Gradient descent is an optimization algorithm that follows the negative gradient of an objective function in order to locate the minimum of the function.
A limitation of gradient descent is that it uses the same step size (learning rate) for each input variable. AdaGrad, for short, is an extension of the gradient descent optimization algorithm that allows the step size in each dimension used by the optimization algorithm to be automatically adapted based on the gradients seen for the variable (partial derivatives) over the course of the search.
A limitation of AdaGrad is that it can result in a very small step size for each parameter by the end of the search that can slow the progress of the search down too much and may mean not locating the optima.
Root Mean Squared Propagation, or RMSProp, is an extension of gradient descent and the AdaGrad version of gradient descent that uses a decaying average of partial gradients in the adaptation of the step size for each parameter. The use of a decaying moving average allows the algorithm to forget early gradients and focus on the most recently observed partial gradients seen during the progress of the search, overcoming the limitation of AdaGrad.
In this tutorial, you will discover how to develop the gradient descent with RMSProp optimization algorithm from scratch.
After completing this tutorial, you will know:
- Gradient descent is an optimization algorithm that uses the gradient of the objective function to navigate the search space.
- Gradient descent can be updated to use an automatically adaptive step size for each input variable using a decaying moving average of partial derivatives, called RMSProp.
- How to implement the RMSProp optimization algorithm from scratch and apply it to an objective function and evaluate the results.
Kick-start your project with my new book Optimization for Machine Learning, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.
Gradient Descent With RMSProp from Scratch
Photo by pavel ahmed, some rights reserved.
Tutorial Overview
This tutorial is divided into three parts; they are:
- Gradient Descent
- Root Mean Squared Propagation (RMSProp)
- Gradient Descent With RMSProp
- Two-Dimensional Test Problem
- Gradient Descent Optimization With RMSProp
- Visualization of RMSProp
Gradient Descent
Gradient descent is an optimization algorithm.
It is technically referred to as a first-order optimization algorithm as it explicitly makes use of the first order derivative of the target objective function.
First-order methods rely on gradient information to help direct the search for a minimum …
— Page 69, Algorithms for Optimization, 2019.
The first order derivative, or simply the “derivative,” is the rate of change or slope of the target function at a specific point, e.g. for a specific input.
If the target function takes multiple input variables, it is referred to as a multivariate function and the input variables can be thought of as a vector. In turn, the derivative of a multivariate target function may also be taken as a vector and is referred to generally as the gradient.
- Gradient: First order derivative for a multivariate objective function.
The derivative or the gradient points in the direction of the steepest ascent of the target function for a specific input.
Gradient descent refers to a minimization optimization algorithm that follows the negative of the gradient downhill of the target function to locate the minimum of the function.
The gradient descent algorithm requires a target function that is being optimized and the derivative function for the objective function. The target function f() returns a score for a given set of inputs, and the derivative function f'() gives the derivative of the target function for a given set of inputs.
The gradient descent algorithm requires a starting point (x) in the problem, such as a randomly selected point in the input space.
The derivative is then calculated and a step is taken in the input space that is expected to result in a downhill movement in the target function, assuming we are minimizing the target function.
A downhill movement is made by first calculating how far to move in the input space, calculated as the step size (called alpha or the learning rate) multiplied by the gradient. This is then subtracted from the current point, ensuring we move against the gradient, or down the target function.
- x = x – step_size * f'(x)
The steeper the objective function at a given point, the larger the magnitude of the gradient, and in turn, the larger the step taken in the search space. The size of the step taken is scaled using a step size hyperparameter.
- Step Size (alpha): Hyperparameter that controls how far to move in the search space against the gradient each iteration of the algorithm.
If the step size is too small, the movement in the search space will be small and the search will take a long time. If the step size is too large, the search may bounce around the search space and skip over the optima.
Now that we are familiar with the gradient descent optimization algorithm, let’s take a look at RMSProp.
Want to Get Started With Optimization Algorithms?
Take my free 7-day email crash course now (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
Root Mean Squared Propagation (RMSProp)
Root Mean Squared Propagation, or RMSProp for short, is an extension to the gradient descent optimization algorithm.
It is an unpublished extension, first described in Geoffrey Hinton’s lecture notes for his Coursera course on neural networks, specifically Lecture 6e titled “rmsprop: Divide the gradient by a running average of its recent magnitude.”
RMSProp is designed to accelerate the optimization process, e.g. decrease the number of function evaluations required to reach the optima, or to improve the capability of the optimization algorithm, e.g. result in a better final result.
It is related to another extension to gradient descent called Adaptive Gradient, or AdaGrad.
AdaGrad is designed to specifically explore the idea of automatically tailoring the step size (learning rate) for each parameter in the search space. This is achieved by first calculating a step size for a given dimension, then using the calculated step size to make a movement in that dimension using the partial derivative. This process is then repeated for each dimension in the search space.
Adagrad calculates the step size for each parameter by first summing the partial derivatives for the parameter seen so far during the search, then dividing the initial step size hyperparameter by the square root of the sum of the squared partial derivatives.
The calculation of the custom step size for one parameter is as follows:
- cust_step_size = step_size / (1e-8 + sqrt(s))
Where cust_step_size is the calculated step size for an input variable for a given point during the search, step_size is the initial step size, sqrt() is the square root operation and s is the sum of the squared partial derivatives for the input variable seen during the search so far.
This has the effect of smoothing out the oscillations in the search for optimization problems that have a lot of curvature in the search space.
AdaGrad shrinks the learning rate according to the entire history of the squared gradient and may have made the learning rate too small before arriving at such a convex structure.
— Pages 307-308, Deep Learning, 2016.
A problem with AdaGrad is that it can slow the search down too much, resulting in very small learning rates for each parameter or dimension of the search by the end of the run. This has the effect of stopping the search too soon, before the minimal can be located.
RMSProp extends Adagrad to avoid the effect of a monotonically decreasing learning rate.
— Page 78, Algorithms for Optimization, 2019.
RMSProp can be thought of as an extension of AdaGrad in that it uses a decaying average or moving average of the partial derivatives instead of the sum in the calculation of the learning rate for each parameter.
This is achieved by adding a new hyperparameter we will call rho that acts like momentum for the partial derivatives.
RMSProp maintains a decaying average of squared gradients.
— Page 78, Algorithms for Optimization, 2019.
Using a decaying moving average of the partial derivative allows the search to forget early partial derivative values and focus on the most recently seen shape of the search space.
RMSProp uses an exponentially decaying average to discard history from the extreme past so that it can converge rapidly after finding a convex bowl, as if it were an instance of the AdaGrad algorithm initialized within that bowl.
— Page 308, Deep Learning, 2016.
The calculation of the mean squared partial derivative for one parameter is as follows:
- s(t+1) = (s(t) * rho) + (f'(x(t))^2 * (1.0-rho))
Where s(t+1) is the decaying moving average of the squared partial derivative for one parameter for the current iteration of the algorithm, s(t) is the decaying moving average squared partial derivative for the previous iteration, f'(x(t))^2 is the squared partial derivative for the current parameter, and rho is a hyperparameter, typically with the value of 0.9 like momentum.
Given that we are using a decaying average of the partial derivatives and calculating the square root of this average gives the technique its name, e.g, square root of the mean squared partial derivatives or root mean square (RMS). For example, the custom step size for a parameter may be written as:
- cust_step_size(t+1) = step_size / (1e-8 + RMS(s(t+1)))
Once we have the custom step size for the parameter, we can update the parameter using the custom step size and the partial derivative f'(x(t)).
