Classification is a predictive modeling problem that involves assigning a label to a given input data sample.
The problem of classification predictive modeling can be framed as calculating the conditional probability of a class label given a data sample. Bayes Theorem provides a principled way for calculating this conditional probability, although in practice requires an enormous number of samples (very large-sized dataset) and is computationally expensive.
Instead, the calculation of Bayes Theorem can be simplified by making some assumptions, such as each input variable is independent of all other input variables. Although a dramatic and unrealistic assumption, this has the effect of making the calculations of the conditional probability tractable and results in an effective classification model referred to as Naive Bayes.
In this tutorial, you will discover the Naive Bayes algorithm for classification predictive modeling.
After completing this tutorial, you will know:
- How to frame classification predictive modeling as a conditional probability model.
- How to use Bayes Theorem to solve the conditional probability model of classification.
- How to implement simplified Bayes Theorem for classification, called the Naive Bayes algorithm.
Kick-start your project with my new book Probability for Machine Learning, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.
- Updated Oct/2019: Fixed minor inconsistency issue in math notation.
- Update Jan/2020: Updated for changes in scikit-learn v0.22 API.

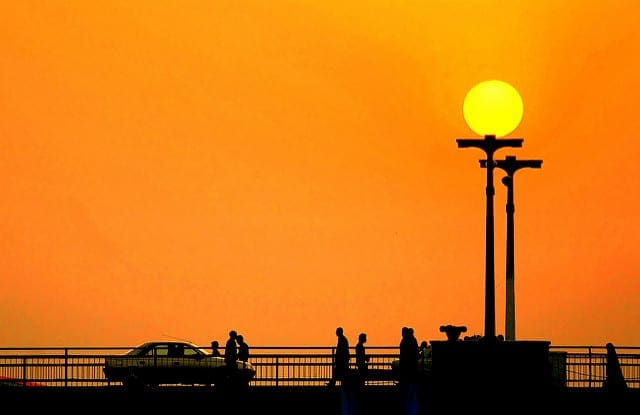
How to Develop a Naive Bayes Classifier from Scratch in Python
Photo by Ryan Dickey, some rights reserved.
Tutorial Overview
This tutorial is divided into five parts; they are:
- Conditional Probability Model of Classification
- Simplified or Naive Bayes
- How to Calculate the Prior and Conditional Probabilities
- Worked Example of Naive Bayes
- 5 Tips When Using Naive Bayes
Conditional Probability Model of Classification
In machine learning, we are often interested in a predictive modeling problem where we want to predict a class label for a given observation. For example, classifying the species of plant based on measurements of the flower.
Problems of this type are referred to as classification predictive modeling problems, as opposed to regression problems that involve predicting a numerical value. The observation or input to the model is referred to as X and the class label or output of the model is referred to as y.
Together, X and y represent observations collected from the domain, i.e. a table or matrix (columns and rows or features and samples) of training data used to fit a model. The model must learn how to map specific examples to class labels or y = f(X) that minimized the error of misclassification.
One approach to solving this problem is to develop a probabilistic model. From a probabilistic perspective, we are interested in estimating the conditional probability of the class label, given the observation.
For example, a classification problem may have k class labels y1, y2, …, yk and n input variables, X1, X2, …, Xn. We can calculate the conditional probability for a class label with a given instance or set of input values for each column x1, x2, …, xn as follows:
- P(yi | x1, x2, …, xn)
The conditional probability can then be calculated for each class label in the problem and the label with the highest probability can be returned as the most likely classification.
The conditional probability can be calculated using the joint probability, although it would be intractable. Bayes Theorem provides a principled way for calculating the conditional probability.
The simple form of the calculation for Bayes Theorem is as follows:
- P(A|B) = P(B|A) * P(A) / P(B)
Where the probability that we are interested in calculating P(A|B) is called the posterior probability and the marginal probability of the event P(A) is called the prior.
