The Long Short-Term Memory (LSTM) network in Keras supports time steps.
This raises the question as to whether lag observations for a univariate time series can be used as time steps for an LSTM and whether or not this improves forecast performance.
In this tutorial, we will investigate the use of lag observations as time steps in LSTMs models in Python.
After completing this tutorial, you will know:
- How to develop a test harness to systematically evaluate LSTM time steps for time series forecasting.
- The impact of using a varied number of lagged observations as input time steps for LSTM models.
- The impact of using a varied number of lagged observations and matching numbers of neurons for LSTM models.
Kick-start your project with my new book Deep Learning for Time Series Forecasting, including step-by-step tutorials and the Python source code files for all examples.
Let’s get started.
- Updated Apr/2019: Updated the link to dataset.

How to Use Timesteps in LSTM Networks for Time Series Forecasting
Photo by YoTuT, some rights reserved.
Tutorial Overview
This tutorial is divided into 4 parts. They are:
- Shampoo Sales Dataset
- Experimental Test Harness
- Experiments with Time Steps
- Experiments with Time Steps and Neurons
Environment
This tutorial assumes you have a Python SciPy environment installed. You can use either Python 2 or 3 with this example.
This tutorial assumes you have Keras v2.0 or higher installed with either the TensorFlow or Theano backend.
This tutorial also assumes you have scikit-learn, Pandas, NumPy, and Matplotlib installed.
If you need help setting up your Python environment, see this post:
Need help with Deep Learning for Time Series?
Take my free 7-day email crash course now (with sample code).
Click to sign-up and also get a free PDF Ebook version of the course.
Shampoo Sales Dataset
This dataset describes the monthly number of sales of shampoo over a 3-year period.
The units are a sales count and there are 36 observations. The original dataset is credited to Makridakis, Wheelwright, and Hyndman (1998).
The example below loads and creates a plot of the loaded dataset.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 |
# load and plot dataset from pandas import read_csv from pandas import datetime from matplotlib import pyplot # load dataset def parser(x): return datetime.strptime('190'+x, '%Y-%m') series = read_csv('shampoo-sales.csv', header=0, parse_dates=[0], index_col=0, squeeze=True, date_parser=parser) # summarize first few rows print(series.head()) # line plot series.plot() pyplot.show() |
Running the example loads the dataset as a Pandas Series and prints the first 5 rows.
|
1 2 3 4 5 6 7 |
Month 1901-01-01 266.0 1901-02-01 145.9 1901-03-01 183.1 1901-04-01 119.3 1901-05-01 180.3 Name: Sales, dtype: float64 |
A line plot of the series is then created showing a clear increasing trend.
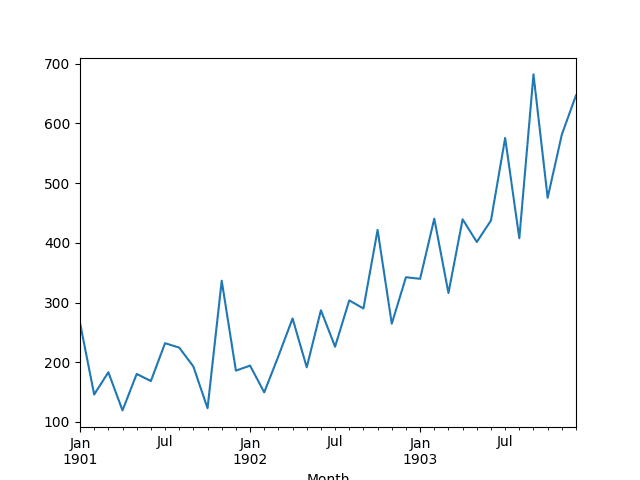
Line Plot of Shampoo Sales Dataset
Next, we will take a look at the LSTM configuration and test harness used in the experiment.
Experimental Test Harness
This section describes the test harness used in this tutorial.
Data Split
We will split the Shampoo Sales dataset into two parts: a training and a test set.
The first two years of data will be taken for the training dataset and the remaining one year of data will be used for the test set.
Models will be developed using the training dataset and will make predictions on the test dataset.
The persistence forecast (naive forecast) on the test dataset achieves an error of 136.761 monthly shampoo sales. This provides a lower acceptable bound of performance on the test set.
Model Evaluation
A rolling-forecast scenario will be used, also called walk-forward model validation.
Each time step of the test dataset will be walked one at a time. A model will be used to make a forecast for the time step, then the actual expected value from the test set will be taken and made available to the model for the forecast on the next time step.
This mimics a real-world scenario where new Shampoo Sales observations would be available each month and used in the forecasting of the following month.
This will be simulated by the structure of the train and test datasets.
All forecasts on the test dataset will be collected and an error score calculated to summarize the skill of the model. The root mean squared error (RMSE) will be used as it punishes large errors and results in a score that is in the same units as the forecast data, namely monthly shampoo sales.
Data Preparation
Before we can fit an LSTM model to the dataset, we must transform the data.
The following three data transforms are performed on the dataset prior to fitting a model and making a forecast.
- Transform the time series data so that it is stationary. Specifically, a lag=1 differencing to remove the increasing trend in the data.
- Transform the time series into a supervised learning problem. Specifically, the organization of data into input and output patterns where the observation at the previous time step is used as an input to forecast the observation at the current time timestep
- Transform the observations to have a specific scale. Specifically, to rescale the data to values between -1 and 1 to meet the default hyperbolic tangent activation function of the LSTM model.
These transforms are inverted on forecasts to return them into their original scale before calculating and error score.
LSTM Model
We will use a base stateful LSTM model with 1 neuron fit for 500 epochs.
A batch size of 1 is required as we will be using walk-forward validation and making one-step forecasts for each of the final 12 months of test data.
A batch size of 1 means that the model will be fit using online training (as opposed to batch training or mini-batch training). As a result, it is expected that the model fit will have some variance.
Ideally, more training epochs would be used (such as 1000 or 1500), but this was truncated to 500 to keep run times reasonable.
The model will be fit using the efficient ADAM optimization algorithm and the mean squared error loss function.
Experimental Runs
Each experimental scenario will be run 10 times.
The reason for this is that the random initial conditions for an LSTM network can result in very different results each time a given configuration is trained.
Let’s dive into the experiments.
Experiments with Time Steps
We will perform 5 experiments, each will use a different number of lag observations as time steps from 1 to 5.
A representation with 1 time step would be the default representation when using a stateful LSTM. Using 2 to 5 timesteps is contrived. The hope would be that the additional context from the lagged observations may improve the performance of the predictive model.
The univariate time series is converted to a supervised learning problem before training the model. The specified number of time steps defines the number of input variables (X) used to predict the next time step (y). As such, for each time step used in the representation, that many rows must be removed from the beginning of the dataset. This is because there are no prior observations to use as time steps for the first values in the dataset.
The complete code listing for testing 1 time step is listed below.
