra Trees are another modification of bagging where random trees are constructed from samples of the training dataset.
You can construct an Extra Trees model for classification using the ExtraTreesClassifier class.
The example below provides a demonstration of extra trees with the number of trees set to 100 and splits chosen from 7 random features.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 |
# Extra Trees Classification import pandas from sklearn import model_selection from sklearn.ensemble import ExtraTreesClassifier url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/pima-indians-diabetes.data.csv" names = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class'] dataframe = pandas.read_csv(url, names=names) array = dataframe.values X = array[:,0:8] Y = array[:,8] seed = 7 num_trees = 100 max_features = 7 kfold = model_selection.KFold(n_splits=10, random_state=seed) model = ExtraTreesClassifier(n_estimators=num_trees, max_features=max_features) results = model_selection.cross_val_score(model, X, Y, cv=kfold) print(results.mean()) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running the example provides a mean estimate of classification accuracy.
|
1 |
0.760269993165 |
Boosting Algorithms
Boosting ensemble algorithms creates a sequence of models that attempt to correct the mistakes of the models before them in the sequence.
Once created, the models make predictions which may be weighted by their demonstrated accuracy and the results are combined to create a final output prediction.
The two most common boosting ensemble machine learning algorithms are:
- AdaBoost
- Stochastic Gradient Boosting
1. AdaBoost
AdaBoost was perhaps the first successful boosting ensemble algorithm. It generally works by weighting instances in the dataset by how easy or difficult they are to classify, allowing the algorithm to pay or or less attention to them in the construction of subsequent models.
You can construct an AdaBoost model for classification using the AdaBoostClassifier class.
The example below demonstrates the construction of 30 decision trees in sequence using the AdaBoost algorithm.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 |
# AdaBoost Classification import pandas from sklearn import model_selection from sklearn.ensemble import AdaBoostClassifier url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/pima-indians-diabetes.data.csv" names = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class'] dataframe = pandas.read_csv(url, names=names) array = dataframe.values X = array[:,0:8] Y = array[:,8] seed = 7 num_trees = 30 kfold = model_selection.KFold(n_splits=10, random_state=seed) model = AdaBoostClassifier(n_estimators=num_trees, random_state=seed) results = model_selection.cross_val_score(model, X, Y, cv=kfold) print(results.mean()) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running the example provides a mean estimate of classification accuracy.
|
1 |
0.76045796309 |
2. Stochastic Gradient Boosting
Stochastic Gradient Boosting (also called Gradient Boosting Machines) are one of the most sophisticated ensemble techniques. It is also a technique that is proving to be perhaps of the the best techniques available for improving performance via ensembles.
You can construct a Gradient Boosting model for classification using the GradientBoostingClassifier class.
The example below demonstrates Stochastic Gradient Boosting for classification with 100 trees.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 |
# Stochastic Gradient Boosting Classification import pandas from sklearn import model_selection from sklearn.ensemble import GradientBoostingClassifier url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/pima-indians-diabetes.data.csv" names = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class'] dataframe = pandas.read_csv(url, names=names) array = dataframe.values X = array[:,0:8] Y = array[:,8] seed = 7 num_trees = 100 kfold = model_selection.KFold(n_splits=10, random_state=seed) model = GradientBoostingClassifier(n_estimators=num_trees, random_state=seed) results = model_selection.cross_val_score(model, X, Y, cv=kfold) print(results.mean()) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running the example provides a mean estimate of classification accuracy.
|
1 |
0.764285714286 |
Voting Ensemble
Voting is one of the simplest ways of combining the predictions from multiple machine learning algorithms.
It works by first creating two or more standalone models from your training dataset. A Voting Classifier can then be used to wrap your models and average the predictions of the sub-models when asked to make predictions for new data.
The predictions of the sub-models can be weighted, but specifying the weights for classifiers manually or even heuristically is difficult. More advanced methods can learn how to best weight the predictions from submodels, but this is called stacking (stacked generalization) and is currently not provided in scikit-learn.
You can create a voting ensemble model for classification using the VotingClassifier class.
The code below provides an example of combining the predictions of logistic regression, classification and regression trees and support vector machines together for a classification problem.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 |
# Voting Ensemble for Classification import pandas from sklearn import model_selection from sklearn.linear_model import LogisticRegression from sklearn.tree import DecisionTreeClassifier from sklearn.svm import SVC from sklearn.ensemble import VotingClassifier url = "https://raw.githubusercontent.com/jbrownlee/Datasets/master/pima-indians-diabetes.data.csv" names = ['preg', 'plas', 'pres', 'skin', 'test', 'mass', 'pedi', 'age', 'class'] dataframe = pandas.read_csv(url, names=names) array = dataframe.values X = array[:,0:8] Y = array[:,8] seed = 7 kfold = model_selection.KFold(n_splits=10, random_state=seed) # create the sub models estimators = [] model1 = LogisticRegression() estimators.append(('logistic', model1)) model2 = DecisionTreeClassifier() estimators.append(('cart', model2)) model3 = SVC() estimators.append(('svm', model3)) # create the ensemble model ensemble = VotingClassifier(estimators) results = model_selection.cross_val_score(ensemble, X, Y, cv=kfold) print(results.mean()) |
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
Running the example provides a mean estimate of classification accuracy.
|
1 |
0.712166780588 |
Summary
In this post you discovered ensemble machine learning algorithms for improving the performance of models on your problems.