We can frame classification as a conditional classification problem with Bayes Theorem as follows:
- P(yi | x1, x2, …, xn) = P(x1, x2, …, xn | yi) * P(yi) / P(x1, x2, …, xn)
The prior P(yi) is easy to estimate from a dataset, but the conditional probability of the observation based on the class P(x1, x2, …, xn | yi) is not feasible unless the number of examples is extraordinarily large, e.g. large enough to effectively estimate the probability distribution for all different possible combinations of values.
As such, the direct application of Bayes Theorem also becomes intractable, especially as the number of variables or features (n) increases.
Want to Learn Probability for Machine Learning
Take my free 7-day email crash course now (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
Simplified or Naive Bayes
The solution to using Bayes Theorem for a conditional probability classification model is to simplify the calculation.
The Bayes Theorem assumes that each input variable is dependent upon all other variables. This is a cause of complexity in the calculation. We can remove this assumption and consider each input variable as being independent from each other.
This changes the model from a dependent conditional probability model to an independent conditional probability model and dramatically simplifies the calculation.
First, the denominator is removed from the calculation P(x1, x2, …, xn) as it is a constant used in calculating the conditional probability of each class for a given instance and has the effect of normalizing the result.
- P(yi | x1, x2, …, xn) = P(x1, x2, …, xn | yi) * P(yi)
Next, the conditional probability of all variables given the class label is changed into separate conditional probabilities of each variable value given the class label. These independent conditional variables are then multiplied together. For example:
- P(yi | x1, x2, …, xn) = P(x1|yi) * P(x2|yi) * … P(xn|yi) * P(yi)
This calculation can be performed for each of the class labels, and the label with the largest probability can be selected as the classification for the given instance. This decision rule is referred to as the maximum a posteriori, or MAP, decision rule.
This simplification of Bayes Theorem is common and widely used for classification predictive modeling problems and is generally referred to as Naive Bayes.
The word “naive” is French and typically has a diaeresis (umlaut) over the “i”, which is commonly left out for simplicity, and “Bayes” is capitalized as it is named for Reverend Thomas Bayes.
How to Calculate the Prior and Conditional Probabilities
Now that we know what Naive Bayes is, we can take a closer look at how to calculate the elements of the equation.
The calculation of the prior P(yi) is straightforward. It can be estimated by dividing the frequency of observations in the training dataset that have the class label by the total number of examples (rows) in the training dataset. For example:
- P(yi) = examples with yi / total examples
The conditional probability for a feature value given the class label can also be estimated from the data. Specifically, those data examples that belong to a given class, and one data distribution per variable. This means that if there are K classes and n variables, that k * n different probability distributions must be created and maintained.
A different approach is required depending on the data type of each feature. Specifically, the data is used to estimate the parameters of one of three standard probability distributions.
In the case of categorical variables, such as counts or labels, a multinomial distribution can be used. If the variables are binary, such as yes/no or true/false, a binomial distribution can be used. If a variable is numerical, such as a measurement, often a Gaussian distribution is used.
- Binary: Binomial distribution.
- Categorical: Multinomial distribution.
- Numeric: Gaussian distribution.
These three distributions are so common that the Naive Bayes implementation is often named after the distribution. For example:
- Binomial Naive Bayes: Naive Bayes that uses a binomial distribution.
- Multinomial Naive Bayes: Naive Bayes that uses a multinomial distribution.
- Gaussian Naive Bayes: Naive Bayes that uses a Gaussian distribution.
A dataset with mixed data types for the input variables may require the selection of different types of data distributions for each variable.
Using one of the three common distributions is not mandatory; for example, if a real-valued variable is known to have a different specific distribution, such as exponential, then that specific distribution may be used instead. If a real-valued variable does not have a well-defined distribution, such as bimodal or multimodal, then a kernel density estimator can be used to estimate the probability distribution instead.