- x(t+1) = x(t) – cust_step_size(t+1) * f'(x(t))
This process is then repeated for each input variable until a new point in the search space is created and can be evaluated.
RMSProp is a very effective extension of gradient descent and is one of the preferred approaches generally used to fit deep learning neural networks.
Empirically, RMSProp has been shown to be an effective and practical optimization algorithm for deep neural networks. It is currently one of the go-to optimization methods being employed routinely by deep learning practitioners.
— Page 308, Deep Learning, 2016.
Now that we are familiar with the RMSprop algorithm, let’s explore how we might implement it and evaluate its performance.
Gradient Descent With RMSProp
In this section, we will explore how to implement the gradient descent optimization algorithm with adaptive gradients using the RMSProp algorithm.
Two-Dimensional Test Problem
First, let’s define an optimization function.
We will use a simple two-dimensional function that squares the input of each dimension and define the range of valid inputs from -1.0 to 1.0.
The objective() function below implements this function
|
1 2 3 |
# objective function def objective(x, y): return x**2.0 + y**2.0 |
We can create a three-dimensional plot of the dataset to get a feeling for the curvature of the response surface.
The complete example of plotting the objective function is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 |
# 3d plot of the test function from numpy import arange from numpy import meshgrid from matplotlib import pyplot # objective function def objective(x, y): return x**2.0 + y**2.0 # define range for input r_min, r_max = -1.0, 1.0 # sample input range uniformly at 0.1 increments xaxis = arange(r_min, r_max, 0.1) yaxis = arange(r_min, r_max, 0.1) # create a mesh from the axis x, y = meshgrid(xaxis, yaxis) # compute targets results = objective(x, y) # create a surface plot with the jet color scheme figure = pyplot.figure() axis = figure.gca(projection='3d') axis.plot_surface(x, y, results, cmap='jet') # show the plot pyplot.show() |
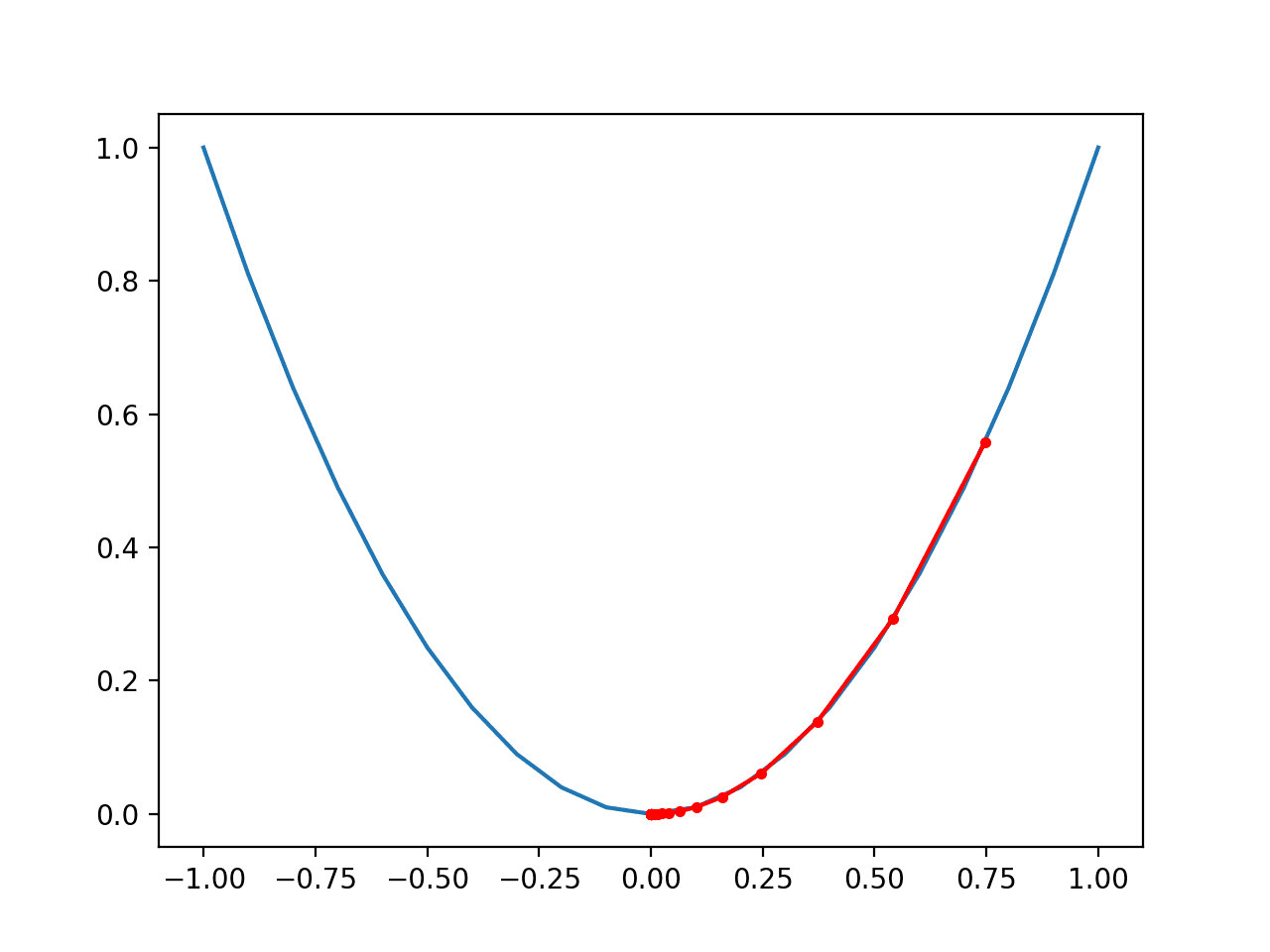
Running the example creates a three-dimensional surface plot of the objective function.
We can see the familiar bowl shape with the global minima at f(0, 0) = 0.