You learned about:
- Bagging Ensembles including Bagged Decision Trees, Random Forest and Extra Trees.
- Boosting Ensembles including AdaBoost and Stochastic Gradient Boosting.
- Voting Ensembles for averaging the predictions for any arbitrary models.
Do you have any questions about ensemble machine learning algorithms or ensembles in scikit-learn? Ask your questions in the comments and I will do my best to answer them.





Informative post.
Once you identify and finalize the best ensemble model, how would you score a future sample with such model? I am referring to the productionalization of the model in a data base.
Would you use something like the pickle package? Or is there a way to spell out the scoring algorithm (IF-ELSE rules for decision tree, or the actual formula for logistic regression) and use the formula for future scoring purposes?
After you finalize the model you can incorporate it into an application on service.
The model would be used directly.
It would be provided input patterns and make predictions that you could use in some operational way.
Hi Jason,
I wrote the code below. However, in your snippet, I see that you did not specify “base_estimator” in the AdaBoostClassifier. Any particular reason? Is there a default value for this parameter (CART??)?
#Boosting – AdaBoost algo
num_trees4 = 30
cart2 = DecisionTreeClassifier()
model4 = AdaBoostClassifier(base_estimator=cart2, n_estimators=num_trees4,random_state=seed)
results4 = cross_val_score(model4, X, Y, cv=kfold, scoring=scoring)
print(‘AdaBoost – Accuracy: %f’)%(results4.mean())
Thank you so much !
Sarra
The default is DecisionTreeClassifier, see:
http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.AdaBoostClassifier.html
Very well written post! Is there any email we could send you some questions about the ensemble methods?
Thanks Christos.
You can contact me directly here:
https://machinelearningmastery.com/contact/
HI Jason,
I,ve copy and paste your Random Forest and then result is:
0.766814764183.
We don’t have the same result, could you tell me why?
Thanks
I try to fix the random number seed Kamagne, but sometimes things get through.
The performance of any machine learning algorithm is stochastic, we estimate performance in the range. It is best practice to run a give configuration many times and take the mean and standard deviation – reporting the range of expected performance on unseen data.
Does that help?
The ensembeled model gave lower accuracy compared to the individual models. Isn’t strange?
This can happen. Ensembles are not a sure-thing to better performance.
In your Bagging Classifier you used Decision Tree Classifier as your base estimator. I was wondering what other algorithms can be used as base estimators?
Good question Marido,
With bagging, the goal is to use a method that has high variance when trained on different data.
Un-pruned decision trees can do this (and can be made to do it even better – see random forest).
Another idea would be knn with a small k.
In fact, take your favorite algorithm and configure it to have a high variance, then bag it.
I hope that helps.
Hi Jason,
Thanks so much for your insightful replies.
What I understand is that ensembles improve the result if they make different mistakes.
In my below result of two models. The first model performs well in one class while the second model performs well on the other class. When I ensemble them, I get lower accuracy. Is that possible or I am doing something wrong.
GBC
[[922035 266]
[ 2 5]]
cart
[[895914 26387]
[ 0 7]]
This is how how I am doing it.
estimators = []
model1 = GradientBoostingClassifier()
estimators.append((‘GBC’, model1))
model2 = DecisionTreeClassifier()
estimators.append((‘cart’, model2))
ensemble = VotingClassifier(estimators)
ensemble.fit(X_train, Y_train)
predictions = ensemble.predict(X_test)
accuracy1 = accuracy_score(Y_test, predictions)
Hi Natheer,
There is no guarantee for ensembles to lift performance.
Perhaps you can try a more sophisticated method to combine the predictions or perhaps try more or different submodels.
Hello Jason,
Many thanks for your post. It is possible to have two different base estimators (i.e. decision tree, knn) in AdaBoost model?
Thank you.
Not that I am aware in Python Djib.
I don’t see why not in theory. You could develop your own implementation and see how it fairs.
Thank you Jason
Hello Jason,
Why max_features is 3? what is it exactly? Thank you 😀
Just for demonstration purposes. It limits the number of selected features to 3.
I found one slight mishap. In the voting ensemble code, I notice is that in the voting ensemble code, on lines 22 and 23 it has
model3 = SVC()
estimators.append((‘svm’, model2))
I believe it should instead say model3 instead of model2, as model 3 is the svm stuff.
Thanks, fixed.
Hey! Thanks for the great post.
I would like to use voting with SVM as you did, however scaling data SVM gives me better results and it’s simply much faster. And from here comes the question: How can I scale just parto of the data for algorithms such as SVM, and leave non-slcaed data for XGB/Random forest and on top of it use ensembles. I have tried using Pipeline to first scale the data for SVM and then use Voting but it seams not working. Any comment would be helpful
Great question.
You could leave the pipeline behind and put the pieces together yourself with a scaled version of the data for SVM and non-scaled for the other methods.
Could we take it further and build a Neural Network model with Keras and use it in the Voting based Ensemble learning? (After wrapping the Neural Network model into a Scikit Learn classifier)
Sure, try it and let me know how you go.
Hi Jason…Thanks for the wonderful post. Is there a way to perform bagging on neural net regression? A sample code or example would be much appreciated. Thanks.
Yes. It may perform quite well. Sorry I do not have an example.
Hi Jason!
Thank you for the great post!
I would like to make soft voting for a convolutional neural network and a gru recurrent neural network, but i have 2 problems.
