Differential Evolution is a global optimization algorithm.
It is a type of evolutionary algorithm and is related to other evolutionary algorithms such as the genetic algorithm.
Unlike the genetic algorithm, it was specifically designed to operate upon vectors of real-valued numbers instead of bitstrings. Also unlike the genetic algorithm it uses vector operations like vector subtraction and addition to navigate the search space instead of genetics-inspired transforms.
In this tutorial, you will discover the Differential Evolution global optimization algorithm.
After completing this tutorial, you will know:
- Differential Evolution optimization is a type of evolutionary algorithm that is designed to work with real-valued candidate solutions.
- How to use the Differential Evolution optimization algorithm API in python.
- Examples of using Differential Evolution to solve global optimization problems with multiple optima.
Kick-start your project with my new book Optimization for Machine Learning, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.
Differential Evolution Global Optimization With Python
Photo by Gergely Csatari, some rights reserved.
Tutorial Overview
This tutorial is divided into three parts; they are:
- Differential Evolution
- Differential Evolution API
- Differential Evolution Worked Example
Differential Evolution
Differential Evolution, or DE for short, is a stochastic global search optimization algorithm.
It is a type of evolutionary algorithm and is related to other evolutionary algorithms such as the genetic algorithm. Unlike the genetic algorithm that represents candidate solutions using sequences of bits, Differential Evolution is designed to work with multi-dimensional real-valued candidate solutions for continuous objective functions.
Differential evolution (DE) is arguably one of the most powerful stochastic real-parameter optimization algorithms in current use.
— Differential Evolution: A Survey of the State-of-the-Art, 2011.
The algorithm does not make use of gradient information in the search, and as such, is well suited to non-differential nonlinear objective functions.
The algorithm works by maintaining a population of candidate solutions represented as real-valued vectors. New candidate solutions are created by making modified versions of existing solutions that then replace a large portion of the population each iteration of the algorithm.
New candidate solutions are created using a “strategy” that involves selecting a base solution to which a mutation is added, and other candidate solutions from the population from which the amount and type of mutation is calculated, called a difference vector. For example, a strategy may select a best candidate solution as the base and random solutions for the difference vector in the mutation.
DE generates new parameter vectors by adding the weighted difference between two population vectors to a third vector.
Base solutions are replaced by their children if the children have a better objective function evaluation.
Finally, after we have built up a new group of children, we compare each child with the parent which created it (each parent created a single child). If the child is better than the parent, it replaces the parent in the original population.
— Page 51, Essentials of Metaheuristics, 2011.
Mutation is calculated as the difference between pairs of candidate solutions that results in a difference vector that is then added to the base solution, weighted by a mutation factor hyperparameter set in the range [0,2].
Not all elements of a base solution are mutated. This is controlled via a recombination hyperparameter and is often set to a large value such as 80 percent, meaning most but not all variables in a base solution are replaced. The decision to keep or replace a value in a base solution is determined for each position separately by sampling a probability distribution such as a binomial or exponential.
A standard terminology is used to describe differential strategies with the form:
- DE/x/y/z
Where DE stands for “differential evolution,” x defines the base solution that is to be mutated, such as “rand” for random or “best” for the best solution in the population. The y stands for the number of difference vectors added to the base solution, such as 1, and z defines the probability distribution for determining if each solution is kept or replaced in the population, such as “bin” for binomial or “exp” for exponential.
The general convention used above is DE/x/y/z, where DE stands for “differential evolution,” x represents a string denoting the base vector to be perturbed, y is the number of difference vectors considered for perturbation of x, and z stands for the type of crossover being used (exp: exponential; bin: binomial).
— Differential Evolution: A Survey of the State-of-the-Art, 2011.
The configuration DE/best/1/bin and DE/best/2/bin are popular configurations as they perform well for many objective functions.
The experiments performed by Mezura-Montes et al. indicate that DE/best/1/bin (using always the best solution to find search directions and also binomial crossover) remained the most competitive scheme, regardless the characteristics of the problem to be solved, based on final accuracy and robustness of results.