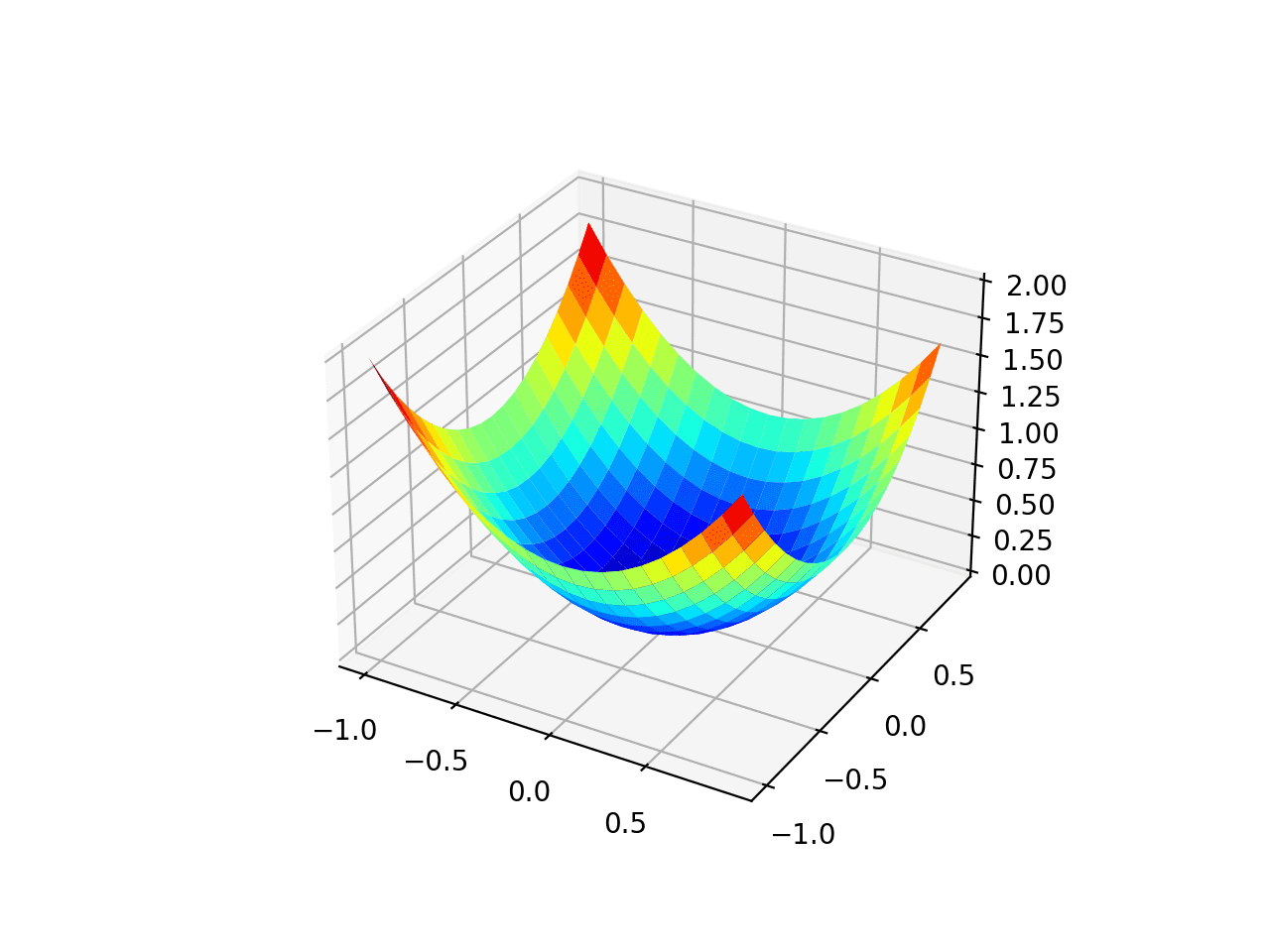
Three-Dimensional Plot of the Test Objective Function
We can also create a two-dimensional plot of the function. This will be helpful later when we want to plot the progress of the search.
The example below creates a contour plot of the objective function.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 |
# contour plot of the test function from numpy import asarray from numpy import arange from numpy import meshgrid from matplotlib import pyplot # objective function def objective(x, y): return x**2.0 + y**2.0 # define range for input bounds = asarray([[-1.0, 1.0], [-1.0, 1.0]]) # sample input range uniformly at 0.1 increments xaxis = arange(bounds[0,0], bounds[0,1], 0.1) yaxis = arange(bounds[1,0], bounds[1,1], 0.1) # create a mesh from the axis x, y = meshgrid(xaxis, yaxis) # compute targets results = objective(x, y) # create a filled contour plot with 50 levels and jet color scheme pyplot.contourf(x, y, results, levels=50, cmap='jet') # show the plot pyplot.show() |
Running the example creates a two-dimensional contour plot of the objective function.
We can see the bowl shape compressed to contours shown with a color gradient. We will use this plot to plot the specific points explored during the progress of the search.
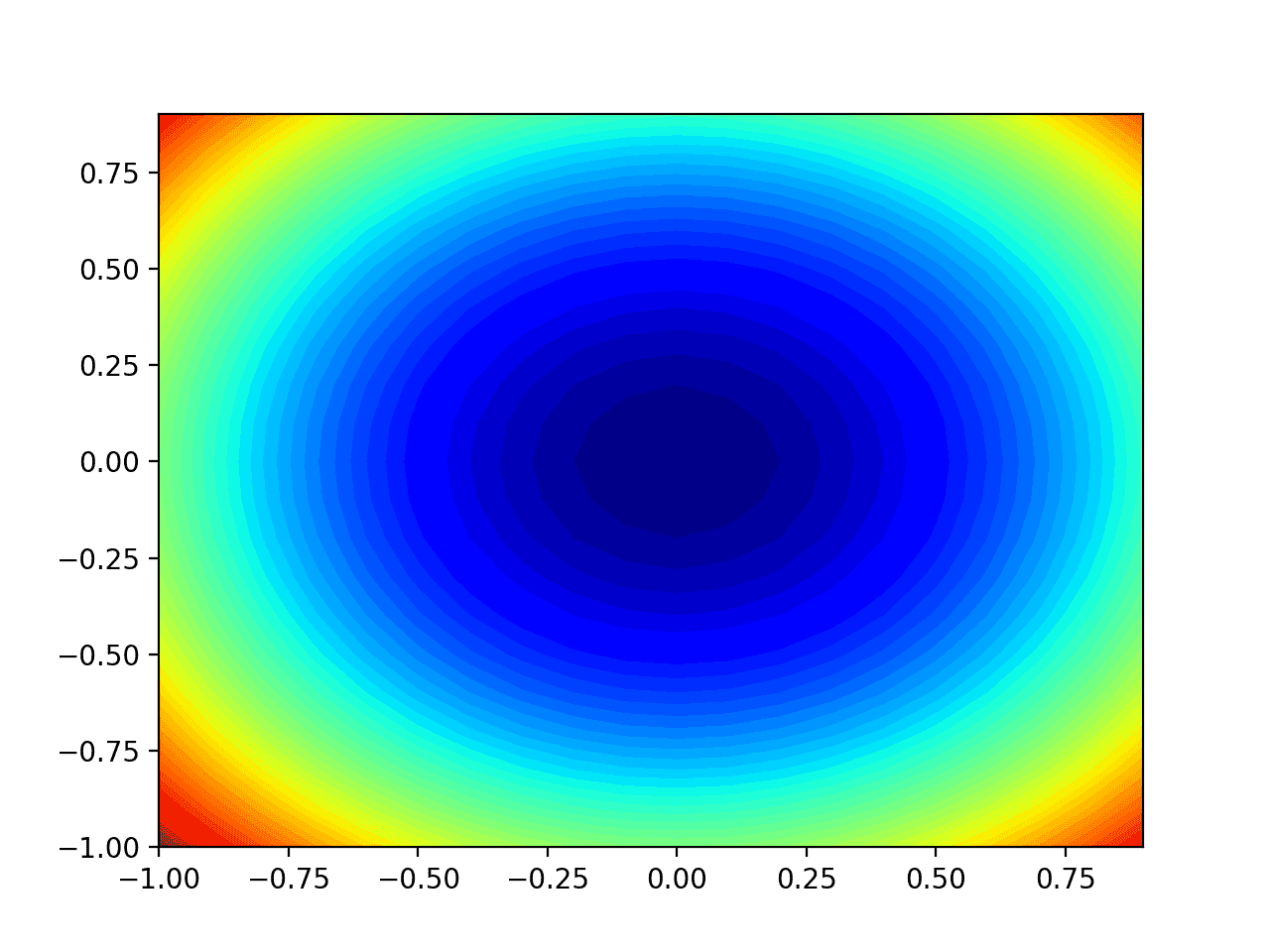
Two-Dimensional Contour Plot of the Test Objective Function
Now that we have a test objective function, let’s look at how we might implement the RMSProp optimization algorithm.