The Naive Bayes algorithm has proven effective and therefore is popular for text classification tasks. The words in a document may be encoded as binary (word present), count (word occurrence), or frequency (tf/idf) input vectors and binary, multinomial, or Gaussian probability distributions used respectively.
Worked Example of Naive Bayes
In this section, we will make the Naive Bayes calculation concrete with a small example on a machine learning dataset.
We can generate a small contrived binary (2 class) classification problem using the make_blobs() function from the scikit-learn API.
The example below generates 100 examples with two numerical input variables, each assigned one of two classes.
|
1 2 3 4 5 6 7 8 |
# example of generating a small classification dataset from sklearn.datasets import make_blobs # generate 2d classification dataset X, y = make_blobs(n_samples=100, centers=2, n_features=2, random_state=1) # summarize print(X.shape, y.shape) print(X[:5]) print(y[:5]) |
Running the example generates the dataset and summarizes the size, confirming the dataset was generated as expected.
The “random_state” argument is set to 1, ensuring that the same random sample of observations is generated each time the code is run.
The input and output elements of the first five examples are also printed, showing that indeed, the two input variables are numeric and the class labels are either 0 or 1 for each example.
|
1 2 3 4 5 6 7 |
(100, 2) (100,) [[-10.6105446 4.11045368] [ 9.05798365 0.99701708] [ 8.705727 1.36332954] [ -8.29324753 2.35371596] [ 6.5954554 2.4247682 ]] [0 1 1 0 1] |
We will model the numerical input variables using a Gaussian probability distribution.
This can be achieved using the norm SciPy API. First, the distribution can be constructed by specifying the parameters of the distribution, e.g. the mean and standard deviation, then the probability density function can be sampled for specific values using the norm.pdf() function.
We can estimate the parameters of the distribution from the dataset using the mean() and std() NumPy functions.
The fit_distribution() function below takes a sample of data for one variable and fits a data distribution.
|
1 2 3 4 5 6 7 8 9 |
# fit a probability distribution to a univariate data sample def fit_distribution(data): # estimate parameters mu = mean(data) sigma = std(data) print(mu, sigma) # fit distribution dist = norm(mu, sigma) return dist |
Recall that we are interested in the conditional probability of each input variable. This means we need one distribution for each of the input variables, and one set of distributions for each of the class labels, or four distributions in total.
First, we must split the data into groups of samples for each of the class labels.
|
1 2 3 4 5 |
... # sort data into classes Xy0 = X[y == 0] Xy1 = X[y == 1] print(Xy0.shape, Xy1.shape) |
We can then use these groups to calculate the prior probabilities for a data sample belonging to each group.
This will be 50% exactly given that we have created the same number of examples in each of the two classes; nevertheless, we will calculate these priors for completeness.
|
1 2 3 4 5 |
... # calculate priors priory0 = len(Xy0) / len(X) priory1 = len(Xy1) / len(X) print(priory0, priory1) |
Finally, we can call the fit_distribution() function that we defined to prepare a probability distribution for each variable, for each class label.
|
1 2 3 4 5 6 7 |
... # create PDFs for y==0 X1y0 = fit_distribution(Xy0[:, 0]) X2y0 = fit_distribution(Xy0[:, 1]) # create PDFs for y==1 X1y1 = fit_distribution(Xy1[:, 0]) X2y1 = fit_distribution(Xy1[:, 1]) |
Tying this all together, the complete probabilistic model of the dataset is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 |
# summarize probability distributions of the dataset from sklearn.datasets import make_blobs from scipy.stats import norm from numpy import mean from numpy import std # fit a probability distribution to a univariate data sample def fit_distribution(data): # estimate parameters mu = mean(data) sigma = std(data) print(mu, sigma) # fit distribution dist = norm(mu, sigma) return dist # generate 2d classification dataset X, y = make_blobs(n_samples=100, centers=2, n_features=2, random_state=1) # sort data into classes Xy0 = X[y == 0] Xy1 = X[y == 1] print(Xy0.shape, Xy1.shape) # calculate priors priory0 = len(Xy0) / len(X) priory1 = len(Xy1) / len(X) print(priory0, priory1) # create PDFs for y==0 X1y0 = fit_distribution(Xy0[:, 0]) X2y0 = fit_distribution(Xy0[:, 1]) # create PDFs for y==1 X1y1 = fit_distribution(Xy1[:, 0]) X2y1 = fit_distribution(Xy1[:, 1]) |
Running the example first splits the dataset into two groups for the two class labels and confirms the size of each group is even and the priors are 50%.