1: I have 2 different training datasets to train my networks on: vectors of prosodic data, and word embeddings of textual data. The 2 training sets are stored in two different np.arrays with different dimensionality. Is there any way to make VotingClassifier accept X1,and X2 except of a single X? (y is the same for both X1 and X2, and naturally they are of the same length)
2: where do i compile my networks?
ensemble = VotingClassifier(estimators)
ensemble.compile()
?
I would be very grateful for any help.
Thank you!
You have a few options.
You could get each model to generate the predictions, save them to a file, then have another model learn how to combine the predictions, perhaps with or without the original inputs.
If the original inputs are high-dimensional (images and sequences), you could try training a neural net to combine the predictions as part of training each sub-model. You can merge each network using a Merge layer in Keras (deep learning library), if your sub-models were also developed in Keras.
I hope that helps as a start.
Thanks for the quick reply! I ve already tried the layer merging. It works, but not giving good results because one of my feature sets yields significantly better recognition accuracy than the other.
But the first solution looks good! I will try and implement it!
Thanks for the help!
Nice work Peter!
I have a doubt. The above result is for training model accuracy. How to find the testing model accuracy for bagging classifier
from sklearn import model_selection
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
kfold=model_selection.KFold(n_splits=10)
dt=DecisionTreeClassifier()
model=BaggingClassifier(base_estimator=dt,n_estimators=10,random_state=5)
result=model_selection.cross_val_score(model,x,y,cv=kfold)
print(result.mean())
I am getting the accuracy for training model . But when I tried to get the testing accuracy for the model. I tried the below model. But I am getting the error. Please correct it
from sklearn.metrics import classification_report,confusion_matrix
from sklearn.metrics import accuracy_score
print(classification_report(ytest,result.predict(xtest)))
No, it evaluates the model using cross-validation:
https://machinelearningmastery.com/k-fold-cross-validation/
Can I use more than one base estimator in Bagging and adaboost eg Bagging(Knn, Logistic Regression, etc)?
You can do anything, but really this is only practical with bagging.
You want to pick base estimators that have low bias/high variance, like k=1 kNN, decision trees without pruning or decision stumps, etc.
what is the meaning of seed here? can you explain the importance of seed and how can some changes in the seed will affect the model? Thanks
Seed makes the example reproducible so you get the same results as me.
Machine learning algorithms are stochastic, meaning they give different results each time they are run. This is a feature, not a bug. See this post for more details:
https://machinelearningmastery.com/randomness-in-machine-learning/
Does that help?
hi sir
i am unable to run the gradient boosting code on my dataset.
please help me.
see my parameters.
AGE Haemoglobin RBC Hct Mcv Mch Mchc Platelets WBC Granuls Lymphocytes Monocytes disese
3 9.6 4.2 28.2 67 22.7 33.9 3.75 5800 44 50 6 Positive
11 12.1 4.3 33.7 78 28.2 36 2.22 6100 73 23 4 Positive
2 9.5 4.1 27.9 67 22.8 34 3.64 5100 64 32 4 Positive
4 9.9 3.9 27.8 71 25.3 35.6 2.06 4900 65 32 3 Positive
14 10.7 4.4 31.2 70 24.2 34.4 3 7600 50 44 6 Negative
7 9.8 4.2 28 66 23.2 35.1 1.95 3800 28 63 9 Negative
8 14.6 5 39.2 77 28.7 37.2 3.06 4400 58 36 6 Negative
4 12 4.5 33.3 74 26.5 35.9 5.28 9500 40 54 6 Negative
2 11.2 4.6 32.7 70 24.1 34.3 2.98 8800 38 58 4 Negative
1 9.1 4 27.2 67 22.4 33.3 3.6 5300 40 55 5 Negative
11 14.8 5.8 42.5 72 25.1 34.8 4.51 17200 75 20 5 Negative
What is the problem?
hi sir ,
problem is first i want to balance the dataset with SMOTE algorithm but it is not happening.
see my code and help me.
import pandas
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.datasets import make_classification
from sklearn.decomposition import PCA
from imblearn.over_sampling import SMOTE
print(__doc__)
sns.set()
# Define some color for the plotting
almost_black = ‘#262626’
palette = sns.color_palette()
data = (‘mdata.csv’)
dataframe = pandas.read_csv(data)
array = dataframe.values
X = array[:,0:12]
y = array[:,12]
#print (X_train, Y_train)
# Generate the dataset
#X, y = make_classification(n_classes=2, class_sep=2, weights=[0.1, 0.9],
# n_informative=3, n_redundant=1, flip_y=0,
# n_features=10, n_clusters_per_class=1,
# n_samples=500, random_state=10)
print (X, y)
plt.show()
# Instanciate a PCA object for the sake of easy visualisation
pca = PCA(n_components=2)
# Fit and transform x to visualise inside a 2D feature space
X_vis = pca.fit_transform(X)
# Apply regular SMOTE
sm = SMOTE(kind=’regular’)
X_resampled, y_resampled = sm.fit_sample(X, y)
X_res_vis = pca.transform(X_resampled)
# Two subplots, unpack the axes array immediately
f, (ax1, ax2) = plt.subplots(1, 2)
ax1.scatter(X_vis[y == 0, 0], X_vis[y == 0, 1], label=”Class #0″, alpha=0.5,
edgecolor=almost_black, facecolor=palette[0], linewidth=0.15)
ax1.scatter(X_vis[y == 1, 0], X_vis[y == 1, 1], label=”Class #1″, alpha=0.5,
edgecolor=almost_black, facecolor=palette[2], linewidth=0.15)
ax1.set_title(‘Original set’)
ax2.scatter(X_res_vis[y_resampled == 0, 0], X_res_vis[y_resampled == 0, 1],
label=”Class #0″, alpha=.5, edgecolor=almost_black,
facecolor=palette[0], linewidth=0.15)
ax2.scatter(X_res_vis[y_resampled == 1, 0], X_res_vis[y_resampled == 1, 1],
label=”Class #1″, alpha=.5, edgecolor=almost_black,
facecolor=palette[2], linewidth=0.15)
ax2.set_title(‘SMOTE ALGORITHM – Malaria regular’)
“””
daata after resample
“””
print (X_resampled, y_resampled)
plt.show()
Sorry, I cannot debug your code for you. Perhaps you can post your code to stackoverflow?