The time steps parameter in the run() function is varied from 1 to 5 for each of the 5 experiments. In addition, the results are saved to file at the end of the experiment and this filename must also be changed for each different experimental run; e.g.: experiment_timesteps_1.csv, experiment_timesteps_2.csv, etc.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 |
from pandas import DataFrame from pandas import Series from pandas import concat from pandas import read_csv from pandas import datetime from sklearn.metrics import mean_squared_error from sklearn.preprocessing import MinMaxScaler from keras.models import Sequential from keras.layers import Dense from keras.layers import LSTM from math import sqrt import matplotlib import numpy from numpy import concatenate # date-time parsing function for loading the dataset def parser(x): return datetime.strptime('190'+x, '%Y-%m') # frame a sequence as a supervised learning problem def timeseries_to_supervised(data, lag=1): df = DataFrame(data) columns = [df.shift(i) for i in range(1, lag+1)] columns.append(df) df = concat(columns, axis=1) return df # create a differenced series def difference(dataset, interval=1): diff = list() for i in range(interval, len(dataset)): value = dataset[i] - dataset[i - interval] diff.append(value) return Series(diff) # invert differenced value def inverse_difference(history, yhat, interval=1): return yhat + history[-interval] # scale train and test data to [-1, 1] def scale(train, test): # fit scaler scaler = MinMaxScaler(feature_range=(-1, 1)) scaler = scaler.fit(train) # transform train train = train.reshape(train.shape[0], train.shape[1]) train_scaled = scaler.transform(train) # transform test test = test.reshape(test.shape[0], test.shape[1]) test_scaled = scaler.transform(test) return scaler, train_scaled, test_scaled # inverse scaling for a forecasted value def invert_scale(scaler, X, yhat): new_row = [x for x in X] + [yhat] array = numpy.array(new_row) array = array.reshape(1, len(array)) inverted = scaler.inverse_transform(array) return inverted[0, -1] # fit an LSTM network to training data def fit_lstm(train, batch_size, nb_epoch, neurons, timesteps): X, y = train[:, 0:-1], train[:, -1] X = X.reshape(X.shape[0], timesteps, 1) model = Sequential() model.add(LSTM(neurons, batch_input_shape=(batch_size, X.shape[1], X.shape[2]), stateful=True)) model.add(Dense(1)) model.compile(loss='mean_squared_error', optimizer='adam') for i in range(nb_epoch): model.fit(X, y, epochs=1, batch_size=batch_size, verbose=0, shuffle=False) model.reset_states() return model # make a one-step forecast def forecast_lstm(model, batch_size, X): X = X.reshape(1, len(X), 1) yhat = model.predict(X, batch_size=batch_size) return yhat[0,0] # run a repeated experiment def experiment(repeats, series, timesteps): # transform data to be stationary raw_values = series.values diff_values = difference(raw_values, 1) # transform data to be supervised learning supervised = timeseries_to_supervised(diff_values, timesteps) supervised_values = supervised.values[timesteps:,:] # split data into train and test-sets train, test = supervised_values[0:-12, :], supervised_values[-12:, :] # transform the scale of the data scaler, train_scaled, test_scaled = scale(train, test) # run experiment error_scores = list() for r in range(repeats): # fit the base model lstm_model = fit_lstm(train_scaled, 1, 500, 1, timesteps) # forecast test dataset predictions = list() for i in range(len(test_scaled)): # predict X, y = test_scaled[i, 0:-1], test_scaled[i, -1] yhat = forecast_lstm(lstm_model, 1, X) # invert scaling yhat = invert_scale(scaler, X, yhat) # invert differencing yhat = inverse_difference(raw_values, yhat, len(test_scaled)+1-i) # store forecast predictions.append(yhat) # report performance rmse = sqrt(mean_squared_error(raw_values[-12:], predictions)) print('%d) Test RMSE: %.3f' % (r+1, rmse)) error_scores.append(rmse) return error_scores # execute the experiment def run(): # load dataset series = read_csv('shampoo-sales.csv', header=0, parse_dates=[0], index_col=0, squeeze=True, date_parser=parser) # experiment repeats = 10 results = DataFrame() # run experiment timesteps = 1 results['results'] = experiment(repeats, series, timesteps) # summarize results print(results.describe()) # save results results.to_csv('experiment_timesteps_1.csv', index=False) # entry point run() |
Run the 5 different experiments for the 5 different numbers of time steps.
You can run them in parallel if you have sufficient memory and CPU resources. GPU resources are not required for these experiments and experiments should be complete in minutes to tens of minutes.
After running the experiments, you should have 5 files containing the results, as follows:
|
1 2 3 4 5 |
experiment_timesteps_1.csv experiment_timesteps_2.csv experiment_timesteps_3.csv experiment_timesteps_4.csv experiment_timesteps_5.csv |
We can write some code to load and summarize these results.
Specifically, it is useful to review both descriptive statistics from each run and compare the results for each run using a box and whisker plot.
Code to summarize the results is listed below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 |
from pandas import DataFrame from pandas import read_csv from matplotlib import pyplot # load results into a dataframe filenames = ['experiment_timesteps_1.csv', 'experiment_timesteps_2.csv', 'experiment_timesteps_3.csv','experiment_timesteps_4.csv','experiment_timesteps_5.csv'] results = DataFrame() for name in filenames: results[name[11:-4]] = read_csv(name, header=0) # describe all results print(results.describe()) # box and whisker plot results.boxplot() pyplot.show() |
Running the code first prints descriptive statistics for each set of results.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
We can see from the average performance alone that the default of using a single time step resulted in the best performance. This is also shown when reviewing the median test RMSE (50th percentile).
|
1 2 3 4 5 6 7 8 9 |
timesteps_1 timesteps_2 timesteps_3 timesteps_4 timesteps_5 count 10.000000 10.000000 10.000000 10.000000 10.000000 mean 102.785197 127.308725 136.182907 146.277122 142.631684 std 6.299329 22.171668 7.760161 5.609412 6.611638 min 92.603903 106.124901 124.724903 138.845314 137.359503 25% 98.979692 114.100891 130.719154 141.906083 138.354265 50% 103.904185 114.519986 137.055840 145.865171 141.409855 75% 108.434727 144.328534 139.615541 150.729938 143.604275 max 110.270559 164.880226 150.497130 155.603461 159.948033 |
A box and whisker plot comparing the distributions of results is also created.
The plot tells the same story as the descriptive statistics. There is a general trend of increasing test RMSE as the number of time steps is increased.

Box and Whisker Plot of Timesteps vs RMSE
The expectation of increased performance with the increase of time steps was not observed, at least with the dataset and LSTM configuration used.
This raises the question as to whether the capacity of the network is a limiting factor. We will look at this in the next section.
Experiments with Time Steps and Neurons
The number of neurons (also called blocks) in the LSTM network defines its learning capacity.
It is possible that in the previous experiments the use of one neuron limited the learning capacity of the network such that it was not capable of making effective use of the lagged observations as time steps.
We can repeat the above experiments and increase the number of neurons in the LSTM with the increase in time steps and see if it results in an increase in performance.
This can be achieved by changing the line in the experiment function from:
|
1 |
lstm_model = fit_lstm(train_scaled, 1, 500, 1, timesteps) |
to
|
1 |
lstm_model = fit_lstm(train_scaled, 1, 500, timesteps, timesteps) |
In addition, we can keep the results written to file separate from the results created in the first experiment by adding a “_neurons” suffix to the filenames, for example, changing:
|
1 |
results.to_csv('experiment_timesteps_1.csv', index=False) |
to
|
1 |
results.to_csv('experiment_timesteps_1_neurons.csv', index=False) |
Repeat the same 5 experiments with these changes.