— Differential Evolution: A Survey of the State-of-the-Art, 2011.
Now that we are familiar with the differential evolution algorithm, let’s look at how we might use the SciPy API implementation.
Want to Get Started With Optimization Algorithms?
Take my free 7-day email crash course now (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
Differential Evolution API
The Differential Evolution global optimization algorithm is available in Python via the differential_evolution() SciPy function.
The function takes the name of the objective function and the bounds of each input variable as minimum arguments for the search.
|
1 2 3 |
... # perform the differential evolution search result = differential_evolution(objective, bounds) |
There are a number of additional hyperparameters for the search that have default values, although you can configure them to customize the search.
A key hyperparameter is the “strategy” argument that controls the type of differential evolution search that is performed. By default, this is set to “best1bin” (DE/best/1/bin), which is a good configuration for most problems. It creates new candidate solutions by selecting random solutions from the population, subtracting one from the other, and adding a scaled version of the difference to the best candidate solution in the population.
- new = best + (mutation * (rand1 – rand2))
The “popsize” argument controls the size or number of candidate solutions that are maintained in the population. It is a factor of the number of dimensions in candidate solutions and by default, it is set to 15. That means for a 2D objective function that a population size of (2 * 15) or 30 candidate solutions will be maintained.
The total number of iterations of the algorithm is maintained by the “maxiter” argument and defaults to 1,000.
The “mutation” argument controls the number of changes made to candidate solutions each iteration. By default, this is set to 0.5. The amount of recombination is controlled via the “recombination” argument, which is set to 0.7 (70 percent of a given candidate solution) by default.
Finally, a local search is applied to the best candidate solution found at the end of the search. This is controlled via the “polish” argument, which by default is set to True.
The result of the search is an OptimizeResult object where properties can be accessed like a dictionary. The success (or not) of the search can be accessed via the ‘success‘ or ‘message‘ key.
The total number of function evaluations can be accessed via ‘nfev‘ and the optimal input found for the search is accessible via the ‘x‘ key.
Now that we are familiar with the differential evolution API in Python, let’s look at some worked examples.
Differential Evolution Worked Example
In this section, we will look at an example of using the differential evolution algorithm on a challenging objective function.
The Ackley function is an example of an objective function that has a single global optima and multiple local optima in which a local search might get stuck.
As such, a global optimization technique is required. It is a two-dimensional objective function that has a global optima at [0,0], which evaluates to 0.0.
The example below implements the Ackley and creates a three-dimensional surface plot showing the global optima and multiple local optima.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 |
# ackley multimodal function from numpy import arange from numpy import exp from numpy import sqrt from numpy import cos from numpy import e from numpy import pi from numpy import meshgrid from matplotlib import pyplot from mpl_toolkits.mplot3d import Axes3D # objective function def objective(x, y): return -20.0 * exp(-0.2 * sqrt(0.5 * (x**2 + y**2))) - exp(0.5 * (cos(2 * pi * x) + cos(2 * pi * y))) + e + 20 # define range for input r_min, r_max = -5.0, 5.0 # sample input range uniformly at 0.1 increments xaxis = arange(r_min, r_max, 0.1) yaxis = arange(r_min, r_max, 0.1) # create a mesh from the axis x, y = meshgrid(xaxis, yaxis) # compute targets results = objective(x, y) # create a surface plot with the jet color scheme figure = pyplot.figure() axis = figure.gca(projection='3d') axis.plot_surface(x, y, results, cmap='jet') # show the plot pyplot.show() |
Running the example creates the surface plot of the Ackley function showing the vast number of local optima.