i am getting the error :
File “/usr/local/lib/python2.7/dist-packages/imblearn/over_sampling/smote.py”, line 360, in _sample_regular
nns = self.nn_k_.kneighbors(X_class, return_distance=False)[:, 1:]
File “/home/sajana/.local/lib/python2.7/site-packages/sklearn/neighbors/base.py”, line 347, in kneighbors
(train_size, n_neighbors)
ValueError: Expected n_neighbors <= n_samples, but n_samples = 5, n_neighbors = 6
It looks like your k is larger than the number of instances in one class. You need to reduce k or increase the number of instances for the least represented class.
I hope that helps.
i cleared the error sir
but when i work with Gradientboosting it doesn’t work even though my dataset contains 2 classes as shown in the above discussion.
error : binomial deviance require 2 classes
and code :
import pandas
import matplotlib.pyplot as plt
from sklearn import model_selection
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import accuracy_score
data = (‘dataset160.csv’)
dataframe = pandas.read_csv(data)
array = dataframe.values
X = array[:,0:12]
Y = array[:,12]
print (X, Y)
plt.show()
seed = 7
num_trees = 100
kfold = model_selection.KFold(n_splits=10, random_state=seed)
model = GradientBoostingClassifier(n_estimators=num_trees, random_state=seed)
results = model_selection.cross_val_score(model, X, Y, cv=kfold)
print(results.mean())
print(results)
Perhaps you need to transform your class variable from numeric to being a label.
sklearn has a label encoder you can use:
http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html
sir, i have already labelled data set ,
i am unable to run the gradient boosting code on my dataset.
please help me.
see my parameters.
AGE Haemoglobin RBC Hct Mcv Mch Mchc Platelets WBC Granuls Lymphocytes Monocytes disese
3 9.6 4.2 28.2 67 22.7 33.9 3.75 5800 44 50 6 Positive
11 12.1 4.3 33.7 78 28.2 36 2.22 6100 73 23 4 Positive
2 9.5 4.1 27.9 67 22.8 34 3.64 5100 64 32 4 Positive
4 9.9 3.9 27.8 71 25.3 35.6 2.06 4900 65 32 3 Positive
14 10.7 4.4 31.2 70 24.2 34.4 3 7600 50 44 6 Negative
7 9.8 4.2 28 66 23.2 35.1 1.95 3800 28 63 9 Negative
8 14.6 5 39.2 77 28.7 37.2 3.06 4400 58 36 6 Negative
4 12 4.5 33.3 74 26.5 35.9 5.28 9500 40 54 6 Negative
2 11.2 4.6 32.7 70 24.1 34.3 2.98 8800 38 58 4 Negative
1 9.1 4 27.2 67 22.4 33.3 3.6 5300 40 55 5 Negative
11 14.8 5.8 42.5 72 25.1 34.8 4.51 17200 75 20 5 Negative
Perhaps this tutorial will help you get started:
https://machinelearningmastery.com/machine-learning-in-python-step-by-step/
thank you sir.
sir i have a small doubt.
is that XGBoost algorithm is best or SMOTEBoost algorithm is best to handle skewed data.
Try both on your specific dataset and see what works best.
thank you so much sir,
finally i have a doubt sir.
when i run the prediction code using adaboost i am getting 0.0%prediction accuracy .
for the code :
model = AdaBoostClassifier(n_estimators=num_trees, random_state=seed)
results = model_selection.cross_val_score(model, X, Y, cv=kfold)
model.fit(X, Y)
print(‘learning accuracy’)
print(results.mean())
predictions = model.predict(A)
print(predictions)
accuracy1 = accuracy_score(B, predictions)
print(“Accuracy % is “)
print(accuracy1*100)
anything wrong in the code . always i am getting 0.0.%accuracy and precision recall also 0% for any ensemble like boosting , bagging.
kindly rectify sir.
It’s not clear. Perhaps try a suite of algorithms and see what works best on your problem. I recommend this process:
https://machinelearningmastery.com/machine-learning-performance-improvement-cheat-sheet/
Dr. Jason you ARE doing a great job in machine learning.
Thanks Amos, I really appreciate your support!
Hi Jason,
Thanks a lot for the great article. You are an inspiration.
Is there a way I could measure the performance impact of the different ensemble methods?
For me, the VotingClassifier took more time than the others. If there is a metric could you please help identify which is faster and has the least performance implications when working with larger datasets?