After running these experiments, you should have 5 result files.
|
1 2 3 4 5 |
experiment_timesteps_1_neurons.csv experiment_timesteps_2_neurons.csv experiment_timesteps_3_neurons.csv experiment_timesteps_4_neurons.csv experiment_timesteps_5_neurons.csv |
As in the previous experiment, we can load the results, calculate descriptive statistics, and create a box and whisker plot. The complete code listing is below.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 |
from pandas import DataFrame from pandas import read_csv from matplotlib import pyplot # load results into a dataframe filenames = ['experiment_timesteps_1_neurons.csv', 'experiment_timesteps_2_neurons.csv', 'experiment_timesteps_3_neurons.csv','experiment_timesteps_4_neurons.csv','experiment_timesteps_5_neurons.csv'] results = DataFrame() for name in filenames: results[name[11:-12]] = read_csv(name, header=0) # describe all results print(results.describe()) # box and whisker plot results.boxplot() pyplot.show() |
Running the code first prints descriptive statistics from each of the 5 experiments.
Note: Your results may vary given the stochastic nature of the algorithm or evaluation procedure, or differences in numerical precision. Consider running the example a few times and compare the average outcome.
The results tell a similar story to the first set of experiments with a one neuron LSTM. The average test RMSE appears lowest when the number of neurons and the number of time steps is set to one.
|
1 2 3 4 5 6 7 8 9 |
timesteps_1 timesteps_2 timesteps_3 timesteps_4 timesteps_5 count 10.000000 10.000000 10.000000 10.000000 10.000000 mean 109.484374 133.195856 133.432933 145.843701 149.854229 std 9.663732 36.328757 19.347675 19.389278 30.194324 min 91.803241 91.791014 87.739484 113.808683 103.612424 25% 104.757265 119.269854 127.937277 137.417983 131.278548 50% 108.464050 129.775765 134.076721 147.222168 151.999097 75% 114.265381 132.796259 147.557091 159.518828 164.741625 max 126.581011 226.396127 156.019616 171.570206 208.030615 |
A box and whisker plot is created to compare the distributions.
The trend in spread and median performance almost shows a linear increase in test RMSE as the number of neurons and time steps is increased.
The linear trend may suggest that the increase in network capacity is not given sufficient time to fit the data. Perhaps an increase in the number of epochs would be required as well.
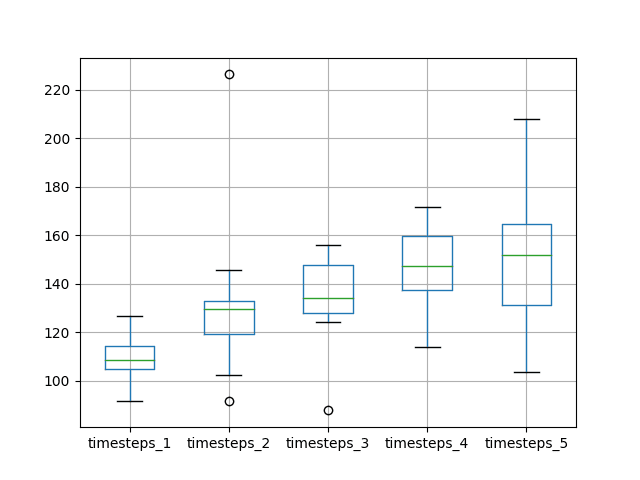
Box and Whisker Plot of Timesteps and Neurons vs RMSE
Extensions
This section lists some areas for further investigation that you may consider exploring.
- Lags as Features. The use of lagged observations as time steps also raises the question as to whether lagged observations can be used as input features. It is not clear whether time steps and features are treated the same way internally by the Keras LSTM implementation.
- Diagnostic Run Plots. It may be helpful to review plots of train and test RMSE over epochs for multiple runs for a given experiment. This might help tease out whether overfitting or underfitting is taking place, and in turn, methods to address it.
- Increase Training Epochs. An increase in neurons in the LSTM in the second set of experiments may benefit from an increase in the number of training epochs. This could be explored with some follow-up experiments.
- Increase Repeats. Using 10 repeats results in a relatively small population of test RMSE results. It is possible that increasing repeats to 30 or 100 (or even higher) may result in a more stable outcome.
Did you explore any of these extensions?
Share your findings in the comments below; I’d love to hear what you found.
Summary
In this tutorial, you discovered how to investigate using lagged observations as input time steps in an LSTM network.
Specifically, you learned:
- How to develop a robust test harness for experimenting with input representation with LSTMs.
- How to use lagged observations as input time steps for time series forecasting with LSTMs.
- How to increase the learning capacity of the network with the increase of time steps.
You discovered that the expectation that the use of lagged observations as input time steps did not decrease the test RMSE on the chosen problem and LSTM configuration.
Do you have any questions?
Ask your questions in the comments below and I will do my best to answer them.








The problem with using lagged values as predictors is that the model misses out the subtle time dependencies which are usually captured by the time series models.
Agreed. The promise of LSTMS is to learn the temporal dependence.
So LSTM will work for all kinds of time series?
Yes, but test other methods and double down on what works best on your problem.
Just wanted to let you know that this is the most lucid explanation of how these LSTM’s work under the hood. Thank you for that. I purchased your Practical Machine Learning Book with the Excel Samples and that was great too! I will certainly try to spread the word.
Thanks.
Hi Jason,
I am really thankful for your articles and posts. It has helped all of us.
I have used walkforward validation and splitted the data in train and test.
after splitting I again used loop and made X_train, y_train with 60 timestep for X_train and y_train as it is.
While fitting the model in LSTM using keras with epoch and batch size, I didnt solve the accuracy. It is constant in all the epoch.
After aplying LSTM also, it has been overfitted. It dont give good result in new.
I have trained with 85% training data. Used scaling in both the sets too still couldnt figure out the solution.
You’re welcome!
Generally, we cannot measure accuracy for regression problems, you must use an error metric:
https://machinelearningmastery.com/faq/single-faq/how-do-i-calculate-accuracy-for-regression
Perhaps some of the suggestions here will help you improve your model performance:
https://machinelearningmastery.com/start-here/#better
Hi Jason,
Your posts are always helpful.
Now, I get two similar data sets. I’d like to train this data using multitask model in keras. To be percise, I have two input data sets and I want to get two output separately in one train model.
Is it possible in keras? I get some content. https://keras.io/getting-started/functional-api-guide/
But I still do not figure it out how. Could you give me some advice?
Almost all neural nets can have multiple output values.
Just frame your dataset and set the number of outputs you require in the output layer of the network.
Another question. Compared with tensorflow, a fine-tuned keras model will get a better result or a worse one? Is it comparable?
Keras is built on top of TensorFlow. Comparing results from the two does not make sense (at least to me).
Thank you for your reply.
Have a good day.
Hi Jason,
could you elaborate this line
train = train.reshape(train.shape[0], train.shape[1])
isn’t this the same?
It does look that way, I may have been too excited with all the resizing. Try removing it and see if all is well.
“Lags as Features. The use of lagged observations as time steps also raises the question as to whether lagged observations can be used as input features. It is not clear whether time steps and features are treated the same way internally by the Keras LSTM implementation.”
Any further thoughts on this?