Gradient Descent Optimization With RMSProp
We can apply the gradient descent with RMSProp to the test problem.
First, we need a function that calculates the derivative for this function.
- f(x) = x^2
- f'(x) = x * 2
The derivative of x^2 is x * 2 in each dimension. The derivative() function implements this below.
|
1 2 3 |
# derivative of objective function def derivative(x, y): return asarray([x * 2.0, y * 2.0]) |
Next, we can implement gradient descent optimization.
First, we can select a random point in the bounds of the problem as a starting point for the search.
This assumes we have an array that defines the bounds of the search with one row for each dimension and the first column defines the minimum and the second column defines the maximum of the dimension.
|
1 2 3 |
... # generate an initial point solution = bounds[:, 0] + rand(len(bounds)) * (bounds[:, 1] - bounds[:, 0]) |
Next, we need to initialize the decay average of the squared partial derivatives for each dimension to 0.0 values.
|
1 2 3 |
... # list of the average square gradients for each variable sq_grad_avg = [0.0 for _ in range(bounds.shape[0])] |
We can then enumerate a fixed number of iterations of the search optimization algorithm defined by a “n_iter” hyperparameter.
|
1 2 3 4 |
... # run the gradient descent for it in range(n_iter): ... |
The first step is to calculate the gradient for the current solution using the derivative() function.
|
1 2 3 |
... # calculate gradient gradient = derivative(solution[0], solution[1]) |
We then need to calculate the square of the partial derivative and update the decaying average of the squared partial derivatives with the “rho” hyperparameter.
|
1 2 3 4 5 6 7 |
... # update the average of the squared partial derivatives for i in range(gradient.shape[0]): # calculate the squared gradient sg = gradient[i]**2.0 # update the moving average of the squared gradient sq_grad_avg[i] = (sq_grad_avg[i] * rho) + (sg * (1.0-rho)) |
We can then use the moving average of the squared partial derivatives and gradient to calculate the step size for the next point.
We will do this one variable at a time, first calculating the step size for the variable, then the new value for the variable. These values are built up in an array until we have a completely new solution that is in the steepest descent direction from the current point using the custom step sizes.
|
1 2 3 4 5 6 7 8 9 10 |
... # build a solution one variable at a time new_solution = list() for i in range(solution.shape[0]): # calculate the step size for this variable alpha = step_size / (1e-8 + sqrt(sq_grad_avg[i])) # calculate the new position in this variable value = solution[i] - alpha * gradient[i] # store this variable new_solution.append(value) |
This new solution can then be evaluated using the objective() function and the performance of the search can be reported.
|
1 2 3 4 5 6 |
... # evaluate candidate point solution = asarray(new_solution) solution_eval = objective(solution[0], solution[1]) # report progress print('>%d f(%s) = %.5f' % (it, solution, solution_eval)) |
And that’s it.
We can tie all of this together into a function named rmsprop() that takes the names of the objective function and the derivative function, an array with the bounds of the domain and hyperparameter values for the total number of algorithm iterations and the initial learning rate, and returns the final solution and its evaluation.
This complete function is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 |