Probability distributions are then prepared for each variable for each class label and the mean and standard deviation parameters of each distribution are reported, confirming that the distributions differ.
|
1 2 3 4 5 6 |
(50, 2) (50, 2) 0.5 0.5 -1.5632888906409914 0.787444265443213 4.426680361487157 0.958296071258367 -9.681177100524485 0.8943078901048118 -3.9713794295185845 0.9308177595208521 |
Next, we can use the prepared probabilistic model to make a prediction.
The independent conditional probability for each class label can be calculated using the prior for the class (50%) and the conditional probability of the value for each variable.
The probability() function below performs this calculation for one input example (array of two values) given the prior and conditional probability distribution for each variable. The value returned is a score rather than a probability as the quantity is not normalized, a simplification often performed when implementing naive bayes.
|
1 2 3 |
# calculate the independent conditional probability def probability(X, prior, dist1, dist2): return prior * dist1.pdf(X[0]) * dist2.pdf(X[1]) |
We can use this function to calculate the probability for an example belonging to each class.
First, we can select an example to be classified; in this case, the first example in the dataset.
|
1 2 3 |
... # classify one example Xsample, ysample = X[0], y[0] |
Next, we can calculate the score of the example belonging to the first class, then the second class, then report the results.
|
1 2 3 4 5 |
... py0 = probability(Xsample, priory0, distX1y0, distX2y0) py1 = probability(Xsample, priory1, distX1y1, distX2y1) print('P(y=0 | %s) = %.3f' % (Xsample, py0*100)) print('P(y=1 | %s) = %.3f' % (Xsample, py1*100)) |
The class with the largest score will be the resulting classification.
Tying this together, the complete example of fitting the Naive Bayes model and using it to make one prediction is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 |
# example of preparing and making a prediction with a naive bayes model from sklearn.datasets import make_blobs from scipy.stats import norm from numpy import mean from numpy import std # fit a probability distribution to a univariate data sample def fit_distribution(data): # estimate parameters mu = mean(data) sigma = std(data) print(mu, sigma) # fit distribution dist = norm(mu, sigma) return dist # calculate the independent conditional probability def probability(X, prior, dist1, dist2): return prior * dist1.pdf(X[0]) * dist2.pdf(X[1]) # generate 2d classification dataset X, y = make_blobs(n_samples=100, centers=2, n_features=2, random_state=1) # sort data into classes Xy0 = X[y == 0] Xy1 = X[y == 1] # calculate priors priory0 = len(Xy0) / len(X) priory1 = len(Xy1) / len(X) # create PDFs for y==0 distX1y0 = fit_distribution(Xy0[:, 0]) distX2y0 = fit_distribution(Xy0[:, 1]) # create PDFs for y==1 distX1y1 = fit_distribution(Xy1[:, 0]) distX2y1 = fit_distribution(Xy1[:, 1]) # classify one example Xsample, ysample = X[0], y[0] py0 = probability(Xsample, priory0, distX1y0, distX2y0) py1 = probability(Xsample, priory1, distX1y1, distX2y1) print('P(y=0 | %s) = %.3f' % (Xsample, py0*100)) print('P(y=1 | %s) = %.3f' % (Xsample, py1*100)) print('Truth: y=%d' % ysample) |
Running the example first prepares the prior and conditional probabilities as before, then uses them to make a prediction for one example.