Yes, I would recommend a robust test harness such as repeated cross validation, see here:
https://machinelearningmastery.com/evaluate-skill-deep-learning-models/
will you please show how to use CostSensitiveRandomForestClassifier()
on a dataset. i can’t run the code sir.
Thanks for the suggestion.
r u have any sample code ….. on costsensitive ensemble method
Not at this stage, thanks for the suggestion.
Hi Jason,
I have following queries :-
1. As bagging method only works for high variance so don’t you think that while using bagging we actually reducing overfitting as it occurs when we have low bias and high variance in our model?
2. Since random forest is used to lower the correlation between individual classifiers as we have in bagging approach. So, is this also leads to reduce the overfitting in our model by reducing correlation?
3. Can you please refer me a way to study thoroughly about stack aggregation?
Thanks in advance.
Bagging can reduce overfitting.
RF often performs better than bagging.
Here’s a post on stacking:
https://machinelearningmastery.com/implementing-stacking-scratch-python/
Thanks a lot Jason! great content as always!
I have a couple of questions:
1) Does more advanced methods that learn how to best weight the predictions from submodels (i.e Stacking) always give better results than simpler ensembling techniques?
2) I read in your post on stacking that it works better if the predictions of submodels are weakly correlated. Does it mean that it is better to train submodels from different families? (for example: a SVM model, a RF and a neural net)
Can I build an Aggregated model using stacking with Xgboost, LigthGBM, GBM?
Thanks!
No, ensembles are not always better, but if used carefully often are.
Yes, different families of models, different input features, etc.
Sure, you can try anything, just ensure you have a robust test harness.
Jason, thanks for your answer. I have two more questions:
1) What kind of test can I use in order to ensure the robustness of my ensembled model?
2) How do you deal with imbalanced classes in this context?
Thanks in advance.
Repeated cross validation is a good approach to evaluating model skill:
https://machinelearningmastery.com/evaluate-skill-deep-learning-models/
I have some ideas on working with imbalanced data here:
https://machinelearningmastery.com/tactics-to-combat-imbalanced-classes-in-your-machine-learning-dataset/
Hey Jason,
I have a question with regards to a specific hyperparameter the “base_estimator” of AdaBoostClassifier. Is there a way for me to ensemble several models (For instance: DecisionTreeClassifier, KNeighborsClassifier, and SVC) into the base_estimator hyperparameter? Cause I have seen most people implementing only one model but the main concept of AdaBoostClassifiers is to train different classifiers into an ensemble giving more weigh to incorrect classifications and correct prediction models through the use of bagging. Basically, I just want to know if this is possible to add several classifiers into the base_estimator hyperparameter. Thanks for the help and nice post!
Perhaps, but I don’t think so. I’d recommend stacking or voting instead.
Dear Jason,
I am using a simple backpropagation NN with time delays for time series forecasting. My data is heavily skewed with only a few extreme values. When I run e.g. 20 identical models (not ensembles), with random weights for each run, I choose the model with lowest validation error. But the problem then is that the error using the test set for that model may not be the lowest.
So I suppose ensembles might help, but what is the best approach for NN? Perhaps you have already answered this somewhere. I suppose I can e.g. just take the average or median or some other measures for my 20 models, but will this count as ensembles?
Thanks for sharing your knowledge!
You may need a more robust way of selecting models that better captures the skill of the model on out of sample data.
I have some ideas here:
https://machinelearningmastery.com/evaluate-skill-deep-learning-models/
I just tried the classifier on iris dataset its giving accuracy as 1.00.I dont think its classifying properly.
Hope u can help me
Perhaps try running the example a few times. The algorithms are stochastic and by chance it might have achieved 100% accuracy.
Thank you Jason for your helpful tutorial.
I’ve a question about Voting ensembles, I mean what is the difference between average voting and majrity voting (I know how it works), but I want to know in which situation we apply majority voting and the same thing about average voting.
Another question: By applying majority voting, is it obliged to train classifiers on the same training set? I’am working on enhancing prediction’s accuracy of models by updating the training dataset in each iteration (by selecting relevant feautures).
Averaging is for regression problems, majority (statistical mode) is for classification.
Ultimately, try both and see what works best for your specific problem and models.
HI Jason,
If we have both a classification and regression problem that rely on the same input data, is it possible to successfully architect a neural network that gives both classification and regression outputs?
If so, how might the combined output loss and accuracy function be constructed? Do we have to sum up both loss and accuracy function?
Thank you Jason.
Yes, see this post:
https://machinelearningmastery.com/keras-functional-api-deep-learning/
Thank you, Jason! I will definitely look it through.
hello sir, thanks for the great post. I would like to know, after building the ensemble classifier, how do i test it with a new test data? what is the procedure?
See this post:
https://machinelearningmastery.com/train-final-machine-learning-model/
Hello, Jason. Thanks you are doing a great work, I am working on my Master research project in which I am using Random Forest with Sklearn but have to cite this paper “1. Breiman, L., “Random Forests”, Machine Learning. Vol. 45, No. 1, pp. 5–32, 2001.” i.e. base paper of Random Forest and he used Voting method but in sklearn documentation they given “In contrast to the original publication [B2001], the scikit-learn implementation combines classifiers by averaging their probabilistic prediction, instead of letting each classifier vote for a single class.” and I implemented RandomForestClassifier() in my program and works very well.