# gradient descent algorithm with rmsprop def rmsprop(objective, derivative, bounds, n_iter, step_size, rho): # generate an initial point solution = bounds[:, 0] + rand(len(bounds)) * (bounds[:, 1] - bounds[:, 0]) # list of the average square gradients for each variable sq_grad_avg = [0.0 for _ in range(bounds.shape[0])] # run the gradient descent for it in range(n_iter): # calculate gradient gradient = derivative(solution[0], solution[1]) # update the average of the squared partial derivatives for i in range(gradient.shape[0]): # calculate the squared gradient sg = gradient[i]**2.0 # update the moving average of the squared gradient sq_grad_avg[i] = (sq_grad_avg[i] * rho) + (sg * (1.0-rho)) # build a solution one variable at a time new_solution = list() for i in range(solution.shape[0]): # calculate the step size for this variable alpha = step_size / (1e-8 + sqrt(sq_grad_avg[i])) # calculate the new position in this variable value = solution[i] - alpha * gradient[i] # store this variable new_solution.append(value) # evaluate candidate point solution = asarray(new_solution) solution_eval = objective(solution[0], solution[1]) # report progress print('>%d f(%s) = %.5f' % (it, solution, solution_eval)) return [solution, solution_eval] |
Note: we have intentionally used lists and imperative coding style instead of vectorized operations for readability. Feel free to adapt the implementation to a vectorization implementation with NumPy arrays for better performance.
We can then define our hyperparameters and call the rmsprop() function to optimize our test objective function.
In this case, we will use 50 iterations of the algorithm, an initial learning rate of 0.01, and a value of 0.99 for the rho hyperparameter, all chosen after a little trial and error.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 |
... # seed the pseudo random number generator seed(1) # define range for input bounds = asarray([[-1.0, 1.0], [-1.0, 1.0]]) # define the total iterations n_iter = 50 # define the step size step_size = 0.01 # momentum for rmsprop rho = 0.99 # perform the gradient descent search with rmsprop best, score = rmsprop(objective, derivative, bounds, n_iter, step_size, rho) print('Done!') print('f(%s) = %f' % (best, score)) |
Tying all of this together, the complete example of gradient descent optimization with RMSProp is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 |
# gradient descent optimization with rmsprop for a two-dimensional test function from math import sqrt from numpy import asarray from numpy.random import rand from numpy.random import seed # objective function def objective(x, y): return x**2.0 + y**2.0 # derivative of objective function def derivative(x, y): return asarray([x * 2.0, y * 2.0]) # gradient descent algorithm with rmsprop def rmsprop(objective, derivative, bounds, n_iter, step_size, rho): # generate an initial point solution = bounds[:, 0] + rand(len(bounds)) * (bounds[:, 1] - bounds[:, 0]) # list of the average square gradients for each variable sq_grad_avg = [0.0 for _ in range(bounds.shape[0])] # run the gradient descent for it in range(n_iter): # calculate gradient gradient = derivative(solution[0], solution[1]) # update the average of the squared partial derivatives for i in range(gradient.shape[0]): # calculate the squared gradient sg = gradient[i]**2.0 # update the moving average of the squared gradient sq_grad_avg[i] = (sq_grad_avg[i] * rho) + (sg * (1.0-rho)) # build a solution one variable at a time new_solution = list() for i in range(solution.shape[0]): # calculate the step size for this variable alpha = step_size / (1e-8 + sqrt(sq_grad_avg[i])) # calculate the new position in this variable value = solution[i] - alpha * gradient[i] # store this variable new_solution.append(value) # evaluate candidate point solution = asarray(new_solution) solution_eval = objective(solution[0], solution[1]) # report progress print('>%d f(%s) = %.5f' % (it, solution, solution_eval)) return [solution, solution_eval] # seed the pseudo random number generator seed(1) # define range for input bounds = asarray([[-1.0, 1.0], [-1.0, 1.0]]) # define the total iterations n_iter = 50 # define the step size step_size = 0.01 # momentum for rmsprop rho = 0.99 # perform the gradient descent search with rmsprop best, score = rmsprop(objective, derivative, bounds, n_iter, step_size, rho) print('Done!') print('f(%s) = %f' % (best, score)) |
Running the example applies the RMSProp optimization algorithm to our test problem and reports the performance of the search for each iteration of the algorithm.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see that a near optimal solution was found after perhaps 33 iterations of the search, with input values near 0.0 and 0.0, evaluating to 0.0.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 |