The score of the example belonging to y=0 is about 0.3 (recall this is an unnormalized probability), whereas the score of the example belonging to y=1 is 0.0. Therefore, we would classify the example as belonging to y=0.
In this case, the true or actual outcome is known, y=0, which matches the prediction by our Naive Bayes model.
|
1 2 3 |
P(y=0 | [-0.79415228 2.10495117]) = 0.348 P(y=1 | [-0.79415228 2.10495117]) = 0.000 Truth: y=0 |
In practice, it is a good idea to use optimized implementations of the Naive Bayes algorithm. The scikit-learn library provides three implementations, one for each of the three main probability distributions; for example, BernoulliNB, MultinomialNB, and GaussianNB for binomial, multinomial and Gaussian distributed input variables respectively.
To use a scikit-learn Naive Bayes model, first the model is defined, then it is fit on the training dataset. Once fit, probabilities can be predicted via the predict_proba() function and class labels can be predicted directly via the predict() function.
The complete example of fitting a Gaussian Naive Bayes model (GaussianNB) to the same test dataset is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 |
# example of gaussian naive bayes from sklearn.datasets import make_blobs from sklearn.naive_bayes import GaussianNB # generate 2d classification dataset X, y = make_blobs(n_samples=100, centers=2, n_features=2, random_state=1) # define the model model = GaussianNB() # fit the model model.fit(X, y) # select a single sample Xsample, ysample = [X[0]], y[0] # make a probabilistic prediction yhat_prob = model.predict_proba(Xsample) print('Predicted Probabilities: ', yhat_prob) # make a classification prediction yhat_class = model.predict(Xsample) print('Predicted Class: ', yhat_class) print('Truth: y=%d' % ysample) |
Running the example fits the model on the training dataset, then makes predictions for the same first example that we used in the prior example.
In this case, the probability of the example belonging to y=0 is 1.0 or a certainty. The probability of y=1 is a very small value close to 0.0.
Finally, the class label is predicted directly, again matching the ground truth for the example.
|
1 2 3 |
Predicted Probabilities: [[1.00000000e+00 5.52387327e-30]] Predicted Class: [0] Truth: y=0 |
5 Tips When Using Naive Bayes
This section lists some practical tips when working with Naive Bayes models.
1. Use a KDE for Complex Distributions
If the probability distribution for a variable is complex or unknown, it can be a good idea to use a kernel density estimator or KDE to approximate the distribution directly from the data samples.
A good example would be the Gaussian KDE.
2. Decreased Performance With Increasing Variable Dependence
By definition, Naive Bayes assumes the input variables are independent of each other.
This works well most of the time, even when some or most of the variables are in fact dependent. Nevertheless, the performance of the algorithm degrades the more dependent the input variables happen to be.
3. Avoid Numerical Underflow with Log
The calculation of the independent conditional probability for one example for one class label involves multiplying many probabilities together, one for the class and one for each input variable. As such, the multiplication of many small numbers together can become numerically unstable, especially as the number of input variables increases.
To overcome this problem, it is common to change the calculation from the product of probabilities to the sum of log probabilities. For example:
- P(yi | x1, x2, …, xn) = log(P(x1|y1)) + log(P(x2|y1)) + … log(P(xn|y1)) + log(P(yi))
Calculating the natural logarithm of probabilities has the effect of creating larger (negative) numbers and adding the numbers together will mean that larger probabilities will be closer to zero. The resulting values can still be compared and maximized to give the most likely class label.
This is often called the log-trick when multiplying probabilities.
4. Update Probability Distributions
As new data becomes available, it can be relatively straightforward to use this new data with the old data to update the estimates of the parameters for each variable’s probability distribution.
This allows the model to easily make use of new data or the changing distributions of data over time.
5. Use as a Generative Model
The probability distributions will summarize the conditional probability of each input variable value for each class label.
These probability distributions can be useful more generally beyond use in a classification model.