I’m a little confused on how to use timesteps when some input features are lagged and some are not. (really, I’m fundamentally confused as to why timesteps exists at all, given that it would seem any lagged input should just be treated as features). There’s surprisingly little clear information on the matter of LSTM timesteps on the internet… I don’t recall ever coming across the concept of timesteps in any of Schmidhuber, et al papers, either (perhaps I wasn’t paying attention!)
Thanks for the great resource you’ve put together and continue to share, btw.
Yes, I was wrong.
Features are weighted inputs. Timesteps are discrete inputs of features over time. (does that make sense, it reads poorly…)
The key to understanding timesteps is the BPTT algorithm. I have a post on this scheduled.
“I’m a little confused on how to use timesteps when some input features are lagged and some are not. (really, I’m fundamentally confused as to why timesteps exists at all, given that it would seem any lagged input should just be treated as features). There’s surprisingly little clear information on the matter of LSTM timesteps on the internet…”
This is 100% my question, I’ve done so much Googling (and read multiple of Jason’s posts) and I still don’t understand this at all. Cannot figure out how to prep lagged time steps + features for LSTM.
Lagged obs are time steps in LSTMs.
LSTM input is 3d: [samples, time steps, features]. If your series is univariate, you one many time steps and one feature. If you want to classify one day of data, you have one sample, 25 hours of time steps and one feature.
Does that help?
Totally agree with you.
In your previous blog(https://machinelearningmastery.com/time-series-prediction-lstm-recurrent-neural-networks-python-keras/),
you use “trainX = numpy.reshape(trainX, (trainX.shape[0], 1, trainX.shape[1]))”(1),
and now you use “X = X.reshape(X.shape[0], timesteps, 1)” (2).
If the second parameter means timestamp. then in (1), you may use “look_back” in that article instead of 1
If the third parameter means one var, then in (1), you may use 1 instead of trainX.shape[1], because trainX.shape[1] means look_back or timesteps in this article.
I would recommend using past observations as timesteps when inputting to the model.
Could it be overfitting with more neurons? since more neurons means more degrees of freedom, so the model can (over) fit the training data well, while generalize poorly.
If that’s the case, more epochs won’t help though, we need more training data.
Yes, more neurons can result in overfitting.
Hi Jason – thank you for the great content. Really enjoyed your ML Python recipes. I am having some trouble understanding the structure of the input data for the LSTM, since everywhere I look seems to suggest something different.
I understand that the input X has the shape (samples, timesteps, features). My use case is I have about 100 time series, and I’m trying to use them as features to forecast another time series 5 steps ahead at a time (updating as new information in the rolling window method you detailed in a different post). What will the structure of X look like in my case? I currently have something like this:
X Y
[[t0, t1, t2], [[t3, t4]
[t1, t2, t3], [t4, t5]
… …
for each feature, which I’ve then stacked together into a 3D shape using np.stack( ). But it seems like this is incorrect, since the timesteps should be 2, not 3? Am I coming at this the right way? The timestep/feature/lag confusion seems to be prevalent on the Internet. Also each feature might have greater predictive power at different lag/leads, will this LSTM setup potentially bottleneck my accuracy, and is there a better approach to this? Thanks!!
If you have 5 series then that would be 5 features.
I would recommend loading the data as a 2d matrix then using reshape, perhaps with 1 sample.
Does that help?
Hi Jason, great post.
I have been trying to implement Keras LSTM using R. How can I reshape my univariate data frame to the input shape required by LSTM in R.
Sorry, I don’t have material on using Keras in R.
Ohh that’s unfortunate. Although I did find reshape layer in keras, but I am not sure if it is same as numpy.reshape.
Also when i used it to train a model, it converted the train set into 3D array but now i cannot evaluate the model since I am stuck on trying to convert test set in to 3D array. Thanks.
What error are you getting?
I was able to fix the problem by reshaping the train set to 3D array with timesteps = 1 and including lagged values as input.
But I cannot set timesteps more than 1.
E.g. I have a dataset with time interval of every 15 mins. If I set timestep to 96(1 Day) and built a LSTM model then I cannot forecast on test(1 Month) set since I get only (2880/96 = ) 30 values and not 2880 values.
hey thank you very much for this posts !!
they are really useful
I’m glad to hear that.
Hi Jason, great post.
I have a bit of problem to understand the timeseries_to_supervised function however. It seems to shit the data so that the dataframe columns are in the following order [t, t-1, t-2, t-3, t-4, t+1]. T+1 will be used as y, but the rest of the columns are to my understanding in a wrong order (i.e, newest data point is first and vice versa)… Or did I miss something important.
maybe some bugs are here
I think the function is so
Learn more about the function here:
https://machinelearningmastery.com/convert-time-series-supervised-learning-problem-python/
I have used a few prints to better understand my question.
First 6 rows:
0 -120.1
1 37.2
2 -63.8
3 61.0
4 -11.8
5 63.3
First 6 rows after supervised:
0 0 0
0 NaN NaN -120.1
1 -120.1 NaN 37.2
2 37.2 -120.1 -63.8
3 -63.8 37.2 61.0
4 61.0 -63.8 -11.8
5 -11.8 61.0 63.3
And then dropping the NAN’s:
[[ 37.2 -120.1 -63.8]
[ -63.8 37.2 61. ]
[ 61. -63.8 -11.8]
[ -11.8 61. 63.3]
[ 63.3 -11.8 -7.3]
[ -7.3 63.3 -31.7]
[ -31.7 -7.3 -69.9]
…..]
Then scaling:
[[ 0.04828702 -0.83250961 -0.496628 ]
[-0.496628 0.03130148 0.17669274]
[ 0.17669274 -0.52333882 -0.21607769]
[-0.21607769 0.1619989 0.1891017 ]
[ 0.1891017 -0.23778144 -0.1917993 ]
[-0.1917993 0.17462932 -0.32344214]
…… ]
Then dividing into X and y:
X: [[ 0.04828702 -0.83250961]
[-0.496628 0.03130148]
[ 0.17669274 -0.52333882]
[-0.21607769 0.1619989 ]
[ 0.1891017 -0.23778144]
[-0.1917993 0.17462932]
…..]
y: [-0.496628 0.17669274 -0.21607769 0.1891017 -0.1917993 -0.32344214…]
Then reshaping:
[[[ 0.04828702]
[-0.83250961]]
[[-0.496628 ]
[ 0.03130148]]
[[ 0.17669274]
[-0.52333882]]
[[-0.21607769]
[ 0.1619989 ]]
[[ 0.1891017 ]
[-0.23778144]]
[[-0.1917993 ]
[ 0.17462932]]
….]
Now here is my problem and I think it’s the same as in Jussi’s post.
I’m not sure if I’m right, but if I take for example, the first one, I would use:
[[[ 0.04828702]
[-0.83250961]]]
for X and
[-0.496628]
for y, so that i have t2,t1 -> t3
Then I think that I would learn the wrong order, if the timesteps are learned from top to bottom.
Should the order not be as follows:
[[[-0.83250961]
[ 0.04828702]]]
for X and
[-0.496628]
for y, so that i have t1,t2 -> t3
Am I right, that this is wrong? Thank you
Hi jason,
i’m getting this error ValueError: time data ‘1901-Jan’ does not match format ‘%Y-%m’. is there something wrong with it?
Ensure that you remove the footer from the data file.
I am bit confused with case timesteps=1. Isn’t using timestep=1 same as using traditional Neural Network?