... >30 f([-9.61030898e-14 3.19352553e-03]) = 0.00001 >31 f([-3.42767893e-14 2.71513758e-03]) = 0.00001 >32 f([-1.21143047e-14 2.30636623e-03]) = 0.00001 >33 f([-4.24204875e-15 1.95738936e-03]) = 0.00000 >34 f([-1.47154482e-15 1.65972553e-03]) = 0.00000 >35 f([-5.05629595e-16 1.40605727e-03]) = 0.00000 >36 f([-1.72064649e-16 1.19007691e-03]) = 0.00000 >37 f([-5.79813754e-17 1.00635204e-03]) = 0.00000 >38 f([-1.93445677e-17 8.50208253e-04]) = 0.00000 >39 f([-6.38906842e-18 7.17626999e-04]) = 0.00000 >40 f([-2.08860690e-18 6.05156738e-04]) = 0.00000 >41 f([-6.75689941e-19 5.09835645e-04]) = 0.00000 >42 f([-2.16291217e-19 4.29124484e-04]) = 0.00000 >43 f([-6.84948980e-20 3.60848338e-04]) = 0.00000 >44 f([-2.14551097e-20 3.03146089e-04]) = 0.00000 >45 f([-6.64629576e-21 2.54426642e-04]) = 0.00000 >46 f([-2.03575780e-21 2.13331041e-04]) = 0.00000 >47 f([-6.16437387e-22 1.78699710e-04]) = 0.00000 >48 f([-1.84495110e-22 1.49544152e-04]) = 0.00000 >49 f([-5.45667355e-23 1.25022522e-04]) = 0.00000 Done! f([-5.45667355e-23 1.25022522e-04]) = 0.000000 |
Visualization of RMSProp
We can plot the progress of the search on a contour plot of the domain.
This can provide an intuition for the progress of the search over the iterations of the algorithm.
We must update the rmsprop() function to maintain a list of all solutions found during the search, then return this list at the end of the search.
The updated version of the function with these changes is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 |
# gradient descent algorithm with rmsprop def rmsprop(objective, derivative, bounds, n_iter, step_size, rho): # track all solutions solutions = list() # generate an initial point solution = bounds[:, 0] + rand(len(bounds)) * (bounds[:, 1] - bounds[:, 0]) # list of the average square gradients for each variable sq_grad_avg = [0.0 for _ in range(bounds.shape[0])] # run the gradient descent for it in range(n_iter): # calculate gradient gradient = derivative(solution[0], solution[1]) # update the average of the squared partial derivatives for i in range(gradient.shape[0]): # calculate the squared gradient sg = gradient[i]**2.0 # update the moving average of the squared gradient sq_grad_avg[i] = (sq_grad_avg[i] * rho) + (sg * (1.0-rho)) # build solution new_solution = list() for i in range(solution.shape[0]): # calculate the learning rate for this variable alpha = step_size / (1e-8 + sqrt(sq_grad_avg[i])) # calculate the new position in this variable value = solution[i] - alpha * gradient[i] new_solution.append(value) # store the new solution solution = asarray(new_solution) solutions.append(solution) # evaluate candidate point solution_eval = objective(solution[0], solution[1]) # report progress print('>%d f(%s) = %.5f' % (it, solution, solution_eval)) return solutions |
We can then execute the search as before, and this time retrieve the list of solutions instead of the best final solution.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 |
... # seed the pseudo random number generator seed(1) # define range for input bounds = asarray([[-1.0, 1.0], [-1.0, 1.0]]) # define the total iterations n_iter = 50 # define the step size step_size = 0.01 # momentum for rmsprop rho = 0.99 # perform the gradient descent search with rmsprop solutions = rmsprop(objective, derivative, bounds, n_iter, step_size, rho) |