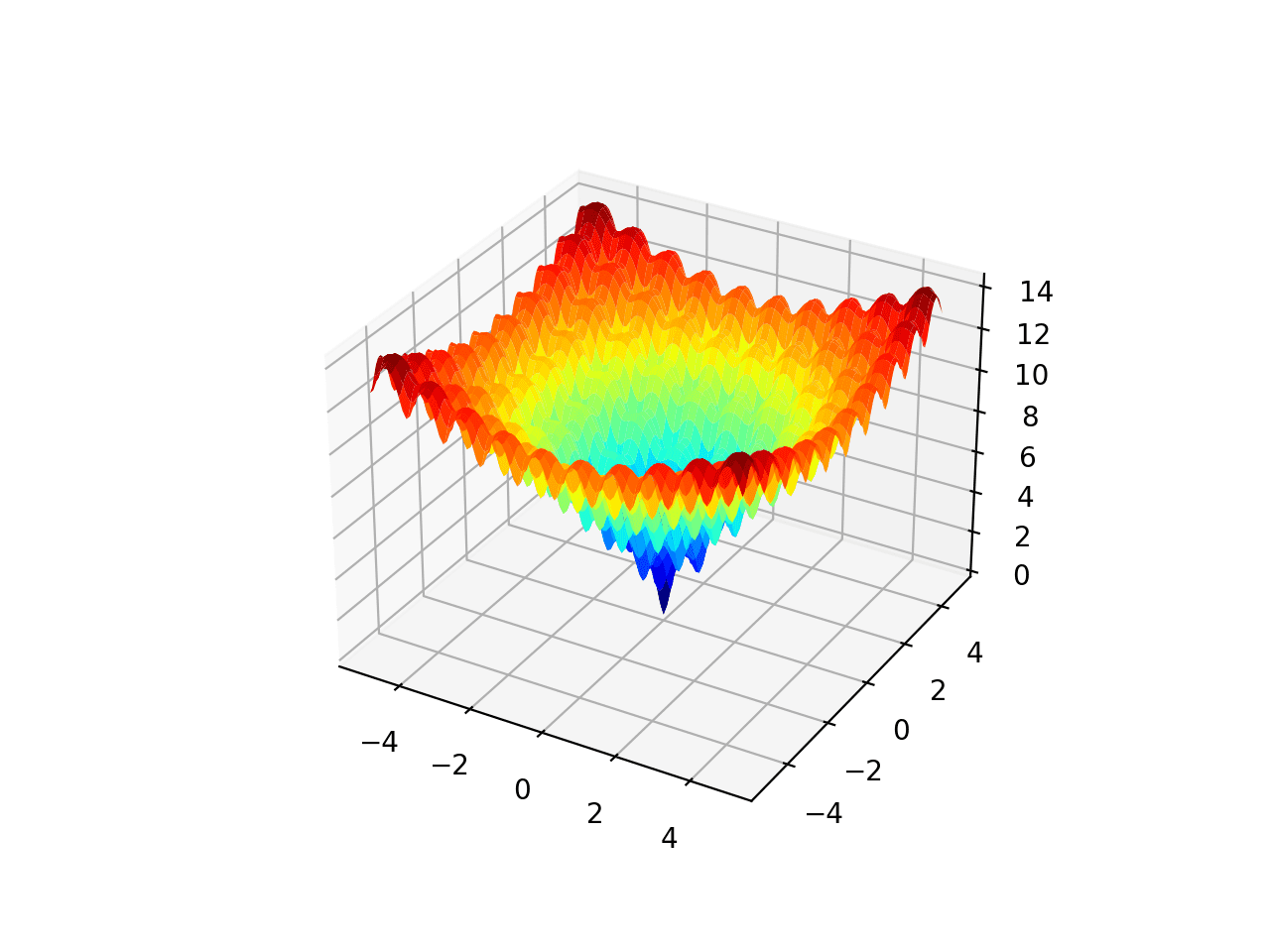
3D Surface Plot of the Ackley Multimodal Function
We can apply the differential evolution algorithm to the Ackley objective function.