Almost. LSTMs still have internal state. To make good use of BPTT, times steps should be more than 1:
https://machinelearningmastery.com/gentle-introduction-backpropagation-time/
Hi Jason, thanks for your great post,i have learned much.
I have a question about the difference, is it necessary to do difference(to make stationary)? In time series analysis it is necessary, but in here Neural Network why should we do that?
Generally, making the problem simpler makes it easier to model which in turn makes the forecast more accurate.
Hi Jason, thanks for sharing this post with us. I am doing similar forecasting analysis and really enjoy reading it.
I have question on the preprocessing part. Please correct if I missed something: I noticed that there is a time shift for all the features including the predicted value(y), and this method also is applied on the test set. From what I saw is that you first do time shift, then do train-test split and finally generate input and output, which will result in a y(t-1) feature for the last “real-value” output.
If this is exact what I thought, I think there should not be a time shift in the test set because the output value doesn’t even exist before you make a prediction on this time stamp. And this may cause that the test set already contains the real value for prediction in its feature.
What I thought is that you only can make predictions step by step in the test set. First generate the prediction of the first time stamp, and use that prediction in computing the output of next time stamp.
Please correct me if I missed something. And this problem has been lingering in my head for several days.
Thanks!
Yes, ideally we would use walk forward validation to test sequence prediction models:
https://machinelearningmastery.com/backtest-machine-learning-models-time-series-forecasting/
Hi Jason,
Thanks for your tutorial really helpful. Do you have other tutorials in the same domain but in Convolutional LSTM in Keras?
Not for time series.
Jason,
First of all, thanks for all your work here – I simply could not have made the progress I have without this site.
With your help I’ve got MLPs, LSTMs and Bidirectional LSTMs up and running. However my LSTMs are all single timestep and it is the multi-timestep step I now want to crack.
I’ve been looking at this code at each stage to see how you build the data and you’ve got me scratching my head. In short, it looks to me as if the time sequences are wrongly ordered:
(borrowing from Chris’ post further up:
First 6 rows:
0 -120.1
1 37.2
2 -63.8
3 61.0
4 -11.8
5 63.3
First 6 rows after supervised:
0 0 0
0 NaN NaN -120.1
1 -120.1 NaN 37.2
2 37.2 -120.1 -63.8
3 -63.8 37.2 61.0
4 61.0 -63.8 -11.8
5 -11.8 61.0 63.3
And then dropping the NAN’s:
[[ 37.2 -120.1 -63.8]
[ -63.8 37.2 61. ]
[ 61. -63.8 -11.8]
[ -11.8 61. 63.3]
[ 63.3 -11.8 -7.3]
[ -7.3 63.3 -31.7]
[ -31.7 -7.3 -69.9]
…..]
My issue is that the first three timestep values are: -120.1, 37.2 and -63.8 and yet the first timestep sequence is: [ 37.2 -120.1 -63.8 ] when I would expect it to be: [-120.1 37.2 -63.8 ].
All the other timestep sequences follow the same pattern (of course). Am I completely misunderstanding this (which is perfectly possible of course) in which case what rule should be followed, especially as the no. of timesteps is increased?
Once again, thanks for a great site and looking forward to your response
Regards,
Steve
Generally neural networks are poor at time series. You can learn more here:
https://machinelearningmastery.com/faq/single-faq/how-do-i-use-lstms-for-time-series-forecasting
Nevertheless, I have an example of multistep time series forecasting with LSTMs that might help as a template:
https://machinelearningmastery.com/multi-step-time-series-forecasting-long-short-term-memory-networks-python/
Further to my previous post. If I’m right and I’m quite prepared to be wrong ….
A simple tweak to timeseries_to_supervised changing:
columns = [df.shift(i) for i in range(1, lag+1)]
to …
columns = [df.shift(i) for i in range(lag, 0, -1)]
… produces the result:
[-120.1, 37.2, -63.8],
[ 37.2, -63.8, 61. ],
[ -63.8, 61. , -11.8],
[ 61. , -11.8, 63.3],
[ -11.8, 63.3, -7.3],
[ 63.3, -7.3, -31.7],
… which makes more sense to me in terms of presenting the steps in the original order of the input data.
One other observation: this result is produced using a timesteps value of 2. A more ‘understandable’ approach might be to tweak the code so that in this case the output is produced with a timesteps value of 3 but that’s just me and my OCD 🙂
I look forward to your response whether I’m right or wrong – I just want to learn the correct way to pre-process my data in order to get what I hope will be the best performance from my LSTM models.
Regards,
Steve
what is the benefits of using timestep for example (N=3) in rnn.. and does N=3 is better than using a timestep of N=1??
It depends on the problem.
What is difference between timesteps and unrolling an lstm network ? When you see the classical picture of an unrolled lstm does this something has to do with timesteps?
You can learn more about unrolling here:
https://machinelearningmastery.com/rnn-unrolling/
What do you recommend now? Should the time steps be as lag variable as n additional features or would prefer the internal time step functionality of the LSTM network in keras? I hope I have not overlooked such a recommendation, but you may be able to give me the clarity.
Try both and see what works best for your problem.
Also, start with an MLP and only use LSTM if it outperforms the MLP.
I am following you post from past 2 months. Thank you for writing such an amazing block.
I was just trying to understand how lags work in a LSTM model, which you explained quite well.
A question came to my mind when I read https://machinelearningmastery.com/multi-class-classification-tutorial-keras-deep-learning-library/ post with regards to what you explained above.
If I want to create lags to predict multi classification time series problem and I ahve three classes in my predictor variable. Do I need to do One hot encoding first and then create time lags on all three variables or do I need to create time lags first and then do one hot encoding over the predictor variable.
It might be easier to encode the output variable after you prepare the lag inputs.
This post might help in preparing the data:
https://machinelearningmastery.com/convert-time-series-supervised-learning-problem-python/
Trying to do waste generation prediction for a time series dataset using lstms. Any ideas of datasets that could help and codes will be grateful
I list datasets here:
https://machinelearningmastery.com/time-series-datasets-for-machine-learning/
lets say i have a data set with 1000 rows and 6 features out of which i want to make prediction.I want to find the regressive value ‘y’ at each row.But i want the network to remember 10 previous observations to make the next observation.How should i change the data
[1000,1,5]
or
[100,10,5]
Sounds like you want 10 time steps. The second example.
Hi Janson,
Thanks for those tuto on LSTM !
There are still a few points that remain unclear for me :
Taking a univariate toy example where you try to predict the next Temperature, if you decide to include lag observations :
1. is it better to include them as features part of a unique timestep or to consider them as several timesteps with one features ?
2. what is the impact on the network training process ?
Thanks in advance for you help on that,
Regards
LSTMs can read multiple timesteps as input.
Therefore, we can provide multiple lag observations to the model and have it predict the next step.
Hi Jason,
Thank you very much for this tutorial. I have one question about lag observations. Is it possible to apply lag observations for sequence classification in LSTM? I mean, given an X=[X1, X2,…, Xn] input sequence, I want to classify current Xt refer also to past observations.
Thanks in advance
Best regards
Yes, lag observations are input as timesteps to the LSTM.
hi jason, can you explain briefly what is time-step?
A sample is a sequence.
A timestep is a point at which observations can be made.
A feature is an observation at a time step.