We can then create a contour plot of the objective function, as before.
|
1 2 3 4 5 6 7 8 9 10 |
... # sample input range uniformly at 0.1 increments xaxis = arange(bounds[0,0], bounds[0,1], 0.1) yaxis = arange(bounds[1,0], bounds[1,1], 0.1) # create a mesh from the axis x, y = meshgrid(xaxis, yaxis) # compute targets results = objective(x, y) # create a filled contour plot with 50 levels and jet color scheme pyplot.contourf(x, y, results, levels=50, cmap='jet') |
Finally, we can plot each solution found during the search as a white dot connected by a line.
|
1 2 3 4 |
... # plot the sample as black circles solutions = asarray(solutions) pyplot.plot(solutions[:, 0], solutions[:, 1], '.-', color='w') |
Tying this all together, the complete example of performing the RMSProp optimization on the test problem and plotting the results on a contour plot is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 |
# example of plotting the rmsprop search on a contour plot of the test function from math import sqrt from numpy import asarray from numpy import arange from numpy.random import rand from numpy.random import seed from numpy import meshgrid from matplotlib import pyplot from mpl_toolkits.mplot3d import Axes3D # objective function def objective(x, y): return x**2.0 + y**2.0 # derivative of objective function def derivative(x, y): return asarray([x * 2.0, y * 2.0]) # gradient descent algorithm with rmsprop def rmsprop(objective, derivative, bounds, n_iter, step_size, rho): # track all solutions solutions = list() # generate an initial point solution = bounds[:, 0] + rand(len(bounds)) * (bounds[:, 1] - bounds[:, 0]) # list of the average square gradients for each variable sq_grad_avg = [0.0 for _ in range(bounds.shape[0])] # run the gradient descent for it in range(n_iter): # calculate gradient gradient = derivative(solution[0], solution[1]) # update the average of the squared partial derivatives for i in range(gradient.shape[0]): # calculate the squared gradient sg = gradient[i]**2.0 # update the moving average of the squared gradient sq_grad_avg[i] = (sq_grad_avg[i] * rho) + (sg * (1.0-rho)) # build solution new_solution = list() for i in range(solution.shape[0]): # calculate the learning rate for this variable alpha = step_size / (1e-8 + sqrt(sq_grad_avg[i])) # calculate the new position in this variable value = solution[i] - alpha * gradient[i] new_solution.append(value) # store the new solution solution = asarray(new_solution) solutions.append(solution) # evaluate candidate point solution_eval = objective(solution[0], solution[1]) # report progress print('>%d f(%s) = %.5f' % (it, solution, solution_eval)) return solutions # seed the pseudo random number generator seed(1) # define range for input bounds = asarray([[-1.0, 1.0], [-1.0, 1.0]]) # define the total iterations n_iter = 50 # define the step size step_size = 0.01 # momentum for rmsprop rho = 0.99 # perform the gradient descent search with rmsprop solutions = rmsprop(objective, derivative, bounds, n_iter, step_size, rho) # sample input range uniformly at 0.1 increments xaxis = arange(bounds[0,0], bounds[0,1], 0.1) yaxis = arange(bounds[1,0], bounds[1,1], 0.1) # create a mesh from the axis x, y = meshgrid(xaxis, yaxis) # compute targets results = objective(x, y) # create a filled contour plot with 50 levels and jet color scheme pyplot.contourf(x, y, results, levels=50, cmap='jet') # plot the sample as black circles solutions = asarray(solutions) pyplot.plot(solutions[:, 0], solutions[:, 1], '.-', color='w') # show the plot pyplot.show() |
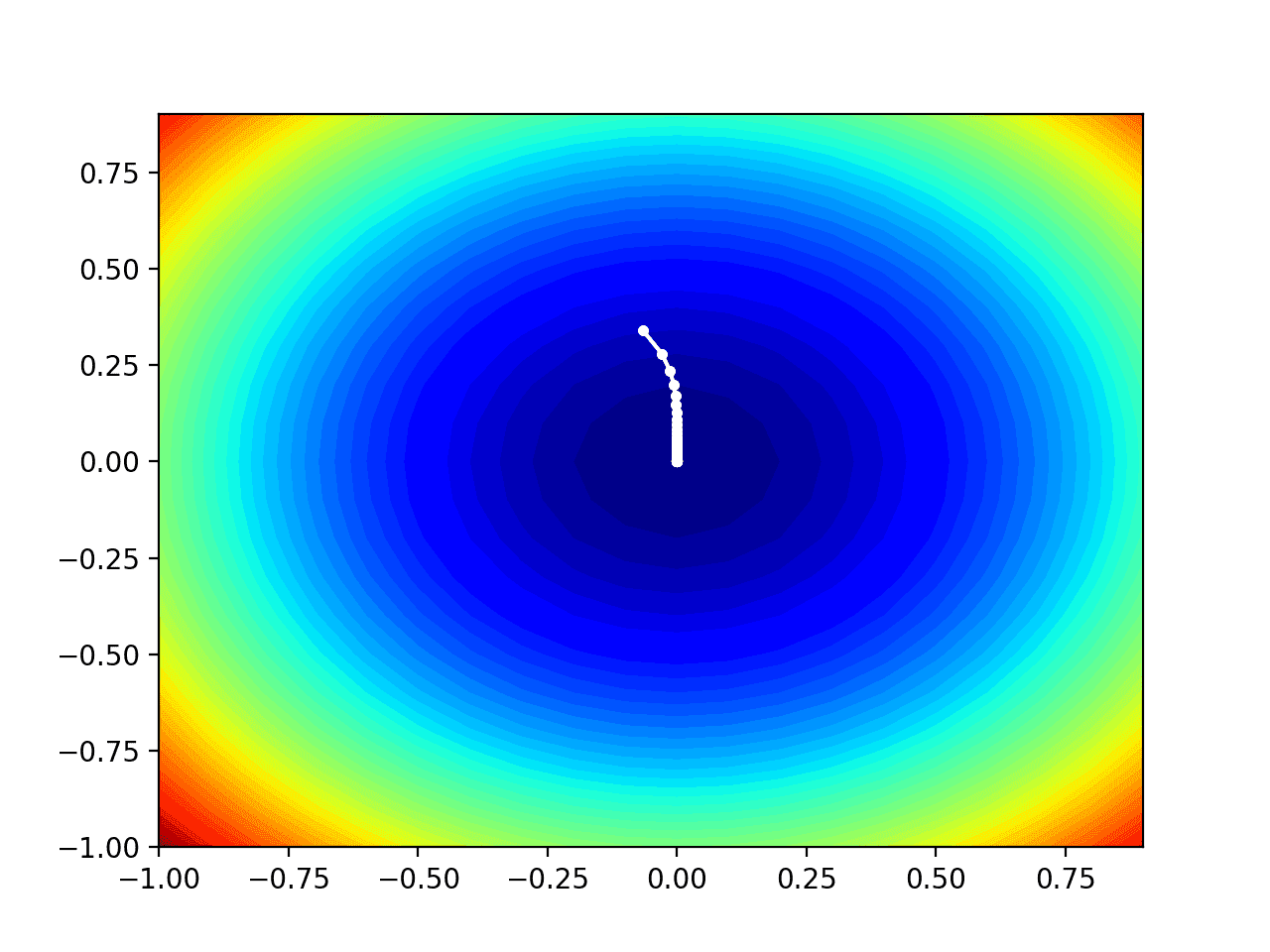
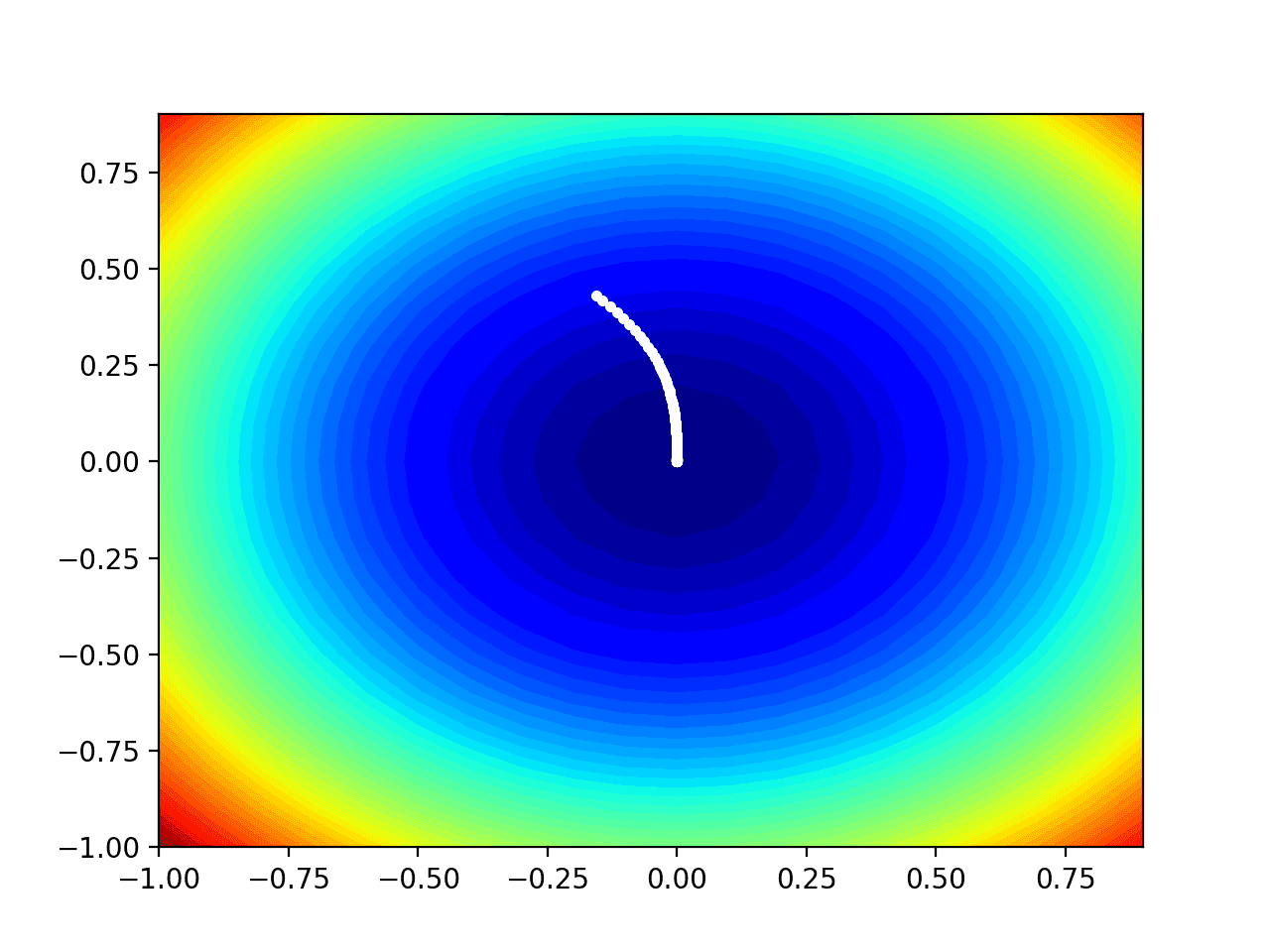
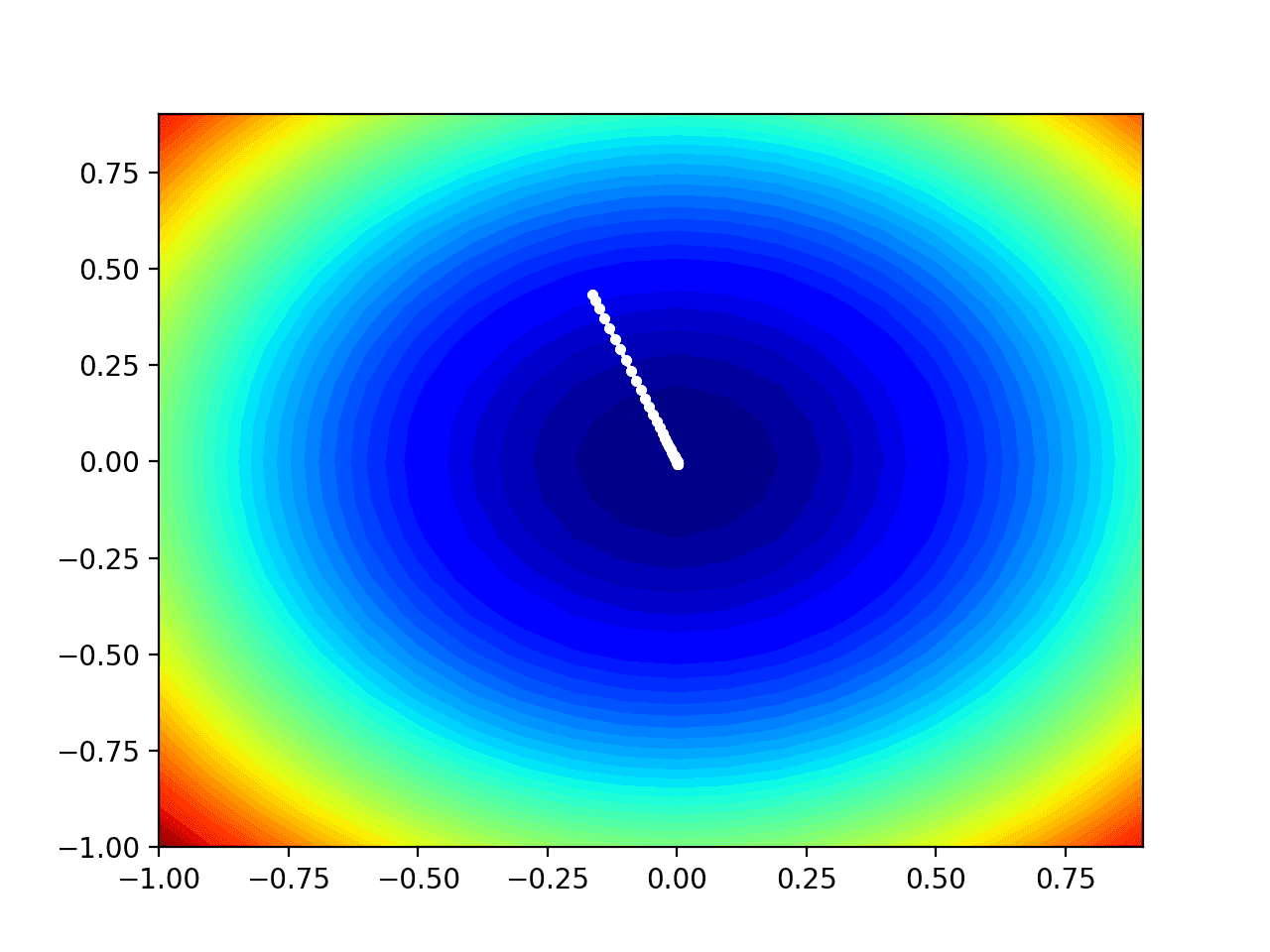
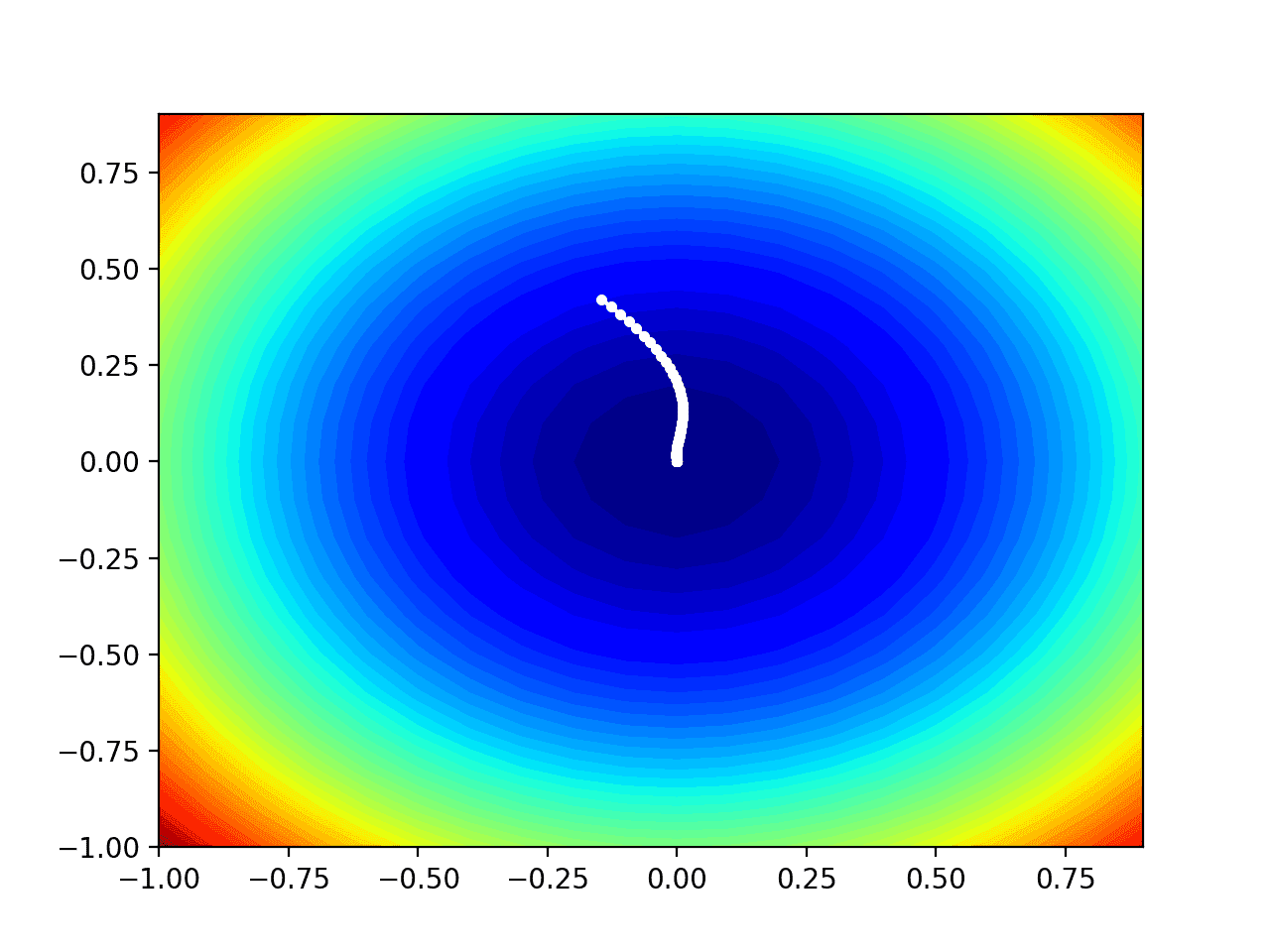
Running the example performs the search as before, except in this case, the contour plot of the objective function is created.