For example, the prepared probability distributions can be randomly sampled in order to create new plausible data instances. The conditional independence assumption assumed may mean that the examples are more or less plausible based on how much actual interdependence exists between the input variables in the dataset.
Further Reading
This section provides more resources on the topic if you are looking to go deeper.
Tutorials
- Naive Bayes Tutorial for Machine Learning
- Naive Bayes for Machine Learning
- Better Naive Bayes: 12 Tips To Get The Most From The Naive Bayes Algorithm
Books
- Machine Learning, 1997.
- Machine Learning: A Probabilistic Perspective, 2012.
- Pattern Recognition and Machine Learning, 2006.
- Data Mining: Practical Machine Learning Tools and Techniques, 4th edition, 2016.
API
- sklearn.datasets.make_blobs API.
- scipy.stats.norm API.
- Naive Bayes, scikit-learn documentation.
- sklearn.naive_bayes.GaussianNB API
Articles
- Bayes’ theorem, Wikipedia.
- Naive Bayes classifier, Wikipedia.
- Maximum a posteriori estimation, Wikipedia.
Summary
In this tutorial, you discovered the Naive Bayes algorithm for classification predictive modeling.
Specifically, you learned:
- How to frame classification predictive modeling as a conditional probability model.
- How to use Bayes Theorem to solve the conditional probability model of classification.
- How to implement simplified Bayes Theorem for classification called the Naive Bayes algorithm.
Do you have any questions?
Ask your questions in the comments below and I will do my best to answer.


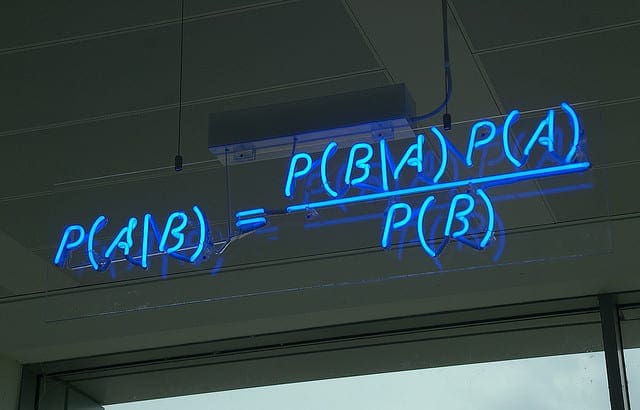




Dr Dr Jason,
We’ve all seen the scatter plots of the “iris data” of sepal length vs sepal width, for the three different iris species, setosa, versicolor and virginica, an example is at https://bigqlabsdotcom.files.wordpress.com/2016/06/iris_data-scatter-plot-11.png?w=620 source https://bigqlabs.com/2016/06/27/training-a-naive-bayes-classifier-using-sklearn/ .
We see in the plots where there is some overlap in lengths, that is some of versicolors’s lengths are the same as the virginica.
Where there is overlap, is there any algorithm that can make a distinction between a virginica and versicolor?
Put it another way, are there any additional variables needed to increase the accuracy of prediction between the three species?
This is because so far relying on sepal length and sepal width is not enough.
Thank you,
Anthony of Sydney
Perhaps you can use the available data to engineer new input features?
Dear Sir
i am very thankful of your valued information which you send me. These are very useful for my education and I am waiting for you more coperation like this
Thanks, I’m happy the tutorials are helpful!
Good work sir. Pls, how do I apply naive Bayes rule in predicting road traffic congestion. Eg, if I am monitoring traffic in about 3 different routes say, a, b and c. Reply me pls.
I recommend following this process:
https://machinelearningmastery.com/start-here/#process
What is the base value of the logarithm of probabilities?