First, we can define the bounds of the search space as the limits of the function in each dimension.
|
1 2 3 |
... # define the bounds on the search bounds = [[r_min, r_max], [r_min, r_max]] |
We can then apply the search by specifying the name of the objective function and the bounds of the search. In this case, we will use the default hyperparameters.
|
1 2 3 |
... # perform the differential evolution search result = differential_evolution(objective, bounds) |
After the search is complete, it will report the status of the search and the number of iterations performed, as well as the best result found with its evaluation.
|
1 2 3 4 5 6 7 8 |
... # summarize the result print('Status : %s' % result['message']) print('Total Evaluations: %d' % result['nfev']) # evaluate solution solution = result['x'] evaluation = objective(solution) print('Solution: f(%s) = %.5f' % (solution, evaluation)) |
Tying this together, the complete example of applying differential evolution to the Ackley objective function is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 |
# differential evolution global optimization for the ackley multimodal objective function from scipy.optimize import differential_evolution from numpy.random import rand from numpy import exp from numpy import sqrt from numpy import cos from numpy import e from numpy import pi # objective function def objective(v): x, y = v return -20.0 * exp(-0.2 * sqrt(0.5 * (x**2 + y**2))) - exp(0.5 * (cos(2 * pi * x) + cos(2 * pi * y))) + e + 20 # define range for input r_min, r_max = -5.0, 5.0 # define the bounds on the search bounds = [[r_min, r_max], [r_min, r_max]] # perform the differential evolution search result = differential_evolution(objective, bounds) # summarize the result print('Status : %s' % result['message']) print('Total Evaluations: %d' % result['nfev']) # evaluate solution solution = result['x'] evaluation = objective(solution) print('Solution: f(%s) = %.5f' % (solution, evaluation)) |
Running the example executes the optimization, then reports the results.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
In this case, we can see that the algorithm located the optima with inputs equal to zero and an objective function evaluation that is equal to zero.
We can see that a total of 3,063 function evaluations were performed.
|
1 2 3 |
Status: Optimization terminated successfully. Total Evaluations: 3063 Solution: f([0. 0.]) = 0.00000 |
Further Reading
This section provides more resources on the topic if you are looking to go deeper.
Papers
- Differential Evolution – A Simple and Efficient Heuristic for global Optimization over Continuous Spaces, 1997.
- Differential Evolution: A Survey of the State-of-the-Art, 2011.
Books
- Algorithms for Optimization, 2019.
- Essentials of Metaheuristics, 2011.
- Computational Intelligence: An Introduction, 2007.
APIs
Articles
Summary
In this tutorial, you discovered the Differential Evolution global optimization algorithm.
Specifically, you learned:
- Differential Evolution optimization is a type of evolutionary algorithm that is designed to work with real-valued candidate solutions.
- How to use the Differential Evolution optimization algorithm API in python.
- Examples of using Differential Evolution to solve global optimization problems with multiple optima.
Do you have any questions?
Ask your questions in the comments below and I will do my best to answer.






Thanks Dr. Brownlee,
I would love to see a book with this topics in such a practical way.
Thanks! Me too!
Thanks Dr. Brownlee
You’re welcome.
Great tutorial! Would love to see an implementation of this for a more challenging problem!
Thanks.
can I apply constraints in this function?
Sure.
Can I perform multiobjective optimization with available multiple equality and inequality constraints?
I’m not sure, I think you will need to use a multi-object algorithm/API directly.
Is it possible to use differential evolution algorithms to find the global minimum/maximum of a multivariate function of a machine learning model?
Sure. It will depend on the complexity of the function and the configuration of the optimization algorithm.
Yes.Use
optimize.NonLinearConstraintto do that.Nice!
Thanks Jason! Great example to show how to set it up.
How can I pass an input parameter to the input function that should not be optimised? I’m asking because in my case I’ll calculate other parameters beforehand that are to be used in the objective, but not optimised over.
Best,
Thomas
Sorry, I hadn’t read the documentation properly. I found it 🙂