Hi, I’m working on a time-series prediction system lately. And I wondered the timestep used for the inputs could change to the output, for example enter 10 timesteps and optimize the prediction for 3 timesteps.
Is that possible?
Now if I enter 10 timesteps I can only get 10 at the output, and I can only optimize the last one (return sequence = false) or optimize all of them (return sequence = true).
There would be some way to optimize only for 3 timesteps
I think you’re referring to multi-step prediction?
I have general information here:
https://machinelearningmastery.com/multi-step-time-series-forecasting/
I have many tutorials on the topic, for example:
https://machinelearningmastery.com/how-to-develop-lstm-models-for-multi-step-time-series-forecasting-of-household-power-consumption/
Hi Jason,
I thank you so much for this tutorial, I have a question that I posted in SO if you don’t mind passing by, https://datascience.stackexchange.com/q/41305/33279 , my specific case is that for each train data for each time point, there are many entries. each is a learning entry, so I can’t sample (group) and sum, mean, max or min any entry. Knowing that time index was construct from two columns: year and month, and I want learn from other features along with these two columns.
Many thanks !
Perhaps you can summarize your problem in a sentence or two?
Hi, Jason.
The LSTM posts you have posted are very useful.
However, since the accuracy of the model cannot be printed, it is questionable in reliability.
Data is only output in late 500 to late 600 values in the range of 500 to 10000.
Can you tell me how to increase the scope of data in these areas and how to add the output of accuracy?
Thanks, Jason.
Classification accuracy is not appropriate for regression problems. Instead we must use error.
I recommend reading this:
https://machinelearningmastery.com/classification-versus-regression-in-machine-learning/
Hi Jason,
Thank you for another great post and for sharing your knowledge with the community.
I’m having some difficulty to understand the difference between an LSTM with 1 timestep and an MLP. Although everything I read so far indicates that they should have the same behavior, in my experimental results the LSTM with 1 timestep performs significantly better than the MLP. Also, increasing the timestep to values greater than 1 does not improve performance.
I’m training the networks with batches of 128 samples with 126 features each. Is it possible that the LSTM training is behaving as if the number of timesteps was equal to the batch size?
Thank you for your help!
The main difference is the internals of the nodes, and the shared internal state of the units over the samples in the batch.
Hi Jason, what if my time series stamp is not unique. For example, at one specific time stamp, I have different combination of features and the corresponding outputs. Thanks!
Perhaps normalize the dataset so you have the same features at all time steps, even if some are 0 values?
Hi Jason,
First of all, thank you so much for this amazing website. Although this is the first time I am posting a comment, I have been using its many resources for a while now !
I have a question about LSTM in time series prediction tasks. It is my understanding that one of the advantage of LSTMs is the capacity to remember past examples and control what is stored in memory with the gated units. Knowing this, how come that we need to include past features and why can’t we limit ourselves to using only one feature corresponding to the last timestep that we have? I know that using larger timesteps increases the performance, my question is why?
Thank you
The time steps define the past you are providing to the model that it might remember for future predictions.
Perhaps I don’t understand your question?
My conception of how LSTMs and RNNs process data was flawed, but I now understand why it is necessary. The only concept of time in a LSTM is with regard to one sequence. How all the different sequences fit together is not relevant.
There is still something that bothers me though: how can we process sequences of variable length if the LSTM expects a fixed size sequence? Of course, we can use padding but I see this more as a workaround than as a real solution… Any idea?
LSTMs can be used dynamically, e.g. one time step at a time, but it very slow.
So, instead we vectorize the data into fixed length sequences, use padding and use a masking layer to ignore the padded values.
Hi !
I can’t understand your “timeseries_to_supervise function” .
Why time-steps are like for instance, ( t-1,t-2,t-3,t) and not (t-3,t-2,t-1,t) ?
Our sequences are in this way : in X : (t-1,t-2,t-3) and y : (t) and it seems weird to me.
Thanks
Perhaps start with this post:
https://machinelearningmastery.com/convert-time-series-supervised-learning-problem-python/
Hi Jason,
Thank you for this post, I have a question, I have an LSTM for predicting energy consumption using time series, I thought I had a good model because I had an RMSE of 0.005 but when I tried to calculate the R2 score I had 0.44. Do you think that this is a bad model and I should change it? or should I just try to modify the architecture of my LSTM and tune my model.
I recommend comparing the skill of the model to a baseline model, such as a naive persistence forecast. Skill is relative to the baseline.
when we test with a trained lstm model, do we need to make the test timestep is the same as train
No.
Hi Jason.
Thanks for the post.
I didn’t understant why we need to run one epoch 500 times on a loop instead of 500 epochs.
If i change the looped activation function to tanh (i think the default is sigmoid, is it?), could I use 500 epochs?
Thanks.
When I wrote looped activations function, I meant ‘recurrent_activation’ parameter from keras LSTM layer, which is set to default ‘hard_sigmoid’.
Thanks.
They are equivalent.
I am curious why you used reshape in “X = X.reshape(X.shape[0], timesteps, 1)”
If you simply use reshape it for more than 1 timestep, it seems to lose time dependency between batches and it ruins whole training X dataset because it is stateful learning which needs time dependency between batches… I think, for more than 1 timestep, it needs to use window sliding for X and stateful=False for the model. what do you think about it?
This might help understand the nature of time steps in a sample:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Hi Jason. Found your site after searching for “LSTM time series forecasting output is lagged”.
We have built our data set in what seems the same as you, where input features, and the output have their t0 input features and their t+60 values also as inputs.
Some of our models seemed to have worked, but recently, all training seems to produce a forecast that is lagged by t+60. Example output of what i mean. The AI is catching the inflections exactly t+60 after the event we want to predict.
Data: ______/
AI: ________/
Is this common? How do you guide the NN back into shape?
it removed my spaces and made my ascii graph look like it is exactly working…
Data: ______/
…..AI: ________/
and specifically, do you cover this problem in any of your materials?
thanks
Yes, this is common, it suggests that the model is not skillful, see this:
https://machinelearningmastery.com/faq/single-faq/why-is-my-forecasted-time-series-right-behind-the-actual-time-series
Hey Jason amazing job, I have a question.
Why we need to run one epoch 500 times on a loop instead of 500 epochs.(take it from Thiago)
Thanks.
We are doing walk-forward validation to evaluate the model AFTER it is fit.
Sorry I am new at Deep Learning, and programing, how does walk-foward validation works, does this improve the performance of my model?
Good question, see this tutorial:
https://machinelearningmastery.com/backtest-machine-learning-models-time-series-forecasting/
So what will be the best method to prepare the data:
First:
Inputs Outputs
1,2,3 4,5,6
2,3,4 5,6,7
3,4,5 6,7,8
Second:
Inputs Outputs
1,2,3 4,5,6
1,2,3,4 5,6,7
1,2,3,4,5 6,7,8
Perhaps experiment and see what framing makes sense and/or is learnable.
Sorry, now is clearly:
First:
_Inputs_____Outputs
__1,2,3_______4, 5,6
__2,3,4_______5, 6,7
__3,4,5_______6, 7,8
Second:
_Inputs______Outputs
__1,2,3________4,5,6
__1,2,3,4______5,6,7
__1,2,3,4,5____6,7,8
You typically do not expand your input shape, you limit it at a fixed size.