We use add log probs to avoid multiplying many small numbers which can result in an underflow.
easier naive bayes implementation is below:
https://github.com/Bhavya112298/Machine-Learning-algorithms-scratch-implementation
Thanks for sharing.
hey i have question what mean fit_prior and class_prior i dont understand, i have read the documentation and i dont understand. thx before
hat don’t you understand exactly?
yes i dont understand what is mean can u make it simple or example what fit_prior and class_prior used to? sorry for bad english
A class prior is the probability of any observation belonging to that class, give no information.
No idea what you mean by the fit prior, I’ve not heard of that before.
thx for answer sir!
A very well written article. Well I am working on a PGM query model and was hunting around for ideas on how best to represent a CPD. Should it be a dict of dicts or dataframe etc or a combination of both where the indexes are the enumeration of the “inedges” with the possible node values being the columns. Will probably look around or if not build a custom object. And if I’m really upset I’ll do it in Java ????
Perhaps prototype a few approaches?
Thank you so much for this, Sir!
You’re welcome!
Thank you for the tutorial. I appreciate the naive Bayes concept, but still have issues while trying to classify dataset from user ratings of products into two labels [similar ratings; dissimilar rating] using the Naive Bayes classifier.
Note, am using ‘AppleStore.csv’ dataset.
kindly, help, am very new in this territory.
Perhaps one of the methods here will help:
https://scikit-learn.org/stable/modules/naive_bayes.html
Can it be used for 4 classes case?
Sure.
Hi Jason, Can we apply conditional probability and Bayes classifier in case of non-Gaussian distributions for class distributions? Do we have any cost functions that measure the classification model uncertainty with respect to non-Gaussian densities of the class distributions?
You can use another distribution for a feature, even a kernel or empirical distribution.
Not sure about using a conditional distribution, sorry.
Think of each feature/distribution as a model and naive bayes as an ensemble of these smaller models.
Hi Jason,
I have a data set in which I have both categorical variables and numerical variables. I decided to use binomial distribution for the categorical variables(A,B,C) and gaussian distribution for the numerical variables(D,E,F).
I don’t want to construct the naive bayes model from scratch. I there any way in sklearn or any other library in python to tell the naive bayes model that it has to treat variable A,B,C as binomial distribution and variable D,E,F as gaussian distribution?
Thanks for reply
Great question.
Off the cuff, I don’t think the sklearn implementations support mixed types. You could have one model for each variable type and ensemble their predictions together.
Hi Jason,
Following you I read an article on ensemble learning.
Below is the link:
https://www.datacamp.com/community/tutorials/ensemble-learning-python
In this article it is written, “Also, if you need to work in a probabilistic setting, ensemble methods may not work either. ”
You can check this by yourself. So ensemble method is not going to work for naive bayes. Is there any alternative to mixed types in python ?
Thanks for reply.
I disagree with the statement and I have not read the post. Probabilistic methods are used in ensembles all the time and are effective.
Hi,
Nice article
But I think I see a problem:
When you use the norm.pdf function, you get the distribution value y at the supplied x value – the likelihood. The height of the distribution is not normalized to 1 (the area under it, is) and this means you can get all kind of numbers that have no true relation to probability between the distributions, as one pdf value can be lower than another even though it is more likely to be a sample of the first.
This will not be a hugh problem when two distribution of the same feature (for two classes) have a similar standard deviation. But if, for example, one distribution has a SD that is 10 times bigger than the 2nd one, it will return a pdf value that is 10 time lower for the a sample with the same z score.
Would be happy to hear your thoughts.
That is the nature of pdf. It is a density function (which can be any value), and shouldn’t be interpreted as probability (which is between 0 and 1). If you want a probability-like value, you may consider norm.cdf (cumulative distribution function) but you must understand what you’re doing. It is not always a direct substitution.
Hi Jason, very thankful for the valuable information you have shared in the article. The concepts proved very helpful still waiting for more content like this. The tutorials are helpful. I appreciate the naive Bayes concept. I have also gone through https://www.edvanza.com/ perhaps one of the methods here will help.
Thank you for the feedback and suggestion Ernst!