Now, my question is as I have to write some details of Random Forest in my research paper and will explain about voting method too so, should I use your above “Voting Ensemble” method or simple sklearn implementaiton is fine.?
This really depends on your goals.
If you use the sklearn method you must document how it works.
Hi Jason,
how can I use ensemble machine learning algorithm for regression problem?
For example: how we can ensemble tow regression models such as SVR and linear regression to improve the regression result?
The same as for classification, just with a different output. Most ensemble algorithms work for regression and classification (e.g. random forests, bagging, stacking, voting, etc.)
Is there a specific problem you’re having?
I’m trying to use the GradientBoostingRegressor function to combine the predictions of two machine learning algorithms ( linear regression and SVR algorithms) to predict the popularity of the image. As I know In case of regression it takes an average of results instead of voting. I wrote the following code :
# coding: utf-8
import numpy
import pandas
from keras.models import Sequential
from keras.layers import Dense
from keras.wrappers.scikit_learn import KerasRegressor
from sklearn.model_selection import cross_val_score,cross_val_predict
from sklearn.model_selection import KFold
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn import model_selection
from sklearn import metrics
import matplotlib
#from matplotlib import pyplot as PLT
matplotlib.use(‘Agg’)
import matplotlib.pyplot
import matplotlib.pyplot as plt
import time
import cPickle
#from keras.utils.visualize_util import plot
import os
import theano
from PIL import Image
from numpy import *
import scipy
import numpy as np
from sklearn.metrics import mean_squared_error
from sklearn.datasets import make_friedman1
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.linear_model import LinearRegression
from sklearn.svm import SVR
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# load dataset
dataframe = pandas.read_csv(‘/home/fatmasaid/regression_code/user_features.csv’, delim_whitespace=True, header=None)
dataset = dataframe.values
# split into input (X) and output (Y) variables
X = dataset[:,0:5]
Y = dataset[:,5]
seed = 7
numpy.random.seed(seed)
kfold = model_selection.KFold(n_splits=10, random_state=seed)
# Initializing models
lr = LinearRegression()
svr_lin = SVR(kernel=’linear’)
model1 = GradientBoostingRegressor( lr ,n_estimators=100, learning_rate=0.1, max_depth=1, random_state=seed, loss=’ls’)
result1 = model_selection.cross_val_score(model1, X, Y, cv=kfold)
print(result1.mean())
model2 = GradientBoostingRegressor( svr_lin ,n_estimators=100, learning_rate=0.1, max_depth=1, random_state=seed, loss=’ls’)
result2 = model_selection.cross_val_score(model2, X, Y, cv=kfold)
print(result2.mean())
# Make cross validated predictions & compute Sperman
p1 = cross_val_predict(model1, X, Y, cv=kfold)
plt.scatter(Y, p1)
print scipy.stats.stats.spearmanr(Y, p1)[0]
p2 = cross_val_predict(model2, X, Y, cv=kfold)
plt.scatter(Y, p2)
print scipy.stats.stats.spearmanr(Y, p2)[0]
p = np.mean([p1, p2], axis=0)
mse = mean_squared_error(Y, p)
print(“MSE: %.4f” % mse)
I got the following error:
TypeError: __init__() got multiple values for keyword argument ‘loss’
I don’t if my idea is right or not. So, please if you have any example then you can upload it.
Thanks
I’m eager to help, but I cannot debug your code for you.
Perhaps post it to stackoverflow?
Hello Jason,
I have been posted the code to stackoverflow. you can find on the following link:
https://stackoverflow.com/questions/49792812/gradient-boosting-regression-algorithm
Thanks
Thanks for your great post
Do you have any post for ensemble classifier while Multi-Label?
I don’t have any examples of multi-label classification, sorry.
Hi Jason,
This is a fantastic post! Thank you for posting it.
You use
kfold = model_selection.KFold(n_splits=10, random_state=seed)
to generate the value for cv in your cross_val_score calculations.
Is this really necessary for regression estimators, as cross_val_score and cross_val_predict already use KFold by default for regression and other cases. Is there an advantage to your implementation of KFold?
Cheers!
Thanks. You can use either way. This approach offers more control/insight into what is going on.
Jason,
Also, have you used VotingClassifier to combine regression estimators? So far it looks like it only excepts integer model predictions, not continuous.
Thank you…
No, it is for classifiers only.
Thank you for both answers, Jason! Keep up the good work.
Dear Jason,
Thank you very much for your blogs they are great.
I have the following task and do not know how to accomplish it:
1. I have legacy code which is not well-done looks like this:
clf = BaggingRegressor(svm.SVR(C=10.0), n_estimators=64, max_samples=0.9, max_features=0.8)
predicted = cross_val_predict(clf, X_standard, y_standard.ravel(), cv=10, n_jobs=10)
predicted = y_scaler.inverse_transform(predicted)
gMAE, gMRE = evaluate(j, predicted, y[i][j])
In the code as you can see the person has done cross_val_predict to train and predict svr model.
My task is using the same data but dnn models to predict and prove that my dnn models are better.
How should I do that since I think initially this project has not been done well.
I recommend developing a suite of different models in order to discover what works best for your specific dataset. Perhaps start with an MLP.
I have the MLP-models (done in TF). However, I do not know how to compare them because in my TF models I do not use CrossValidation and in order to compare the results, I need to use the same training and validation sets, which from this function before looks like are created randomly. I do not know if you understand better my question now.