In this case, we can see that a white dot is shown for each solution found during the search, starting above the optima and progressively getting closer to the optima at the center of the plot.
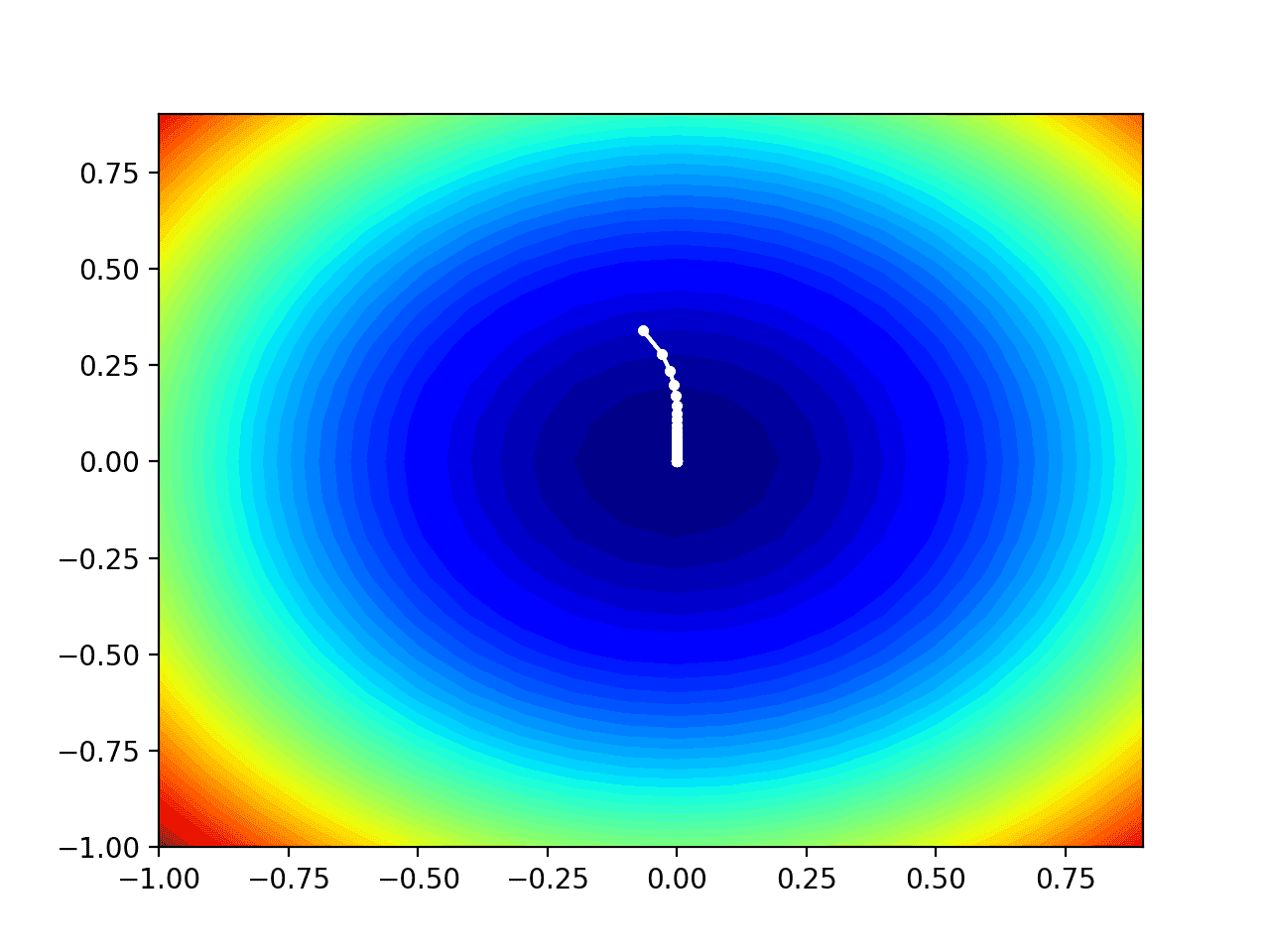
Contour Plot of the Test Objective Function With RMSProp Search Results Shown
Further Reading
This section provides more resources on the topic if you are looking to go deeper.
Papers
- Lecture 6e, rmsprop: Divide the gradient by a running average of its recent magnitude, Neural Networks for Machine Learning, Geoffrey Hinton.
Books
- Algorithms for Optimization, 2019.
- Deep Learning, 2016.
APIs
Articles
- Gradient descent, Wikipedia.
- Stochastic gradient descent, Wikipedia.
- An overview of gradient descent optimization algorithms, 2016.
Summary
In this tutorial, you discovered how to develop gradient descent with RMSProp optimization algorithm from scratch.
Specifically, you learned:
- Gradient descent is an optimization algorithm that uses the gradient of the objective function to navigate the search space.
- Gradient descent can be updated to use an automatically adaptive step size for each input variable using a decaying average of partial derivatives, called RMSProp.
- How to implement the RMSProp optimization algorithm from scratch and apply it to an objective function and evaluate the results.
Do you have any questions?
Ask your questions in the comments below and I will do my best to answer.

Really great article!
I am also interested in swarm particle optimization, do you have an article on that?
If not, what are most applications is it used for or in which cases?
Thanks.
Not at this stage.
PSO is used for optimizing high-dimensional real-valued functions without a derivative.
Thank you!! I was interested in switching it for in a neural network system since back propagation takes more time. What’s you thoughts on this?
You would not remove backprop, it would be used with RMSprop.
Thanks sir
You’re welcome.
Hi Jason,
Is it possbile to run RMSProp, Adadelta, Adamax and Nadam scratch formula in sklearn for optimization?
I am working on Sklearn MLP regressor and I would like to compare the above optimizer results with Adam in Sklearn?
If it is not possbile, could I replicate my Sklearn MLP model to Tensorflow and compare Optimizers there?
Pleased to hear from your side.
Best regards,
Faraz