Son, after that I have build the model with an lstm layer with 10 neurons and a dense layer with 3,does it looks fine? or should I do any change? , and after that how do I fit the model? , should I use validation data?, sorry for asking that much as I say I’m new and programing and doing it in R so I train to have everything you explained clear to code it in R
“Does a model look fine” is subjective for your problem, this can help in terms of diagnostics:
https://machinelearningmastery.com/learning-curves-for-diagnosing-machine-learning-model-performance/
How to improve performance – that is a massive topic, you can start here:
https://machinelearningmastery.com/start-here/#better
Validation data can be used normally, but for sequence data / time series data it’s challenging/not reasonable as you must use walk-forward validation:
https://machinelearningmastery.com/backtest-machine-learning-models-time-series-forecasting/
I don’t have examples of deep learning in R and cannot offer advice on the topic.
Thanks for all your help
You’re welcome.
Thanks
You’re welcome.
Dear Mr Jason Brownlee,
I really enjoyed your blog and I learnt a lot from it.
You said that you are using a walk forward validation like this one
https://machinelearningmastery.com/backtest-machine-learning-models-time-series-forecasting/
Could you please clarify where did you use it in your code ?
Thank you
Thanks.
Line 99 of the complete code example where we walk across the test set.
Thank you !
Your blog is so helpful !
Hello Jason,
I have one question:
why do I need to specifiy the time step? Isn’t the nature of a LSTM that it uses cell state information of previous observations anyway?
E.g. if I use timestep = 1, does the LSTM not use any cellstate and hidden state information of previous observations?
Good question, LSTM expect inputs to have be provided with a structure of time steps and features, you can learn more here:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Hi Jason,
Thank you for the insightful post.
So, if we want to use lags as features because the logic of the problem requires to use past features, would you recommend using MLP with lagged features over LSTMs with lagged features?
I would recommend testing a suite of models such as linear methods, ml methods, mlp, cnn, lstm, hybrids and discover what works best for your specific dataset.
hi jason
why you dont plot your models schema?
im really confused in the models whether this is a one to one or many to one
You can, call model.summary()
I had a small doubt, While I have time series data, I have varying lengths of data recorded at different time steps, For instance I have 10 records for year 2020 in different features and 5 records from 2010 and increasing so on. Due to this while I’m splitting data, I’m having large errors at new features in timesteps. Is it ok If I randomly scatter the sequence and lower the loss to get more accurate predictions.
You could try zero padding to make all input samples the same length, then use a masking layer to skip the padded values.
Hi Jason,I have came across with different examples when talking about walk forward validation and it have raised in my this doubt. How should the data by prepare.
1st Example.
______Training___________________Testing
Inputs________Outputs______Input_______Outputs
_1,2,3__________4,5,6________4,5,6________7,8,9
_2,3,4__________5,6,7________5,6,7________8,9,10
_3,4,5__________6,7,8________6,7,8________9,10,11
2nd Example.
______Training
Inputs________Outputs
_1,2,3__________4,5,6
_2,3,4__________5,6,7
_3,4,5__________6,7,8
______Testing
Inputs_______Outputs
_4,5,6_________7,8,9
_5,6,7_________8,9,10
_6,7,8_________9,10,11
Perhaps this code will help:
https://machinelearningmastery.com/convert-time-series-supervised-learning-problem-python/
Thanks, another great job, but doesn’t clarify at all the doubt. The essential part of the doubt is regarding the testing or validation data, should be part in each window? or should be out of sample? or perhaps both ways could be call as walk forward validation?
A model is always evaluated on out of sample data.
The way that this can be achieved for time series is via walk-forward validation comprised of a training set and a test set where each step or window of the test set becomes part of the training set progressively. There is no validation dataset.
Perhaps I don’t understand your question, if so, could you please rephrase or elaborate it?
Thanks again, that it’s exactly what I was asking. So, should the data be pre-processed as in the first example?
I have read again and see that the questions aren’t clearly? Sorry for that. I gonna try explain myself better.
When I build a model how should I test it’s effectiveness before real world aplication?
Examples:
First
Training a window and test it with the consecutive one. And the expected effectiveness should be equal to the average of all the test results.
Second
Training the model whit a sample of windows and testing it whit a consecutive sample of windows. And the expected effectiveness should be equal to the average of the test results.
Define a test period, say the last week/month/year of data.
Use walk-forward validation test harness and report the mean error, e.g. RMSE or MAE. Size of the window could be one time step or whatever is most appropriate to your project/dataset.
Establish a baseline using a persistence model, any model that does better than this has skill.
Does that help?
Yes, thanks for all your great work.
Hi Jason, do we necessarily need to make a time series stationary before modelling it with a LSTM model?
No, but it is a good idea generally (yes).
Hi Jason,
If we have a time series in which samples do not have equivalent lags, what do we have to do? Is it important that the time interval between 2 consecutive samples be the same?
Bests
Zero padding to the same length or truncate to the same length, more here:
https://machinelearningmastery.com/data-preparation-variable-length-input-sequences-sequence-prediction/
Thanks for your reply. The dataset I am using contains the trajectory of taxicabs in Rome city over 320 taxis. It has four columns: “car id, time, latitude, longitude.” Even for each car time interval between 2 consecutive samples is not the same. It varies from 1 second to 20 seconds. I read the information in the link above, but this kind of padding is not working here. Do you have any suggestions? If I consider a fixed timestamp, for example 10 seconds, and get rid of extra data, do you think it is a good idea, or will lose some useful information?
Perhaps you can use a model to transform the sequence to a fixed size, e.g. like an autoencoder:
https://machinelearningmastery.com/lstm-autoencoders/
I have a dataset with 1000 samples, 1 timestep, and 1 feature (1000,1,1). I see a lot of discussions where they create subsequences (split one timestep column into multiple by using some X number – ex: (100,10,1)) and use that as input to LSTM autoencoder and argue that LSTMs/autoencoder can only learn when the data has multiple timesteps. But, I get good results with just 1 timestep column. What is the correct input here? Is it incorrect if I use univariate (1000,1,1) input?
You must discover what works best for your specific model and dataset.
Thank you for the. reply. So is it okay using just one timestep for an LSTM autoencoder? Does it still learn temporal dependencies?
In this post (https://towardsdatascience.com/lstm-autoencoder-for-anomaly-detection-e1f4f2ee7ccf), the author used a one time stamp but all the comments were related to the question I asked earlier.
It can if your model maintains state across samples.
I recommend this tutorial:
https://machinelearningmastery.com/how-to-develop-lstm-models-for-time-series-forecasting/
But I see you split the univariate timestep into subsequences in this link. Am I missing something here?
Yes, it is called a window approach, see this:
https://machinelearningmastery.com/time-series-forecasting-supervised-learning/
That’s’ great! So just to be sure, if you have 100 rows (100,1) of univariate data, you transform them using the window approach and keep the tensor as (98,2,1)?
That seems right, or something like that.
Hi Jason,
Thank you for another great work!
I am still confused about time step as lag vs lagged features with time step =1.
To clarify that, let say I have 3 time series x1, x2 and x3 as inputs and y as output. Now I would like tov implement lag into the lstm model.
Scenario 1 is that I use ( sample, time step = lag, features =3) as shape for dataset.