You can evaluate models using the same train/test sets.
It’s possible to use decisiontree + adapboost or it’s only for bagging?
Yes.
sir, instead of directly using extratreeclassifier, i want to call it as user defined bulit in function, but it won’t works. can u please suggest me how to write or use extratreeclassfier as user own defined function
Sorry, I don’t have the capacity to review your code.
Hello Jason,
First of all thank you for these awesome tutorials. I have used the pima indians diabetes dataset and applied modeling using MLP neural networks, and got an accuracy of around 73%. Now I want to boost my accuracy using ensembles, so shall I discard MLP and depend only on either Trees, Random Forests, etc. ?
Thanks in advance.
It is a good idea to test a suite of algorithms for a given dataset in order to discover what works best.
Hello Jason, thank you for these aesome tutorials.
May I ask you that after we did the ensembles and got better accuracy, how could we get again this accuracy in the initial models we used before doing ensembles ? (Sorry if my question seems dumb I’m still a beginner)
Thanks in adavance
The idea is that the ensemble offers better performance than a single model. You use the ensemble to make predictions.
How can i use more than one base estimator to bagging in scikit learn python? Doing great work by the way.
Perhaps use two separate bagging models and combine their predictions using voting?
I would like to ensemble multiple binary class models in a way that if at least one model gives class 1 then summary model also gives 1. How I can approach that? Thank you.
Yes, you could manage a voting ensemble manually and use if-statements to check the predictions of the submodels.
Hi!
I need to add a random forest classifier after a simple RNN, How to do this?
thanks.
Perhaps collect the predictions from the RNN and then feed them into a random forest?
e.g. write some code to do it, rather than connect the models directly.
Hi Jason, as always this article has kindled my interest in getting to know more on Machine Learning.
While implementing voting classifier why does the score change for every run?
Good question, this is a common question, I answer it here:
https://machinelearningmastery.com/faq/single-faq/why-do-i-get-different-results-each-time-i-run-the-code
I have the Following error while applying SMOTE
ValueError Traceback (most recent call last)
in
1 sm = SMOTE(random_state=2)
—-> 2 X_train_res, y_train_res = sm.fit_sample(X,y)
~\Anaconda3\lib\site-packages\imblearn\base.py in fit_resample(self, X, y)
83 self.sampling_strategy, y, self._sampling_type)
84
—> 85 output = self._fit_resample(X, y)
86
87 if binarize_y:
~\Anaconda3\lib\site-packages\imblearn\over_sampling\_smote.py in _fit_resample(self, X, y)
794 def _fit_resample(self, X, y):
795 self._validate_estimator()
–> 796 return self._sample(X, y)
797
798 def _sample(self, X, y):
~\Anaconda3\lib\site-packages\imblearn\over_sampling\_smote.py in _sample(self, X, y)
810
811 self.nn_k_.fit(X_class)
–> 812 nns = self.nn_k_.kneighbors(X_class, return_distance=False)[:, 1:]
813 X_new, y_new = self._make_samples(X_class, y.dtype, class_sample,
814 X_class, nns, n_samples, 1.0)
~\Anaconda3\lib\site-packages\sklearn\neighbors\base.py in kneighbors(self, X, n_neighbors, return_distance)
414 “Expected n_neighbors 416 (train_size, n_neighbors)
417 )
418 n_samples, _ = X.shape
ValueError: Expected n_neighbors <= n_samples, but n_samples = 5, n_neighbors = 6
Sorry, I cannot debug your code for you.
how to solve this error
sampling_strategy” can be a float only when the type ‘
‘of target is binary. For multi-class, use a dict.
how can i convert my float object to Dict?
Sorry, i have not seen this error before.
Perhaps post your code and error to stackoverflow?
@Anurag singh
based on your problems, you may want to play around with this paramter within SMOTE function: k_neighbors to suit your situation (e.g. sm=SMOTE(k_neighbors=1)).
Hopefully I am not pointing you away from solving your problems.
Thanks for sharing.
Well written Jason,
I came across this article as am trying to implement a voting classifier,
I have three questions that I wish you have the time to answer:
I am sort of new to this so excuse me if any of my questions sounded silly.
_________________________________________________________________
===============================================================
Question#1- I am regarding the ensembler as a new classifier now with a higher score than the others. I tried to implement the voting classifier to have a (yhat) but it failed.
ensemble=VotingClassifier(estimators)
results = cross_val_score (ensemble, X, y , cv=5)
print (“The ensembler accuracy =”,results.mean())
yhat_ensemble=ensemble.predict(x_test)
The last step gave the following error:
((NotFittedError: This VotingClassifier instance is not fitted yet. Call ‘fit’ with appropriate arguments before using this method))
_________________________________________________________________
==============================================================
Question#2- is there any way to find the probabilities using the ensembler(with soft voting=True)?
i.e. using the following command:
yhat_prob_ensemble = ensemble.predict.proba(x_test)
This also gave me the same (NotFittedError) error as above.
_________________________________________________________________
==============================================================
Question#3 – is it normal to have a classifier with a higher cross-validation score than the ensembler?
I have a Logistic regression (score 0.8) Naive Bayesian (0.73), and Decision Tree (0.71) while the ensembler’s score is (0.74).
_________________________________________________________________
==============================================================
Dear Jason,
as I said earlier, Please execuse my silly questions, I just solved questions 1 and 2 by fitting the new ensembler again.. My previous understanding is that fitting was already done (with the original classifiers) thus we can not do it again. the missing line was:
ensemble = ensemble.fit(X_train, y_train)
However, Quesion#3 still stands. and I hope you find the time to answer it.
Thanks
Happy to hear that Max.
That is very strange, it looks like you are using the VotingClassifier the right way.
Perhaps try debugging – e.g. use a different model, use different ensemble, use a subset of models, etc. try to flush out the cause of the fault.
Typically you only want to adopt the ensemble if it performs better than any single model. Otherwise move on.
i have 6 subset of features sets , i run different machine learning techniques on them and get the results. how can i combine them in ensemble model using python
could you tell me how to do it
Perhaps using hard or soft voting.
hi jason thnx for your poet
i wonder in random forest why you did not fit the model
Sorry, I don’t understand. You can please elaborate your question?
i mean why u did’t write :
classifier.fit(X_train,y_train)
within your code
Because we are evaluating the models many time using cross validation. E.g. the fit is called as part of the cross validation process.
Does that help?
i try to run the random forest code.with num_trees = 50 only cause if i use 100 the program stop running
it gives me : 0.056247265097497987
i don’t know where is the mistake
Perhaps this will help:
https://machinelearningmastery.com/faq/single-faq/why-does-the-code-in-the-tutorial-not-work-for-me
Hi,
I am working on a machine learning project. My data is all about trading(open,high,cos,low) etc. I have constructed some techincal indicators based on those columns. My main goal is to predict the market phase (bullish,bearish,lateral). I have about 15k rows to train the model. I think this problem comes under classification. So I have been using all types of classsification algorithms but they result in 40-50% of accuracy. I want to increase them upto 70%. please let me know about how to increase the accuracy.
Test accuracy is arround 90% but when I use the model on real data it is giving arround 40%
See this:
https://machinelearningmastery.com/faq/single-faq/can-you-help-me-with-machine-learning-for-finance-or-the-stock-market
haha! Never mind! Thanks 🙂
No problem.
I Jason , I am thinking of applying bagging with LSTM , Can you provide me some idea or related links.
-Thnx in advance
Yes, see the tutorials on ensembles with deep learning here:
https://machinelearningmastery.com/start-here/#better
hi Jason , if i want to apply random subspace technique as a first layer then apply ensemble techniques . do you have any materiel in python to learn it
No, sorry.
Hi jason, i want to perform K-fold cross validation for ensemble of classifers with Dynamic Selection (DS) methods. Can you lease suggest me some idea or related links.
What are “dynamic selection methods”?
Hi Jason, Is there any way to plot all ensemble members as well as the final model? for example I want to plot all ensemble members of DescisonTreeRegression base model.
Perhaps, I don’t have an example sorry.
I’m not looking for example just wanted to know is it possible to access the result(model) of ensemble members separately after fitting (in sklearn)?
Perhaps review the API or prepare a prototype and discover the answer directly – in minutes.
Hi Json,
I got the following error while working with AdaBoost,
ValueError: Unknown label type: ‘continuous’
What might be the reason? Please help me
Thank You
It suggests the variable you are trying to predict is numerical rather than a class label.
Perhaps you need to prepare your data before modeling.
Hi Jason,
I found it, It was because the label assigned was a continues to value.
By the way, model (AdaBoost) accuracy by using K-Fold Cross-Validation and Train-Test split methods gave me different figures.
K-Fold Cross-Validation – ~90%
Train-Test split – Overfit 100% (test accuracy ~ 98%)
Any thoughts on this? Please help
Thank you.
Nuwan C
Yes, the train/test split is likely optimistic.
For small datasets, repeated k-fold cross-validation may give a more accurate estimate of model performance.
Thank you for the reply Json
Cheers!!!
You’re welcome!
Hi Jason,
I want to build an ensemble model for time-series forecasting problem, should I use the same techniques listed here ?
Many thanks for your informative website.
Perhaps try them and see if they lift performance on your dataset.
Respected Sir,
Thank you very much for this tutorial. I use your code for my dataset. It works well and gives 100% accuracy while implementing all classifiers. Is it a over fitting problem? Kindly clarify me. Thank you.
If you are getting 100% on a hold out dataset, you are not overfitting.
Also, if you are getting 100% accuracy on any problem, it’s probably too simple and does not require machine learning.
Thanks for the great explanation.
One question: If I have different data with the same length and label and use various classifiers for each data and finally want to fuse their result can I use a similar way? If yes how, do you have a documents for it?
Sorry, I don’t understand your question. Can you please elaborate or rephrase it?
Which base estimators can be used with Bagging and boosting in sklearn?
You can bag any model I believe. Boosting might only be for trees.
Hi Jason
I want to do a fusion of two mlps and two cnns. But I am being unable to do so. Please help.
Perhaps start by averaging their predictions together.
Hi Jason, could you please tell me how does sklearn’s bagging classifier calculate the final prediction score and what kind of voting method does it use?
Good question see this:
https://machinelearningmastery.com/bagging-ensemble-with-python/
Hi Jason, Thank you for the great tutorial!
i want to apply a fusion classifier between BERT, Elmo, and ULMFit language models,
can I do it in the same way that you applied?
and is the fusion classifier the same ensemble classifier and can use votingclassifier() or different?
Perhaps try it and see.