Scenario 2 is that I use lagged features like x1(t-1), x1(t-2), x1(t-3), x2(t-1), x2(t-2), x2(t-3), x3(t-1), x3(t-2), x3(t-3) as inputs with shape of ( sample, time step = 1, features =9).
To me, scenario 1 would make more sense since the essence of lstm is to remember pattern of pervious steps of inputs to predict next time steps of output. However scenario 1 is limited to define one unique value for lag. If I want to use different lags for each feature, like lag = 3 for x1, lag= 5 for x2 and lag = 10 for x3, I cannot use scenario 1 and need to use scenario 2. However scenario 2 is like a MLP network with lagged features and does not use the power of lstm for the pattern recognition. So what would you recommend in the case at which we have different lags for features?
Thank you in advance!
Best regards,
Amir
It can be tricky, I recommend reading this:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
You explanation is not correct.
Based on the API implementation, times steps does not increase the capability of the LSTM. It only decrease the frequency of updating the trainable parameters.
This explains why when time steps is smaller, the accuracy is higher.
Please see more here.
https://stats.stackexchange.com/questions/377091/time-steps-in-keras-lstm
Perhaps this will help:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
HI Jason,
Awesome blog, congrats!
I’m using and studying LSTM for a while and now I’ve a real situation where I can apply that, however I’ve a doubt here about how make future predictions.
For example:
1 – I trained my LSTM model over 4 years of sales, grouped by day, month and year and I sum one specific field which represents the quantity.
2 – I used time steps of 30 (representing 1 month of data);
To evaluate the model I use 1 year of data from 2020, when I predict the test data I-ve to do something like this:
X_test = []
for i in range(30, 317):
X_test.append(inputs[i - 30 : i, 0])
X_test = np.array(X_test)
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
Where inputs it’s my entire dataset (train + test), the problem is , how can i predict the 2021 year for example?
May get the predicted data and fit in the network again?
Thanks,
Antonio
I would recommend using walk-forward validation to evaluate your model, this will help:
https://machinelearningmastery.com/backtest-machine-learning-models-time-series-forecasting/
Hello,
I always wonder about LSTM(RNN) that follow,
there is ‘timestep’ parameter.
it means how many past data you consider to predict future value.
and if timestep = 50, then the RNN loop will be 50
I wonder if timestep=50 means short-term dependency and the whole data means long-term dependency.
This will help:
https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input
Hi Jason! Thanks for the tutorial!
I am working on a problem where I have the future values of one of the features in my dataset (say var2). How can I implement this to predict var1(t)?
Note: var2 is not predictable (e.g. holidays) and var2 is correlated with the value of var1. Hence need to provide var2(t) as input.
In the actual problem, I am using 7 timesteps and 5 features out of which obs(t) for 2 variables is known.
var1(t-1) var2(t-1) var3(t-1) var2(t) var1(t)
1 50.0 0.0 23 1 51
2 51.0 1.0 24 2 52
3 52.0 2.0 25 3 53
4 53.0 3.0 26 4 54
5 54.0 4.0 27 5 55
6 55.0 5.0 28 6 56
7 56.0 6.0 29 7 57
8 57.0 7.0 30 8 58
9 58.0 8.0 31 9 59
You have complete control over the inputs and outputs of your model.
Design your prediction problem and model so that it only uses data that you have available at prediction time.
Sorry, now it’s readable.
|————————INPUT————————|—-output—-|
—-var1(t-1) –var2(t-1) –var3(t-1) –var2(t) —–var1(t)
1 –50.0 ———-0.0 ———-23 ———-1 ———–51
2 –51.0 ———-1.0 ———-24 ———-2 ———–52
3 –52.0 ———-2.0 ———-25 ———-3 ———–53
4 –53.0 ———-3.0 ———-26 ———-4 ———–54
5 –54.0 ———-4.0 ———-27 ———-5 ———–55
6 –55.0 ———-5.0 ———-28 ———-6 ———–56
7 –56.0 ———-6.0 ———-29 ———-7 ———–57
8 –57.0 ———-7.0 ———-30 ———-8 ———–58
9 –58.0 ———-8.0 ———-31 ———-9 ———–59
Hello Jason!
During reading about LSTMs in this page, I came across the explanation (https://machinelearningmastery.com/faq/single-faq/what-is-the-difference-between-samples-timesteps-and-features-for-lstm-input) but some points are still not 100% clear for me.
I am quite confused about the difference between an LSTM with only one time step and a MLP. In what manner do they both differ from each other? Is an LSTM with only one time step able to learn the temporal correlation between samples? If we predict with this LSTM model for 100 sequential samples, does it predict better towards the last test samples, does it gain sensible hidden states which could improve the prediction of next sample? Or will it be the same as the case where we seperately predict every time (which is actually the prediction behavior of a MLP)?
I appreciate your explanatory answers.
Thanks,
Hakan
With one time step, you have no temporal correlation into the network. Hence LSTM cannot learn it. However, MLP and LSTM differ because LSTM has hidden state H and the inner cell C, so it is more complicated than the what MLP would do. But if there’s only one time step, probably you are not using LSTM to the full extent.
Hello Adrian,
thank you for the great explanation (and thank Jason for the great post, whose results I found very surprising).
What I understand here is that, for the dataset used, if we set only one time step (and reset the model every epoch using batches of one sample only, like Jason does in his code), there’s no difference whatsoever between MLP and LSTM in terms of “learning form the past”, the only difference between the models being their structure (so how many weights, how the weights are connected, which is more complicated for LSTM, etc).
You say that “if there’s only one time step, probably you are not using LSTM to the full extent” but Jason experiments demonstrates that, for the shampoo dataset, and for the “goodness” indicator chosen (RMSE) there’s is no point in increasing the number of time steps beyond 1. So there’s no point in using lstm and perhaps a proper choice of hyperparmenters of an MLP would potentially give comparable results.
Is my reasoning correct?
Many thanks, and again congrats for the great work.
Thanks for your response!
But what if we have multiple neurons (‘units’) and only one timestep? E.g. the layer ‘LSTM(32, input_shape=(num_timesteps=1, num_features=10))’. What kind of an advantage do the 32 hidden units bring when we feed only 1 time step data? Does the higher number of units improve the model in terms of temporal correlation, even if we have several time steps or doesn’t it impact on the capability of learning the temporal correlation?
Each LSTM unit is a matrix multiplication to the input. At a high level, you may consider it try to extract one particular feature out of the input. Hence with more units, you attempt to look for more information, and based on that many kind of information, you make your prediction.
hi, i’ve been working on a project and it’s somewhat similar but more complex, instead of forecasting shampoo sales, i’m trying to predict how much a client will spend the next month. i have prepared the data into samples(CSVs for each client containing invoice dates and the total spent on that day). my first impulse was to use an LSTM and it still is, but i’m still trying to figure out how one LSTM will be enough for thousands of clients! do you any ideas that could be useful? thanks anyway, your articles have been guiding me all along my journey of data science 🙂
One LSTM may not be enough — depends on the nature of the data. But if the clients behave homogenous then you will see your forecasting quite accurate.
Hi, Jason
What can I conclude if increasing lookback/timesteps doesn’t effect the loss.
I have hourly load data and I have varied lookback from 3, 6, 12, 24, 48, 72, 120, 144, 168 but all have MAPE between 2 & 3.
Hi Syed…You may find the following of interest:
https://machinelearningmastery.com/get-the-most-out-of-lstms/